Abstract
Background
Bacterial blight (BB) caused by Xanthomonas oryzae Pv. oryzae (Xoo) is one of the most serious diseases of rice worldwide. Oryza officinalis Wall ex Watt, harboring abundant genetic diversity and disease resistance features, are important resources of exploring resistance genes with broad-spectrum resistance to BB. However, the molecular mechanisms and genes of BB resistance in O. officinalis have been rarely explored.
Results
Here, the BB resistance of four different origin O. officinalis populations in Yunnan were identified by seven representative hypervirulent Xoo races, which exhibited different BB resistance among four populations, in which the BB resistance of the Gengma_Lincang population was the strongest. In addition, the pathogenetic ability of seven Xoo races to O. officinalis was different in that the pathogenicity of PXO99 was stronger than that of C5. There were no remarkable differences in leaf microstructures among four O. officinalis populations, revealing the differences in resistance of four O. officinalis to BB are caused by the endogenous resistance genes. Furthermore, our results proved that there were no nine cloned BB resistance genes in four populations but possessed dominant Xa5, dominant Xa13, and recessive xa3/xa26 homologous alleles of xa5, xa13, and Xa3/Xa26 resistance genes. These three homologous genes were isolated and cloned from four populations and named OoXa5, OoXa13, and Ooxa3/xa26. The expression profile revealed that the expression levels of OoXa13 and Ooxa3/xa26 were significantly down-regulated under PXO99 and C5 stress, especially in the Gengma_Lincang population, suggesting the O. officinalis might enhance BB resistance by down-regulating the expression level of OoXa13 and Ooxa3/xa26.
Conclusion
The BB resistance genes of O. officinalis had its own characteristics by expression pattern and BLAST analysis of OoXa5, OoXa13, and Ooxa3/xa26, which indicated that there might be new genes or molecular mechanism of BB resistance in O. officinalis. Our studies provided a solid foundation and reference for revealing the molecular mechanism of BB resistance in O. officinalis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice is the main cereal crop for half of the world’s population, while rice bacterial blight (BB) caused by Xanthomonas oryzae pv. oryzae (Xoo) is the fourth serious bacterial disease in the world (Mansfieldi et al. 2012; Rasabandith 1998). BB has been reported in all major rice-growing regions of the world (Nino-Liu et al. 2006). Nowadays, the most economical and effective strategy to control BB is cultivating resistant cultivars, but the number of resistance genes identified from the cultivated rice is very limited, and more cultivars with narrow-spectrum BB resistance genes become susceptible to diseases with the emergence of new physiological races. Therefore, it is very important to identify more resistance resources and to explore new resistance genes for breeding resistant cultivars with broad-spectrum resistance to BB.
Wild rice species, the ancestor of cultivated rice, is an important gene pool harboring lots of good characteristics, such as strong resistances to blight disease, blast, insects, drought, and coldness, and the resistance genes which can be used to improve the genetics and agronomic traits of the cultivated rice (Fan et al. 2000; Fan 2000; Qin et al. 2000). The number of wild rice species in the Yunnan Province is the largest among all provinces in China (Cheng et al. 2004; Zhang 2002; Gao et al. 2000). Oryza officinalis is resistant or tolerant to many common rice diseases. Zhang et al. (1994) found that 50% of the highly resistant materials are O. officinalis among the Chinese wild rice species. The Gengma_Yunnan O. officinalis population has high resistance to BB and brown planthopper and medium resistance to rice blast (Peng et al. 1982). There are several O. officinalis populations in Yunnan, and these populations have some differences in leaf size, plant height, and width. However, the BB resistance levels and the resistance genes of O. officinalis populations remain unclear. Meanwhile, there were many physiological races of Xoo in which their pathogenetic ability to Yunnan O. officinalis was unclear. So far, there is lack of knowledge about systematic identification of Yunnan O. officinalis populations against Xoo races.
Previous studies have identified 42 rice BB resistance genes (Vikal and Bhatia 2017), but only nine genes have been cloned, including Xa1 (Yoshimura et al. 1998), Xa3/Xa26, xa5 (Iyer-Pascuzzi and McCouch 2004), Xa10 (Gu et al. 2008), xa13 (Chu et al. 2006a; Sanchez et al. 1999), Xa21 (Song et al. 1995; Ronald et al. 1992), Xa23 (Fan et al. 2011; Zhang et al. 1998), xa25 (Lee et al. 2003), and Xa27 (Gu et al. 2004). The identification and cloning of BB resistance genes provide gene resources for resistance breeding of rice. However, due to the race specific genes and the evolution of new Xoo races, the existing BB resistance genes are severely restricted in the application of resistance breeding (Bhasin et al. 2012; Kameswara et al. 2002). O. officinalis has high resistance to BB and may possess new BB resistance genes. Seven reported and cloned BB resistance genes, including Xa1, Xa3/Xa26, xa5, xa13, Xa21, Xa23, and Xa27, were detected in O. officinalis using molecular markers and functional markers tightly linked to them (Li et al. 2015). However, it is not clear if these BB resistance genes exist in all the Yunnan O. officinalis populations. What is more, the functional domain and resistance level of BB resistance genes between O. officinalis and cultivated rice have not been reported.
In this study, seven representative hypervirulent Xoo races were used to systematically identify the resistance of four O. officinalis populations, Gengma_Lincang, Menghai_Xishuangbanna, Jingne_Jinghong, and Lancang_Puer, of the Yunnan Province. Meanwhile, the molecular markers linked to nine cloned BB resistance genes were used to detect O. officinalis to confirm whether these genes exist in the four populations. The detected Xa5, Xa13, and xa3/xa26 homologous genes were also cloned from the four populations. Expression patterns of the three homologous genes under Xoo stress were analyzed by qRT-PCR. In addition, the leaf microstructures, relating to the propagation and spreading velocity of Xoo, of four O. officinalis populations were observed by microscopy. Our studies provided a solid foundation and reference for revealing the molecular mechanism of BB resistance in O. officinalis.
Materials and Methods
Plant Materials and Hypervirulent Xoo Races
Oryza officinalis Wall ex Watt populations were collected from four places of Yunnan Province, viz., Gengma_Lincang, Menghai_Xishuangbanna, Jingne_Jinghong, and Lancang_Puer, and planted in a greenhouse. The materials of IRBB1(Xa1), IRBB5 (xa5), IRBB10 (Xa10), IRBB13 (xa13), Oryza longistaminata (Xa21), Oryza rufipogon (Xa23 and Xa27), Minghui63 (Xa25 and Xa3/Xa26), IR24 (Xa5), IR64 (Xa13), and 02428 japonica were also planted in a greenhouse. Seven hypervirulent Xoo races (Table S1) were used to screen the four O. officinalis populations for identification of BB resistance.
