Abstract
Key message
UBIQUITIN-SPECIFIC PROTEASES play important roles in plant development and stress responses.
Abstract
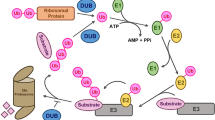
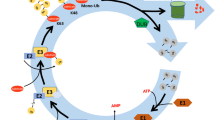
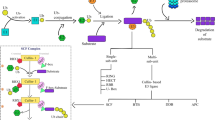
Protein ubiquitination and deubiquitination are reversible processes, which can modulate the stability, activity as well as subcellular localization of the substrate proteins. UBIQUITIN-SPECIFIC PROTEASE (UBP) protein family participates in protein deubiquitination. Members of UBP family are involved in a variety of physiological processes in plants, as evidenced by their functional characterization in model plant Arabidopsis and other plants. UBPs are conserved in plants and distinct UBPs function in different regulatory processes, although functional redundancies exist between some members. Here we briefly reviewed recent advances in understanding the biological functions of UBP protein family in Arabidopsis, particularly the molecular mechanisms by which UBPs regulate plant development and stress responses. We believe that elucidation of UBPs function and regulation in Arabidopsis will provide new insights about protein deubiquitination and might shed light on the understanding of the mechanistic roles of UBPs in general, which will definitely contribute to crop improvement in agriculture.



Similar content being viewed by others
References
Amerik AY, Hochstrasser M (2004) Mechanism and function of deubiquitinating enzymes. BBA 1695:189–207
Aung K, Hu J (2012) Differential roles of Arabidopsis dynamin-related proteins DRP3A, DRP3B, and DRP5B in organelle division. J Integr Plant Biol 54:921–931
Baetz U, Martinoia E (2014) Root exudates: the hidden part of plant defense. Trends Plant Sci 19:90–98
Barberon M, Zelazny E, Robert S et al (2011) Monoubiquitin-dependent endocytosis of the iron-regulated transporter 1 (IRT1) transporter controls iron uptake in plants. Proc Natl Acad Sci USA 108:E450–E458
Bartel B, Citovsky V (2012) Focus on ubiquitin in plant biology. Plant Physiol 160:1
Berger F, Grini PE, Schnittger A (2006) Endosperm: an integrator of seed growth and development. Curr Opin Plant Biol 9:664–670
Bernhardt C, Zhao M, Gonzalez A et al (2005) The bHLH genes GL3 and EGL3 participate in an intercellular regulatory circuit that controls cell patterning in the Arabidopsis root epidermis. Development 132(2):291–298
Bonnet JP, Romier C, Tora L et al (2008) Zinc-finger UBPs: regulators of deubiquitylation. Trend Biochem Sci 33:369–375
Cao Y, Dai Y, Cui S et al (2008) Histone H2B monoubiquitination in the chromatin of FLOWERING LOCUS C regulates flowering time in Arabidopsis. Plant Cell 20:2586–2602
Capron A, Okresz L, Genschik P (2003) First glance at the plant APC/C, a highly conserved ubiquitin-protein ligase. Trends Plant Sci 8:83–89
Chandler J, Mcardle B, Callis J (1997) AtUBP3 and AtUBP4 are two closely related Arabidopsis thaliana ubiquitin-specific proteases present in the nucleus. Mol Genet Genomics 255:302–310
Chen ZJ, Sun LJ (2009) Nonproteolytic functions of ubiquitin in cell signaling. Mol Cell 33:275–286
Cui X, Lu F, Li Y et al (2013) Ubiquitin-specific proteases UBP12 and UBP13 act in circadian clock and photoperiodic flowering regulation in Arabidopsis. Plant Physiol 162:897–906
Cutler SR, Rodriguez PL, Finkelstein RR et al (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679
Dangl J, Jones J (2001) Plant pathogens and integrated defence responses to infection. Nature 411:826–833
Edgar B, Zielke N, Gutierrez C (2014) Endocycles: a recurrent evolutionary innovation for post-mitotic cell growth. Nat Rev Mol Cell Biol 15:197–210
De Gara L, Locato V, Dipierro S et al (2010) Redox homeostasis in plants. The challenge of living with endogenous oxygen production. Respir Physiol Neurobiol 173:S13–S19
Derkacheva M, Liu S, Figueiredo D et al (2016) H2A deubiquitinases UBP12/13 are part of the Arabidopsis Polycomb group protein system. Nat Plant 2:16126