Identification of BB Resistance Among Four O. officinalis Populations
The seven Xoo races were cultured on nutrient agar medium at 28 °C for 48 h. The culture was suspended in sterile water, and the concentration was adjusted to OD600 = 0.8–1.0 using the NanoDrop2000 and, then, used to inoculate four O. officinalis populations along with the susceptible Oryza sativa cultivar 02428 at the booting stage by leaf-clipping method (Kauffman et al. 1973) at 14:00–15:00 time under 28–30 °C room temperature. At least three plants were inoculated by each Xoo race, and each leaf was cut 1–2 cm from the tip of the leaf.
The lesion length and lesion area were measured and calculated after inoculation with Xoo for 21 d. The average of the three plants was taken as the resistance index, and lesion length and lesion area were used to classify resistance or susceptibility by the BB national standard classification (Waheed et al. 2009) (Table S2). Meanwhile, the pathogenetic ability to four O. officinalis populations of the seven Xoo races was analyzed according to the leaf lesion area.
The Microstructure Observation of Four O. officinalis Populations
The middle part of the third fully expanded leaf was selected to crosscut by free-hand section. The thin slices were observed and photographed by inverted fluorescence microscope (Leica DMI4000B). The macro-vascular bundle area, air cavity area, and leaf thickness were measured by the data analysis software of Leica. The numbers of macro-vascular bundle, small-vascular bundle, and air cavity of the leaf midveins were also counted.
The PCR Identification of Nine Cloned BB Resistance Genes in Four O. officinalis Populations
Nine cloned BB resistance genes, including Xa1, xa5, xa13, Xa21, Xa23, Xa26, Xa27, Xa10, and xa25, were detected in four O. officinalis populations. In the identification of Xa23 gene in O. officinalis, the EST molecular marker C189, which was tightly linked to it, was utilized (Li et al. 2015). The Xa21 and Xa27 genes were identified based on the functional markers designated by the polymorphic loci of their sequences (Li et al. 2015; Fan et al. 2011). In the amplification of the other six cloned BB genes, the specific primers designated by their conserved sequences were utilized (Table S3). Meanwhile, two primers provided by Hur et al. (2013) were used to identify the Xa3/Xa26 or xa3/xa26 in O. officinalis. The genomic DNA of all the materials was extracted from fresh leaves followed by the CTAB method (Murray and Thompson 1980). The 50 μL PCR reaction system contained 1 μL DNA template (50 ng L−1), 2 μL each primer (10 μmol L−1), 25 μL 2 × Power Pfu PCR MasterMix (TaKaRa, Dalian, China), and 20 μL ddH2O. The PCR amplification program was as follows: pre-denaturation at 94 °C for 2 min; 94 °C for 15 s, 52–65 °C for 15 s, and 72 °C for 30 s–1 min, at 30 cycles; and 72 °C for 2 min. The PCR products were separated by the 2–4% agarose gel electrophoresis.
Isolation and Cloning of Xa5, Xa13, and xa3/xa26 Homologous Genes from Four O. officinalis Populations
In order to clone the Xa5, Xa13, and xa3/xa26 homologous genes from the four populations, the primers were designated by Primer 5.0 using their ORF sequences (Table S4). Total RNA was extracted from the four populations following the plant RNA kit (Omega Bio-Tek, Georgia, USA), and the first-strand cDNAs were produced by retranscription of total RNA with PrimeScript™ RT reagent (TaKaRa). The cDNA template of the four populations and PrimeSTAR®Max DNA Polymerase (TaKaRa) were used to establish 50 μL PCR reaction system to clone OoXa5, OoXa13, and Ooxa3/xa26. The PCR amplification program was as follows: pre-denaturation at 98 °C for 2 min; 98 °C for 10 s, 50–55 °C for 5–15 s, 72 °C for 5–15 s, at 35 cycles. The PCR products were separated by the 2–4% agarose gel electrophoresis, and, then, the products with the expected sizes were isolated by gel extraction kit (TaKaRa) and cloned into pClone007 vector (TSINGKE) to sequence.
BLAST Analysis of OoXa5, OoXa13, and Ooxa3/xa26
The ORF of OoXa5, OoXa13, and Ooxa3/xa26 was searched through the ORF online software. The three fragments of BF26, SE26, and LA26 were spliced by DNAMAN to obtain the complete ORF of Ooxa3/xa26. The sequences of Xa5/xa5 (Nipponbare and IR24/IRBB5), Xa13/xa13 (IR24 and IR64/IRBB13), and xa3/xa26/Xa3/Xa26 (IR24/Minghui63 and IRBB3) were downloaded from NCBI. The nucleotides and amino acids of OoXa5, OoXa13, and Ooxa3/xa26 were aligned with Xa5/xa5, Xa13/xa13, and xa3/xa26/Xa3/Xa26, respectively, using the DNAMAN. The functional domains of Ooxa3/xa26 from the four populations and xa3/xa26/Xa3/Xa26 were predicted by an online tool (http://smart.embl-heidelberg.de/).
Expression Pattern Analysis of OoXa5, OoXa13, and Ooxa3/xa26 in Response to Xoo
Total RNA was extracted from the leaves after inoculation with ddH2O, PXO99, and C5 for 0, 24, 48, 72, 96, and 120 h according to the manufacturer’s instructions of the plant RNA kit (Omega Bio-Tek). A total of 1 μg RNA was reverse-transcribed into cDNA using PrimeScript™ RT reagent with the gDNA Eraser kit (TaKaRa). The quantitative real-time polymerase chain reaction (qRT-PCR) primers of OoXa5, OoXa13, and Ooxa3/xa26 were designed by an online tool (https://www.ncbi.nlm.nih.gov/tools/primer-blast) based on their ORF sequences (Table S5). Gene expression levels were determined by performing qRT-PCR in Applied Biosystems QuantStudio 6 Flex (ABI, USA) using SYBR Premix Ex Taq II (TaKaRa) according to the manufacturer’s instructions. Data were analyzed by QuantStudio 6 Flex software (ABI, USA) and the 2−△△CT method (Bustamante et al. 2014; Livak and Schmittgen 2001).