Doelling J, Yan N, Kurepa J et al (2001) The ubiquitin-specific protease UBP14 is essential for early embryo development in Arabidopsis thaliana. Plant J 27:393–405
Doelling J, Phillips A, Soyler-Ogretim G et al (2007) The ubiquitin-specific protease subfamily UBP3/UBP4 is essential for pollen development and transmission in Arabidopsis. Plant Physiol 145:801–813
Dreher K, Callis J (2007) Ubiquitin, hormones and biotic stress in plants. Annu Bot 99:787–822
Du L, Li N, Chen L et al (2014) The ubiquitin receptor DA1 regulates seed and organ size by modulating the stability of the ubiquitin-specific protease UBP15/SOD2 in Arabidopsis. Plant Cell 26:665–677
Ewan R, Pangestuti R, Thornber S et al (2011) Deubiquitinating enzymes AtUBP12 and AtUBP13 and their tobacco homologue NtUBP12 are negative regulators of plant immunity. New Phytol 191:92–106
Filimonenko M, Stuffers S, Raiborg C, Yamamoto A et al (2007) Functional multivesicular bodies are required for autophagic clearance of protein aggregates associated with neurodegenerative disease. J Cell Biol 179:485–500
Fischer J (2003) Deubiquitinating enzymes: their roles in development, differentiation, and disease. Int Rev Cytol 229:43–72
Friedman JR, Nunnari J (2014) Mitochondrial form and function. Nature 505:335–343
Gegas VC, Nazari A, Griffiths S et al (2010) A genetic framework for grain size and shape variation in wheat. Plant Cell 22:1046–1056
Gilroy S, Jones DL (2000) Through form to function: root hair development and nutrient uptake. Trends Plant Sci 5:56–60
Glickman M, Ciechanover A (2002) The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev 82:373–428
Gonzalez N, De Bodt S, Sulpice R et al (2010) Increased leaf size: different means to an end. Plant Physiol 153:1261–1279
Haglund K, Dikic I (2005) Ubiquitylation and cell signaling. EMBO J 24:3353–3359
Harmer SL (2009) The circadian system in higher plants. Annu Rev Plant Biol 60:357–377
Hase Y, Trung K, Matsunaga T et al (2006) A mutation in the uvi4 gene promotes progression of endo-reduplication and confers increased tolerance towards ultraviolet B light. Plant J 46:317–326
Hasegawa P (2013) Sodium (Na+) homeostasis and salt tolerance of plants. Environ Exp Bot 92:19–31
Heath M (2000) Hypersensitive response-related death. Plant Mol Biol 44:321–334
Hepler P, Vidali L, Cheung A (2001) Polarized cell growth in higher plants. Annu Rev Cell Dev Biol 17:159–187
Hepler PK, Kunkel JG, Rounds CM et al (2012) Calcium entry into pollen tubes. Trends Plant Sci 17:32–38
Hershko A, Ciechanover A (1998) The ubiquitin system. Annu Rev Biochem 67:425–479
Heyman J, De Veylder L (2012) The anaphase-promoting complex/cyclosome in control of plant development. Mol Plant 5:1182–1194
Higashiyama T, Takeuchi H (2015) The mechanism and key molecules involved in pollen tube guidance. Annu Rev Plant Biol 66:393–413
Hochstrasser M (1996) Ubiquitin-dependent protein degradation. Annu Rev Genet 30:405–439
Hu M, Li P, Li M et al (2002) Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111:1041–1054
Huang OW, Cochran AG (2013) Regulation of deubiquitinase proteolytic activity. Curr Opin Struct Biol 23:806–811
Ishida T, Kurata T, Okada K et al (2008) A genetic regulatory network in the development of trichomes and root hairs. Annu Rev Plant Biol 59(1):365–386
Isono E, Nagel M (2014) Deubiquitylating enzymes and their emerging role in plant biology. Front Plant Sci 5:56
Jamai A, Salome P, Schilling S et al (2009) Arabidopsis photorespiratory serine hydroxymethyltransferase activity requires the mitochondrial accumulation of ferredoxin-dependent glutamate synthase. Plant Cell 21:595–606
Jung C, Muller AE (2009) Flowering time control and applications in plant breeding. Trends Plant Sci 14:563–573
Kanaoka MM, Higashiyama T (2015) Peptide signaling in pollen tube guidance. Curr Opin Plant Biol 28:127–136
Katsiarimpa A, Kalinowska K, Anzenberger F et al (2013) The deubiquitinating enzyme AMSH1 and the ESCRT-III subunit VPS2.1 are required for autophagic degradation in Arabidopsis. Plant Cell 25:2236–2252
Kerscher O, Felberbaum R, Hochstrasser M (2006) Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu Rev Cell Dev Biol 22:159–180
Kwak S, Schiefelbein J (2007) The role of the SCRAMBLED receptor-like kinase in patterning the Arabidopsis root epidermis. Dev Biol 302:118–131
Lafonplacette C, Kohler C (2014) Embryo and endosperm, partners in seed development. Curr Opin Plant Biol 17:64–69
Larson-Rabin Z, Li Z, Masson P et al (2009) FZR2/CCS52A1 expression is a determinant of endoreduplication and cell expansion in Arabidopsis. Plant Physiol 149:874–884
Lee HK, Cho SK, Son O, Xu Z, Hwang I et al (2009) Drought stress-induced Rma1H1, a RING membrane-anchor E3 ubiquitin ligase homolog, regulates aquaporin levels via ubiquitination in transgenic Arabidopsis plants. Plant Cell 21:622–641
Leishman M, Westoby M (1994) The role of seed size in seedling establishment in dry soil-conditions: experimental-evidence from semiarid species. J Ecol 82:249–258
Leitner J, Petrasek J, Tomanov K et al (2012) Lysine63-linked ubiquitylation of PIN2 auxin carrier protein governs hormonally controlled adaptation of Arabidopsis root growth. Proc Natl Acad Sci USA 109:8322–8327
Li Y, Zheng L, Corke F et al (2008) Control of final seed and organ size by the DA1 gene family in Arabidopsis thaliana. Genes Dev 22:1331–1336
Li WF, Perry PJ, Prafulla NN et al (2010) Ubiquitin-specific protease 14 (UBP14) is involved in root responses to phosphate deficiency in Arabidopsis. Mol Plant 3:212–223
Lim K, Lim GG (2011) K63-linked ubiquitination and neurodegeneration. Neurobiol Dis 43:9–16
Linkies A, Graeber K, Knight CA et al (2010) The evolution of seeds. New Phytol 186:817–831
Liu Y, Wang F, Zhang H et al (2008) Functional characterization of the Arabidopsis ubiquitin-specific protease gene family reveals specific role and redundancy of individual members in development. Plant J 55:844–856
Liu L, Cui F, Li Q, Yin B et al (2011) The endoplasmic reticulum-associated degradation is necessary for plant salt tolerance. Cell Res 21:957–969
Lopez-Bucio J, Cruz-Ramirez A, Herrera-Estrella L (2003) The role of nutrient availability in regulating root architecture. Curr Opin Plant Biol 6:280–287
Lu D, Lin W, Gao X et al (2011) Direct ubiquitination of pattern recognition receptor FLS2 attenuates plant innate immunity. Science 332:1439–1442
Luisa I, Iniesto E, Rodriguez L et al (2014) Targeted degradation of abscisic acid receptors is mediated by the ubiquitin ligase substrate adaptor DDA1 in Arabidopsis. Plant Cell 26:712–728
Luo M, Luo M, Buzas D et al (2008) UBIQUITIN-SPECIFIC PROTEASE26 is required for seed development and the repression of PHERES1 in Arabidopsis. Genetics 180:229–236
Lyzenga WJ, Stone SL (2012) Abiotic stress tolerance mediated by protein ubiquitination. J Exp Bot 63:599–616
Ma Y, Szostkiewicz I, Korte A et al (2009) Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324:1064–1068
Makarevich G, Leroy O, Akinci U et al (2006) Different Polycomb group complexes regulate common target genes in Arabidopsis. EMBO Rep 7:947–952
Marino D, Peeters N, Rivas S (2012) Ubiquitination during plant immune signaling. Plant Physiol 160:15–27
McClung C, Hsu M, Painter J et al (2000) Integrated temporal regulation of the photorespiratory pathway. Circadian regulation of two Arabidopsis genes encoding serine hydroxymethyltransferase. Plant Physio 123:381–392
Mcgouran JF, Gaertner SR, Altun M, Kramer HB et al (2013) Deubiquitinating enzyme specificity for ubiquitin chain topology profiled by di-ubiquitin activity probes. Chem Biol 20:1447–1455
Meinke DW (2003) Molecular genetics of plant embryogenesis. Annu Rev Plant Biol 46:369–394
Melaragno JE, Mehrotra BL, Coleman AW (1993) Relationship between endopolyploidy and cell size in epidermal tissue of Arabidopsis. Plant Cell 5:1661–1668
Michaels S, Amasino R (1999) FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11:949–956