Results
The Identification of BB Resistance Among Four Yunnan O. officinalis Populations
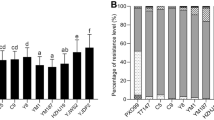
The BB resistance of four O. officinalis populations was different by testing the lesion area, among which the Gengma_Lincang population had the strongest resistance (Figs. 1a and 2). The BB resistance level of the Gengma_Lincang population was HR or R, while the resistance level of the other three populations was similar from MR to R (Figs. 1a–d, 2, and Table S6). Although the resistance level of the four populations against seven Xoo races was different, all of them had strong resistance to BB. The characteristics of high resistance to BB were particularly outstanding in the Gengma_Lincang population, which showed high resistance to multiple Xoo races (BD8438, T7147, C9, and C5), suggesting the Gengma_Lincang population might have richer BB resistance genes than the other three populations or the Gengma_Lincang population had a BB resistance gene family which might confer resistance specific to each Xoo race.
Bacterial blight disease reaction of four different O. officinalis populations and cultivar 02428 against different Xoo races. All the materials were inoculated with seven Xoo races at the booting stage, and the top second leaves were chosen for inoculation. The lesion length and lesion area were analyzed at the 21st day after inoculation with Xoo races
The comparison of leaf lesion area in four O. officinalis populations inoculated with seven Xoo races. The lesion area of the four populations was the largest after inoculation with PXO99, while the lesion area of the four populations was the smallest after inoculation with C5. The lesion area of the Gengma_Lincang population was the smallest among the four populations after inoculation with seven Xoo races. (a) Gengma, Lincang population; (b) Menghai, Xishuangbanna population; (c) Jingne, Jinghong population; (d) Lancang, Puer population; (e) cultivar 02428
Seven Xoo races had different pathogenicity to O. officinalis. The resistance reaction of the same O. officinalis population showed obvious difference to different Xoo races. The lesion length and lesion area of the four populations inoculated with PXO99 were the longest and largest, whereas the lesion length of the four populations inoculated with C5 was the shortest (Fig. 2), especially more than 90% of the Gengma_Lincang population showed high resistance (HR) to C5. The lesion almost extended from the inoculation site to the bottom of the leaf in cultivar 02428, which was highly susceptible to seven Xoo races (Fig. 1e, Table S6). Therefore, it was suggested that the PXO99 might be highly virulent to O. officinalis as compared to C5.
Comparison of Leaf Microstructures Among Four O. officinalis Populations
The resistance of four O. officinalis populations to BB was different, while the numbers and area of vascular bundles in the leaf vein are related to the propagation and spreading velocity of Xoo. Therefore, we compared the leaf microstructures of four O. officinalis populations. We found that there were no significant differences in the microstructures of the leaf midvein and lateral vein among the four populations (Figs. 3 and 4). In addition, there was no remarkable difference in the number of macro-vascular bundles, small-vascular bundles, air cavities of leaf midvein, and leaf thickness among the four populations (Table 1, Fig. S1). The microstructures of the leaf midvein and lateral vein between the four populations and cultivar 02428 were significantly different (Figs. 3e and 4e). There were many bulging vascular bundles in the lateral vein of cultivar 02428 (Fig. 4e showed with elliptic ring), which were not found in O. officinalis populations. Although there were some differences in the arrangement of vascular bundles between O. officinalis and cultivar 02428, there were no significant differences among the four populations. Interestingly, the area of macro-vascular bundles was not the smallest in the Gengma_Lincang population with the strongest BB resistance, while the area of macro-vascular bundles in the Jingne_Jinghong population was larger than that of the susceptible cultivar 02428 (Fig. S1b). These results suggested that the differences in BB resistance among the four O. officinalis populations might not be caused by the leaf microstructures but by the endogenous resistance genes.
The comparison of the microstructures in the leaf midvein among the four O. officinalis populations (50 ×). The middle part of the third leaf at the booting stage was chosen for observation of the leaf microstructures. There were no significant differences in the microstructures of the leaf midvein among the four populations. (a) Gengma, Lincang population; (b) Menghai, Xishuangbanna population; (c) Jingne, Jinghong population; (d) Lancang, Puer population; (e) cultivar 02428
The comparison of the leaf microstructures in the lateral vein among the four O. officinalis populations (100 ×). The middle part of the third leaf at the booting stage was chosen for observation of leaf microstructures. There were no remarkable differences in the microstructures of the lateral vein. (a) Gengma, Lincang population; (b) Menghai, Xishuangbanna population; (c) Jingne, Jinghong population; (d) Lancang, Puer population; (e) cultivar 02428
Detection of Nine Cloned BB Resistance Genes in O. officinalis Populations
The PCR detection of xa5, xa13, Xa21, Xa23, and Xa27 genes showed that Xa21, Xa23, and Xa27 could not be detected any band, while xa5 and xa13 genes were amplified 170 and 280 bp susceptible bands, respectively, from the four O. officinalis populations (Fig. 5). The Xa1, Xa10, Xa3/Xa26, and xa25 genes were identified based on the primers designed by their ORF sequences. The results showed that only Xa3/Xa26 was detected in the 1100 bp bands (Fig. 5), which were further confirmed to belong to the xa3/xa26 homologous gene with 750 bp susceptible bands in the four populations (Fig. 6). Therefore, our results demonstrated that four O. officinalis populations possessed dominant Xa5, dominant Xa13, and recessive xa3/xa26 homologous alleles of xa5, xa13, and Xa3/Xa26 resistance genes.
The detection of nine cloned BB resistance genes in four O. officinalis populations. There were about 170 bp (Xa5) and 280 bp (Xa13) susceptible bands and 1100 bp Xa3/Xa26 or xa3/xa26 bands in the four populations. The other six BB resistance genes were not amplified any band from the four populations. (M) D2000 marker and 50 bp DNA ladder; (1) cultivar 02428; (2) positive controls with the corresponding BB resistance genes; (3) Gengma, Lincang; (4) Menghai, Xishuangbanna; (5) Jingne, Jinghong; (6) Lancang, Puer
The detection of Xa3/Xa26 and xa3/xa26 in four O. officinalis populations. (a) There were about 750 bp bands belonging to xa3/xa26 gene in the four populations. (b) There was no 255 bp resistance band of Xa3/Xa26 in the four populations. (M) D2000 marker; (1) Gengma, Lincang; (2) Menghai, Xishuangbanna; (3) Jingne, Jinghong; (4) Lancang, Puer; (5) IR24 with xa3/xa26; (6) Minghui63 with Xa3/Xa26
The Cloning and BLAST Analysis of OoXa5, OoXa13, and Ooxa3/xa26
The Xa5, Xa13, and xa3/xa26 homologous genes were isolated and cloned from the four populations (Fig. 7) and named OoXa5, OoXa13, and Ooxa3/xa26. The amino acid alignment showed that OoXa5 belonged to Xa5 susceptible gene for the 39th valine (V) (Fig. S2), which was consistent with the 170 bp susceptible band of PCR detection (Fig. 5). One amino acid was different at the 238th site of amino acid sequences between dominant Xa13 and recessive xa13. Although OoXa13 lacked the 238th and 237th sites of amino acids (Fig. S3), we detected 280 bp susceptible bands from the four populations using the primers designed by the different promoter regions between Xa13 and xa13 (Fig. 5). The 280 bp bands from the four populations were sequenced and aligned with the Xa13 gene from IR64 (Fig. S4), which proved that the four populations carried the promoter regions of the Xa13 gene. Therefore, we conjectured that OoXa13 should be the allele of xa13 rather than xa13 resistance gene. However, the coding regions between OoXa13 and reported Xa13 were remarkably different, which indicated the BB resistance genes of O. officinalis might have its own characteristics, and O. officinalis might have new BB resistance genes and molecular mechanism different from the cultivated rice.
The cloning of OoXa5, OoXa13, and Ooxa3/xa26 from the four O. officinalis populations. (M) D2000 marker; (1) Gengma, Lincang; (2) Menghai, Xishuangbanna; (3) Jingne, Jinghong; (4) Lancang, Puer; (BF) the first fragment of Ooxa3/xa26 ORF; (SE) the middle fragment of Ooxa3/xa26 ORF; (LA) the last fragment of Ooxa3/xa26 ORF
There are eight different amino acids between xa3/xa26 and Xa3/Xa26 in the LRR domains (Fig. S5, shown with *) (Hur et al. 2013). Except one amino acid, the other seven amino acids of Ooxa3/xa26 were consistent with xa3/xa26 in the LRR domains. The Xa3/Xa26 resistance gene has the TGCA sequence in 452–456 bp from the start codon, whereas the recessive xa3/xa26 has the AATC sequence at the same sites (Hur et al. 2013). Ooxa3/xa26 had the AATC (429–432 bp) sequence as well as xa3/xa26 at the corresponding sites (Fig. S6). According to the BLAST analysis of amino acids and nucleotides, Ooxa3/xa26 from the four O. officinalis populations should belong to xa3/xa26 homologous allele of Xa3/Xa26 resistance gene, which was consistent with the 750 bp susceptible bands (Fig. 6).
Expression Profile Analysis of OoXa5, OoXa13, and Ooxa3/xa26 in Response to Xoo
To evaluate the functions of OoXa5, OoXa13, and Ooxa3/Ooxa26 in response to Xoo, we analyzed the expression patterns of them in O. officinalis under PXO99 and C5 stress. The expression levels of OoXa13 and Ooxa3/xa26 genes were significantly down-regulated, while the expression of OoXa5 was almost not significantly different under PXO99 and C5 stress (Figs. 8, 9, and 10). The whole expression levels of OoXa5 were higher in CK that dealt with ddH2O than those of under PXO99 and C5 stress (Fig. 8), which indicated that PXO99 and C5 might have no effect on the expression of OoXa5 and the changes of OoXa5 expression level might be caused by mechanical damage during the inoculation. The expression levels of OoXa13 and Ooxa3/xa26 were significantly down-regulated under PXO99 and C5 stress, while their expression level was up-regulated in CK (Figs. 9 and 10). In addition, the expression levels of OoXa13 and Ooxa3/xa26 were lower in the Gengma_Lincang population than those of the three populations after inoculated with Xoo. Meanwhile, the OoXa13 and Ooxa3/xa26 expression levels continually decreased in the Gengma_Lincang population under C5 stress from 24 to 120 h (Figs. 9a and 10a). These results suggested that the expression levels of OoXa13 and Ooxa3/xa26 were remarkably suppressed in four O. officinalis populations under PXO99 and C5 stress, and the suppression effect was the most significant in the Gengma_Lincang population.
The expression levels of OoXa5 gene in the four O. officinalis populations under Xoo stress. The whole expression levels of OoXa5 showed no significant change under PXO99 and C5 stress. * represented the expression levels of OoXa5 at 24, 48, 72, 96, or 120 h which were significantly different compared with 0 h (p < 0.05). The blue, red, and green curves represented the expression trend of OoXa5 under ddH2O, PXO99, and C5 stress, respectively. CK, ddH2O treatment; PXO99, PXO99 stress treatment; C5, C5 stress treatment
The expression levels of OoXa13 gene in the four O. officinalis populations under Xoo stress. The expression of OoXa13 was significantly down-regulated under PXO99 and C5 stress. * represented the expression levels of OoXa13 at 24, 48, 72, 96, or 120 h which were significantly different compared with 0 h (p < 0.05). CK, ddH2O treatment; PXO99, PXO99 stress treatment; C5, C5 stress treatment
The expression levels of Ooxa3/xa26 gene in the four O. officinalis populations under Xoo stress. The expression of Ooxa3/xa26 was significantly down-regulated under PXO99 and C5 stress. * represented the expression levels of Ooxa3/xa26 at 24, 48, 72, 96, or 120 h which were significantly different compared with 0 h (p < 0.05). CK, ddH2O treatment; PXO99, PXO99 stress treatment; C5, C5 stress treatment
The Functional Domain Comparison Between Ooxa3/xa26 and Xa3/Xa26 or xa3/xa26
There were significant differences in the functional domains between Ooxa3/xa26 from the four O. officinalis populations and xa3/xa26 and Xa3/Xa26 from cultivated rice (Fig. S7). The xa3/xa26 (IR24) contained Pkinase and Pkinase_Tyr pfams at amino acid sequence of 800–1103 aa, while the Xa3/Xa26 (Minghui63 and IRBB3) have three pfams of Pkinase, Pkinase_Tyr, and ABC1 at the same amino acid sequence sites. There was, however, only one type of S_TKc pfam in Ooxa3/xa26 at the corresponding sites. In addition, there were three more LRR domains in Ooxa3/xa26 than xa3/xa26 and Xa3/Xa26. Interestingly, the domain of Ooxa3/xa26 was LRR at the amino acid sites of 563–587 aa in Gengma_Lincang population, while it was LRRTYP in the other three populations. The difference of LRR in Gengma_Lincang was due to a different amino acid (Leucine, L) at 585 aa with CTT codon, while the amino acid was phenylalanine (F) with TTT codon in the other three populations.
Discussion
The BB Resistance of Gengma_Lincang Population Was the Strongest Among Four O. officinalis Populations
There are many reported BB resistance identification of O. officinalis. Yunnan Gengma O. officinalis has high resistance to BB (Li et al. 2015; Cheng et al. 2004; Peng et al. 1982). However, there is lack of systematical resistance identification of different Yunnan O. officinalis populations. In this study, seven representative hypervirulent Xoo races (Table S1) were chosen to identify the BB resistance of the four Yunnan O. officinalis populations from different original places, and some of the results were consistent with those of the previous studies. The four populations had strong resistance to seven Xoo races, but the resistance of the four populations was significantly different, in which the Gengma_Lincang population had the strongest resistance among them. In addition, the pathogenicity to the four populations of seven Xoo races was different, in which the pathogenicity of PXO99 was the strongest while C5 was the weakest. Xoo mainly invades the rice host through the leaf water stoma or wound and then propagates and spreads in the vascular bundle (Leach et al. 1989). Therefore, the numbers and area of vascular bundles of the leaf might affect the propagation and spread velocity of Xoo in the leaf. We found that the resistance of the four populations to BB was significantly different, but there were no remarkable differences in their leaf microstructures, deducing the differences in BB resistance among the four populations were not caused by the leaf microstructures but by the endogenous resistance genes. The Gengma_Lincang population had the strongest resistance among the four populations, indicating this population might have a BB resistance gene family which conferred resistance specific to each BB race. Meanwhile, the expression levels of some key resistance genes might be induced under C5 stress and O. officinalis showed high resistance to C5. However, the resistance genes, no matter being induced by PXO99 or C5 in O. officinalis, might have strong resistance to other Xoo races.
There Might Be New BB Resistance Genes or Molecular Mechanism in O. officinalis
The different BB resistance among the four O. officinalis populations were determined by the internal resistance genes but not caused by their leaf microstructures. First of all, we detected nine cloned BB resistance genes to confirm whether they exist in the four O. officinalis populations. Li et al. (2015) found that O. officinalis possesses Xa5 and Xa13 homologous genes; however, it is not clear whether the O. officinalis contains xa3/xa26 or Xa3/Xa26. In this study, we confirmed that the four O. officinalis populations possessed Xa5, Xa13, and xa3/xa26 homologous genes, which were cloned from the four populations and named OoXa5, OoXa13, and Ooxa3/xa26, respectively. Our results proved that there were no nine cloned BB resistance genes in the four populations. Thus, we concluded that the difference in BB resistance among the four populations were not caused by the leaf microstructures or the nine cloned BB resistance genes, which conjectured that O. officinalis might contain new BB resistance genes or molecular mechanism that was different from the cultivated rice.
The Contribution of OoXa5, OoXa13, and Ooxa3/xa26 to the BB Resistance
Both Xa5 and xa5 genes have been shown previously to be constitutively expressed in rice leaf, and their expression levels were not remarkably different after inoculation with Xoo (Jiang et al. 2006; Iyer-Pascuzzi and McCouch 2004). The 39th glutamate of the dominant Xa5 gene, exposed to the surface of the protein crystal structure, interacts with the avirulence protein, and, then, the Avrxa5 binds to the promoters of genes involved in susceptibility. Evidence has been presented that Xa5 may be a nuclear target of Xanthomonas spp. which interacts with one or more Xoo TAL effectors, thus activating disease-promoting genes; however, the resistant protein xa5 in the xa5 homozygote prevents the activation of the disease-promoting genes because there is no interaction between a Xoo effector and xa5, or such interaction is non-productive, and, then, the rice shows resistance to BB (Huang et al. 2016; Gu et al. 2009; Iyer-Pascuzzi et al. 2008; Sugio et al. 2007). The xa5 is a completely recessive gene that mediates resistance in rice by restricting bacterial movement (Iyer-Pascuzzi et al. 2008). Our study found that the whole expression levels of OoXa5 were higher in CK that dealt with ddH2O than those of under PXO99 and C5 stress and the PXO99 and C5 had no significant effect on the expression of OoXa5. This result suggested that the expression change of OoXa5 under Xoo stress might be caused by mechanical damage during inoculation, which was consistent with that of the previous studies (Jiang et al. 2006; Iyer-Pascuzzi and McCouch 2004). However, whether the OoXa5 would activate the expression of susceptible genes like Xa5 gene and then affect the resistance to Xoo in O. officinalis needs further study.
The recessive xa13 resistance gene is a novel R gene with resistance to Xoo race PXO99, but the expression of xa13 was not affected by PXO99 (Yuan et al. 2010, 2011; Chu et al. 2006b). Suppression of Xa13 expression significantly increases the resistance of rice to PXO99, and suppressing the expression of xa13 further enhances the xa13-mediated resistance (Yuan et al. 2011; Chu et al. 2006b). Copper, an essential micronutrient of plants and an important element for a number of pesticides in agriculture, suppresses Xoo growth (Yuan et al. 2010). Xa13 protein cooperates with two other proteins, COPT1 and COPT5, to modulate copper redistribution and promote removal of copper from xylem vessels, where Xoo multiplies and spreads to cause disease (Yuan et al. 2010, 2011). PXO99 is more sensitive to copper than other Xoo races (Yuan et al. 2010, 2011). The expression levels of OoXa13 in the four O. officinalis populations were suppressed significantly under PXO99 and C5 stress. The suppressing expression of OoXa13 would decrease the copper transport complexes of OoXa13 with COPT1 and COPT5, which would greatly reduce the removal of copper from the xylem vessels. Finally, the copper concentration in the xylem vessels was kept at high level which could effectively inhibit the propagation of Xoo and enhance the resistance of O. officinalis to PXO99 and C5. The expression level of OoXa13 was the lowest and significantly suppressed in the Gengma_Lincang population. The lower expression level of OoXa13 suggested that the copper transport complexes formed by OoXa13, COPT1, and COPT5 would be fewer in the Gengma_Lincang population and effectively suppress removal of copper from the xylem vessels. Therefore, we speculated that this might be one of the reasons that the BB resistance of the Gengma_Lincang population was stronger than that of the other three populations. The xa13-mediated resistant/susceptible signaling pathway does not belong to the types currently known and represents a new type of plant disease resistance (Chu et al. 2006b). The sequence alignment and analysis showed OoXa13 from the four O. officinalis populations and Xa13 of the cultivated rice had obvious differences (Fig. S3), suggesting O. officinalis might have its own resistant characteristic or new molecular mechanisms of BB resistance different from the cultivated rice.
Xa3/Xa26 gene, like the most plant disease resistance genes, belongs to constitutive expression genes. Overexpression of Xa3/Xa26 can enhance the resistance of transgenic plant to BB (Cao et al. 2007). The four O. officinalis populations contained recessive xa3/xa26 homologous allele of Xa3/Xa26 resistance gene. The expression levels of Ooxa3/xa26 in the four populations were significantly down-regulated under PXO99 and C5 stress, especially in the Gengma_Lincang population. The inhibiting effect of PXO99 and C5 on Ooxa3/xa26 expression was stronger in the Gengma_Lincang than the other three populations. However, there are no related reports that suppressing the expression of xa3/xa26 can enhance the resistance of rice to BB. The functions of Ooxa3/xa26, which would enhance resistance to PXO99 and C5 by suppressing its own expression level like the Xa13 gene, should be further studied.
The Effect on BB Resistance of Ooxa3/xa26 LRR Domain in Gengma_Lincang O. officinalis Population
Sun et al. (2006) reported that the evolution of the Xa3/Xa26 gene family is rapid and the point mutations of LRR involved in pathogen recognition are the major force of evolution. Xa3/Xa26 belongs to a multigene family, consisting of Xa3/Xa26, MRKa, MRKc, and MRKd. Complementary analysis showed that MRKa and MRKc cannot mediate resistance to BB when regulated by their native promoters, but MRKa confers partial resistance to BB when regulated by a strong constitutive promoter; MRKd however is a pseudogene (Xu et al. 2008; Cao et al. 2007). Overexpression of MRKa can enhance partial resistance of rice to BB (Cao et al. 2007). Although MRKa and MRKc cannot mediate BB resistance, they may be once effective genes for Xoo resistance. Some members of the Xa3/Xa26 gene family once may lose the resistance due to the evolution and variation of the Xoo, and then these genes regain resistance and become new resistance genes for point mutations (Xu et al. 2008; Cao et al. 2007). These results indicated that the Xa3/Xa26 gene family is constantly evolving to respond to the variation of pathogen, suggesting there may be some resistance genes of the Xa3/Xa26 gene family that have not yet been identified or cloned.
LRR participates in the identification of different physiological races to determine the resistant specificity of the Xa3/Xa26 gene (Zhao et al. 2009). The domain was LRR at the sites of 563–587aa in the Gengma_Lincang Ooxa3/xa26, whereas it was LRRTYP in the other populations and cultivar at the corresponding sites. The difference in LRR of the Gengma_Lincang Ooxa3/xa26 was caused by one different amino acid (leucine, L) at 585 aa with CTT codon, while the amino acid was phenylalanine (F) with TTT codon in the other materials. One-point mutation of Ooxa3/xa26 caused a domain change, which might change the BB resistance level of Ooxa3/xa26. Based on the previous studies (Zhao et al. 2009; Cao et al. 2007), the change of the LRR domain in the Gengma_Lincang Ooxa3/xa26 might be due to the point mutations when the LRR identified the pathogen. The expression level of Ooxa3/xa26 in the Gengma_Lincang was the lowest under PXO99 and C5 stress among the four populations. Whether this was related with the changed LRR domain or would the Ooxa3/xa26 of the Gengma_Lincang regain the resistance and become a new BB resistance gene during co-evolution with the pathogen should be further studied.
Conclusion
The BB resistance genes of O. officinalis had its own characteristic, and there might be new BB resistance genes and molecular mechanism in O. officinalis. First, there were no remarkable differences in the leaf microstructures among the four O. officinalis populations, which indicated that the difference in BB resistance of the four populations was caused by the resistance genes. Second, the four populations exhibited various BB resistance, among which the Gengma_Lingcang population had the strongest resistance. Four populations possessed dominant Xa5, dominant Xa13, and recessive xa3/xa26 homologous alleles of xa5, xa13, and Xa3/Xa26 resistance genes. Third, the expression levels of Xa5, Xa13, and xa3/xa26 were different in the four populations under Xoo stress. OoXa13 could enhance the resistance to PXO99 and C5 by down-regulating its own expression level, while the contribution of Ooxa3/xa26 on BB resistance might function like OoXa13. This study provided a solid foundation and reference for studying the molecular mechanism of BB resistance in O. officinalis.
References
Bhasin H, Bhatia D, Raghuvanshi S, Lore JS, Sahi GK, Kaur B, Vikal Y, Singh K (2012) New PCR-based sequence-tagged site marker for bacterial blight resistance gene Xa38 of rice. Mol Breed 30:607–611
Bustamante M, Jin J, Casagran O, Nolan T, Riechmann JL (2014) Gene expression analysis by quantitative real-time PCR for floral tissues. Methods Mol Biol 1110:363
Cao Y, Duan L, Li H, Sun X, Zhao Y, Xu C, Li X, Wang S (2007) Functional analysis of Xa3/Xa26 family members in rice resistance to Xanthomona oryzae pv. oryzae. Theor Appl Genet 115:887–895
Cheng Z, Huang X, Qian J, Zhang Y, Wu C, Wang D, Tang Z, Wang L, Zhou Y (2004) Characteristics and discovery of the newly found natural live material of Oryza officinalis at edge of extinction in Yunnan. Acta Bot Yunnanica 26:267–274
Chu Z, Ouyang Y, Zhang J, Yang H, Wang S (2004) Genome-wide analysis of defense responsive genes in bacterial blight resistance of rice mediated by the recessive R gene xa13. Mol Gen Genomics 271:111–120
Chu Z, Fu B, Yang H, Xu C, Li Z, Sanchez A, Park YJ, Bennetzen JL, Zhang Q, Wang S (2006a) Targeting xa13, a recessive gene for bacterial blight resistance in rice. Theor Appl Genet 112:455–461
Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL, Zhang Q, Wang S (2006b) Promoter mutations of an essential gene for pollen development result in disease resistance in rice. Genes Dev 20:1250–1255
Fan S (2000) The species geographical distribution of wild rice and their characteristics in China. J Wuhan Bot Res 18:417–425
Fan S, Zhang Z, Liu L, Liu H, Liang C (2000) Conservation of wild rice genetic resources in China and their utilization in breeding. Chin Biodivers 8:198–207
Fan H, Wang L, Zhang L, Yu X, Wang X, Jin Q, Wang J (2011) Breeding of rice lines with bacterial blight resistance gene Xa23 by using marker-assisted selection. Chin J Rice Sci 25:331–334
Gao L, Ge S, Hong D (2000) A preliminary study on ecological differentiation within the common wild rice Oryza rufipogon Griff. Acta Agron Sin 26:210–213
Gu K, Tian D, Yang F, Wu L, Sreekala C, Wang D, Wang G, Yin Z (2004) High-resolution genetic mapping of Xa27(t), a new bacterial blight resistance gene in rice, Oryza sativa L. Theor Appl Genet 108:800–807
Gu K, Sangha JS, Li Y, Yin Z (2008) High resolution genetic mapping of bacterial blight resistance gene Xa10. Theor Appl Genet 116:155–163
Gu K, Tian D, Qiu C, Yin Z (2009) Transcription activator-like ype III effector AvrXa27 depends on OsTFIIAgamma5 for the activation of Xa27 transcription in rice that triggers disease resistance to Xanthomonas oryzae pv. Oryzae. Mol Plant Pathol 10:829–835
Huang S, Antony G, Li T, Liu B, Obasa K, Yang B, White FK (2016) The broadly effective recessive resistance gene xa5 of rice is a virulence effector-dependent quantitative trait for bacterial blight. Plant J 86(2):186–194
Hur YJ, Jeung J, Kim SY, Park H, Cho J, Lee JY, Sohn Y, Song YC, Park D, Lee C, Sohn JG, Nam M, Lee JH (2013) Functional markers for bacterial blight resistance gene Xa3 in rice. Mol Breed 31:981–985
Iyer-Pascuzzi AS, McCouch SR (2004) The rice bacterial blight resistance gene xa5 encodes a novel form of disease resistance. Mol Plant-Microbe Interact 17:1348–1354
Iyer-Pascuzzi AS, Jiang H, Huang L, Mccouch S (2008) Genetic and functional characterization of the rice bacterial blight disease resistance gene xa5. Phytopathology 98:289–295
Jiang G, Xia Z, Zhou Y, Wan J, Li D, Chen R, Zhai W, Zhu L (2006) Testifying the rice bacterial blight resistance gene xa5 by genetic complementation and further analyzing xa5 (Xa5) in comparison with its homolog TFIIA gamma 1. Mol Genet Genomics 275:354–366
Kameswara RK, Lakshminarasu M, Jena KK (2002) DNA markers and marker-assisted breeding for durable resistance to bacterial blight disease in rice. Biotechnol Adv 20:33–47
Kauffman HE, Reddy A, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Rep 57:537–541
Leach J, Roberts P, Guo A, Barton-Willis P (1989) Multiplication of Xanthomonas campestris pv. oryzae in rice leaves. International Rice Research Institute, Manila, pp 43–54
Lee KS, Rasabandith S, Angeles ER, Khush GS (2003) Inheritance of resistance to bacterial blight in 21 cultivars of rice. Phytopathology 93:147–152
Li D, Chen L, Li W, Ke X, Yu T, Li E, Huang X, Cheng Z (2015) Identification of bacterial blight resistance gene in Yunnan wild rice. Acta Agron Sin 41:386–393
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25(4):402–408
Mansfieldi J, Genin S, Magori S, Citovsky V, Sriariyanum M, Ronald P, Dow M, Verdier V, Beer SV, Machado MA, Toth I, Salmond G, Foster GD (2012) Top 10 plant pathogenic bacteria in molecular plant pathology. Mol Plant Pathol 13:614–629
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Nino-Liu DO, Ronald PC, Bogdanove AJ (2006) Xanthomonas oryzae pathovars: model pathogens of a model crop. Mol Plant Pathol 7:303–324
Peng S, Wei Z, Mao C, Huang H, Xiao F (1982) Identification of multi resistance of O. meyeriana, O. officinalis and O. sativa growing in Yunnan province. Acta Phytopathol Sin 17:58–60
Qin Q, Li G, Song F, Xu X (2000) The relation of specific traits of wild rice and super high-yield breeding. Hubei Agric Sci 6:16–18
Rasabandith S (1998) Inheritance of resistance to bacterial leaf blight Xanthomonas oryzae pv oryzae (Xoo) in some rice varieties Oryza sativa L. International Rice Research Institute Repository
Ronald PC, Albano B, Tabien R, Abenes L, Wu K, McCouch S, Tanksley SD (1992) Genetic and physical analysis of the rice bacterial blight disease resistance locus, Xa21. Mol Gen Genet 236:113–120
Sanchez AC, Ilag LL, Yang D, Brar DS, Ausubel F, Khush GS, Yano M, Sasaki T, Li Z, Huang N (1999) Genetic and physical mapping of xa13, a recessive gene bacterial blight resistance gene in rice. Theor Appl Genet 98:1022–1028
Song W, Wang G, Chen L, Kim HS, Holsten T, Wang B, Zhai W, Zhu L, Fauquet C, Ronald PC (1995) The rice disease resistance gene, Xa-21, encodes a receptor kinase-like protein. Science 270:1804–1806
Sugio A, Yang B, Zhu T, White FF (2007) Two type III effector genes of Xanthomonas oryzae pv. oryzae control the induction of the host genes OsTFIIAγ1 and OsTFX1 during bacterial blight of rice. Proc Natl Acad Sci 104:10720–10725
Sun X, Cao Y, Wang S (2006) Point mutations with positive selection were a major force during the evolution of a receptor-kinase resistance gene family of rice. Plant Physiol 140:998–1008
Tan G, Ren X, Weng Q, Shi Z, Zhu L, He G (2004) Mapping of a new resistance gene to bacterial blight in rice line introgressed from O. officinalis. J Genet Genomics 31:724–729
Vikal Y, Bhatia D (2017) Genetics and genomics of bacterial blight resistance in rice. Adv Int Rice Res:175–212
Waheed MA, Ullah I, Ahmad H, Sirajuddin AH, Khan A, Khan AZ (2009) Evaluation of rice genotypes for resistance against bacterial leaf blight. Pak J Bot 41:329–335
Xu L, Li X, Wang S (2008) Analysis on expression patterns of the family members of rice bacterial blight resistance gene Xa3/Xa26. Chin J Rice Sci 22:559–563
Yoshimura S, Yamanouchi U, Katayose Y, Toki S, Wang Z, Yano M, Iwata N, Sasaki T (1998) Expression of Xa1, a bacterial blight-resistance gene in rice, is induced by bacterial inoculation. Proc Natl Acad Sci 95:1663–1668
Yuan M, Chu Z, Li X, Xu C, Wang S (2010) The bacterial pathogen Xanthomonas oryzae overcomes rice defenses by regulating host copper redistribution. Plant Cell 22:3164–3176
Yuan M, Li X, Xiao J, Wang S (2011) Molecular and functional analyses of COPT/Ctr-type copper transporter-like gene family in rice. BMC Plant Biol 11:69
Zhang N (2002) Review of the studies relationship of genus Oryza. Nanyang Technol Univ 1:61–76
Zhang Q, Wang C, Shi A, Bai J, Lin S (1994) Evaluation of wild rice species resistance to rice bacterial blight (Xanthomonas oryzae pv. oryzae). Sci Agric Sin 27:1–9
Zhang Q, Lin S, Zhao B, Wang C, Yang W, Zhou Y, Li D, Chen C, Zhu L (1998) Identification and tagging a new gene for resistance to bacterial blight (Xanthomonas oryzae pv. oryzae) from O. rufipogon. Rice Genet Newsl 15:138
Zhao J, Fu J, Li X, Xu C, Wang S (2009) Dissection of the factors affecting development-controlled and race-specific disease resistance conferred by leucine-rich repeat receptor kinase-type R genes in rice. Theor Appl Genet 119:231–239
Acknowledgments
This work was supported by the Key Project of China Scientific Ministry (2017YFD0100202), grants from the National Natural Science Foundation of China—Yunnan Project (U1302265), (31560384, NSFC project 31560384), and the Key Projects of Applied Basic Research in Yunnan (2016RA002,2017FG001-007,2018FG001-068).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Fig. S1
The comparative analysis of leaf microstructures among four O. officinalis populations. (a) There was no significant difference in the numbers of macro-vascular bundles, small-vascular bundles, and air cavity of leaf midvein among four populations. (b) The area comparison of macro-vascular bundles of leaf midvein among four populations. The different letters above the bars represented significant difference (p < 0.05). (c) There was no significant difference in leaf thickness among four populations. The different letters above the bar represented the leaf thickness of cultivar 02428 which was thicker than that of the four populations (p < 0.05). (1) Gengma, Lincang population; (2) Menghai, Xishuangbanna population; (3) Jingne, Jinghong population; (4) Lancang, Puer population; (5) cultivar 02428. (PNG 263 kb)
Fig. S2
The amino acid alignment of OoXa5 among four O. officinalis populations. (Nipponbare and IR24) with susceptible gene Xa5; (IRBB5) with recessive resistance gene xa5. The 39th amino acid of Xa5 is valine (V), while the 39th amino acid of resistant gene xa5 is glutamic acid. (E) The 39th amino acid of OoXa5 in four populations was valine (V), suggesting OoXa5 belonged to Xa5 homologous allele of xa5 resistance gene. (PNG 265 kb)
Fig. S3
The amino acid alignment of OoXa13 among four O. officinalis populations. (IR64 and IR24) with dominant gene Xa13; (IRBB13) with recessive resistance gene xa13. The 238th amino acid of Xa13 is alanine (A), while the 238th amino acid of xa13 is threonine (T). The OoXa13 of four populations had many different amino acid sites compared with IR64, IR24, and IRBB13; interestingly, OoXa13 simultaneously missed the 237th and 238th amino acids. (PNG 699 kb)
Fig. S4
The comparison of promoters in Xa13 gene between O. officinalis and IR64 cultivated rice. The 280 bp bands from four populations were sequenced and aligned with Xa13 gene from IR64. We found that the alignment similarity of gene sequences was 99.71% between O. officinalis and IR64, which proved four populations carried with the promoter regions of Xa13 gene. (PNG 105 kb)
Fig. S5
The amino acid alignment of Ooxa3/xa26 among four O. officinalis populations. (IR24) with recessive xa3/xa26 gene; (Minghui 63 and IRBB3) with resistance gene Xa3/Xa26; (double underline) signal peptide; (region of solid arrow) leucine-rich repeat (LRR) domain; (single underline) transmembrane domain; (region of hollow arrow) serine-threonine kinase domain; * represented the eight differential amino acid sites between Xa3/Xa26 and xa3/xa26 in the LRR domain. (PNG 4071 kb)
Fig. S6
The nucleotide alignment of Ooxa3/xa26 among four O. officinalis populations. (IR24) with recessive xa3/xa26 gene; (Minghui 63 and IRBB3) with resistance gene Xa3/Xa26. The sequence of 452–456 bp from the start codon (ATG) is AATC in xa3/xa26 gene, while Xa3/Xa26 gene has the TGCA sequence at the same site. Four populations and the reported xa3/xa26 (AEM430421) had the AATC sequence in the correspondent sites (429–432 bp). (PNG 2086 kb)
Fig. S7
The comparison of OoXa3/Xa26 domains between O. officinalis and cultivated rice. There were three more LRR domains in four populations than cultivated rice. (a) A-1, A-2, A-3, and A-4 represented the Xa3/Xa26 domains of Gengma_Lincang, Menghai_Xishuangbanna, Jingne_Jinghong, and Lancang_Puer population, respectively. There was one LRR domain in Gengma_Lincang different from the other materials; (b) the two types of Xa3/Xa26 domain in IR24; (c) the three types of Xa3/Xa26 domain in Minghui63/IRBB3. (PNG 755 kb)
Table S1
The Xoo races for BB resistance identification of Yunnan O. officinalis populations. (XLSX 11 kb)
Table S2
Grading criteria and resistance type of rice bacterial blight. (XLSX 11 kb)
Table S3
The PCR primers for identification of bacterial blight resistance genes from four O. officinalis populations. (XLSX 12 kb)
Table S4
The cloning primers of OoXa5, OoXa13, and Ooxa3/xa26. (XLSX 12 kb)
Table S5
The qRT-PCR primers of OoXa5, OoXa13, and Ooxa3/xa26 genes for expression pattern analysis. (XLSX 11 kb)
Table S6
Resistance data of O. officinalis inoculated with seven Xoo races after 21 d. (XLSX 12 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, C., Xiao, S., Li, D. et al. Identification and Expression Pattern Analysis of Bacterial Blight Resistance Genes in Oryza officinalis Wall ex Watt Under Xanthomonas oryzae Pv. oryzae Stress. Plant Mol Biol Rep 37, 436–449 (2019). https://doi.org/10.1007/s11105-019-01164-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-019-01164-3