Miyakawa T, Fujita Y, Yamaguchishinozaki K et al (2013) Structure and function of abscisic acid receptors. Trends Plant Sci 18:259–266
Moon J, Parry G, Estelle M (2004) The ubiquitin-proteasome pathway and plant development. Plant Cell 16:3181–3195
Moon YK, Hong J, Cho Y et al (2009) Structure and expression of OsUBP6, an ubiquitin-specific protease 6 homolog in rice (Oryza sativa L.). Mol Cell 28:463–472
Moreno J, Martín R, Castresana C (2005) Arabidopsis SHMT1, a serine hydroxymethyltransferase that functions in the photorespiratory pathway influences resistance to biotic and abiotic stress. Plant J 41:451–463
Nijman SM, Lunavargas MP, Velds A et al (2005) A genomic and functional inventory of deubiquitinating enzymes. Cell 123:773–786
Nowack MK, Harashima H, Dissmeyer N et al (2012) Genetic framework of cyclin-dependent kinase function in Arabidopsis. Dev Cell 22:1030–1040
Pan R, Kaur N, Hu J (2014) The Arabidopsis mitochondrial membrane-bound ubiquitin protease UBP27 contributes to mitochondrial morphogenesis. Plant J 78:1047–1059
Park SY, Fung P, Nishimura N et al (2009) Abscisic acid inhibits type 2 C protein phosphatases via the PYR/PYL family of START proteins. Science 324:1068–1071
Paul MH (2003) Plant cellular and molecular responses to high salinity. Annu Rev Plant Biol 51:463–499
Peret B, Clement M, Nussaume L et al (2011) Root developmental adaptation to phosphate starvation: better safe than sorry. Trends Plant Sci 16:442–450
Pickart CM (2003) Mechanisms underlying ubiquitination. Annu Rev Biochem 70:503–533
Pickart C, Eddins M (2004) Ubiquitin: structures, functions, mechanisms. BBA Mol Cell Res 1695:55–72
Pien S, Fleury D, Mylne JS et al (2008) ARABIDOPSIS TRITHORAX1 dynamically regulates FLOWERING LOCUS C activation via histone 3 lysine 4 trimethylation. Plant Cell 20:580–588
Piper RC, Dikic I, Lukacs GL (2014) Ubiquitin-dependent sorting in endocytosis. CSH Perspect Biol 6:a016808
Qiu Q, Guo Y, Dietrich M et al (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci USA 99:8436–8441
Qu X, Jiang Y, Chang M et al (2015) Organization and regulation of the actin cytoskeleton in the pollen tube. Front Plant Sci 5:786
Quan R, Lin H, Mendoza I et al (2007) SCABP8/CBL10, a putative calcium sensor, interacts with the protein kinase SOS2 to protect Arabidopsis shoots from salt stress. Plant Cell 19:1415–1431
Racioppi J, Dahlman D, Neukranz R (1981) Effects of L-Canavanine on arginine catabolism in Manduca-sexta (Sphingidae,Lepidoptera). Comp Biochem Physiol B 70:639–642
Raghavendra AS, Gonugunta VK, Christmann A et al (2010) ABA perception and signalling. Trends Plant Sci 15:395–401
Raonaik C, Chandler J, Mcardle B et al (2000) Ubiquitin-specific proteases from Arabidopsis thaliana: cloning of AtUBP5 and analysis of substrate specificity of AtUBP3, AtUBP4, and AtUBP5 using Escherichia coli in vivo and in vitro assays. Arch Biochem Biophys 379:198–208
Reyesturcu FE, Ventii KH, Wilkinson KD (2009) Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu Rev Biochem 78:363–397
Rosenthal G (1991) The biochemical basis for the deleterious effects of L-canavanine. Phytochemistry 30:1055–1058
Rubio V, Linhares FR, Solano R et al (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Gene Dev 15:2122–2133
Rusten TE, Vaccari T, Lindmo K, Rodahl LM et al (2007) ESCRTs and Fab1 regulate distinct steps of autophagy. Curr Biol 17:1817–1825
Salazarhenao JE, Velezbermudez IC, Schmidt W (2016) The regulation and plasticity of root hair patterning and morphogenesis. Development 143:1848–1858
Sanchezcalderon L, Lopezbucio J, Chaconlopez A et al (2005) Phosphate starvation induces a determinate developmental program in the roots of Arabidopsis thaliana. Plant Cell Physiol 46:174–184
Schachtman DP, Shin R (2007) Nutrient sensing and signaling: NPKS. Annu Rev Plant Biol 58:47–69
Schiefelbein J, Kwak S, Wieckowski Y et al (2009) The gene regulatory network for root epidermal cell-type pattern formation in Arabidopsis. J Exp Bot 60:1515–1521
Schmitz RJ, Tamada Y, Doyle MR et al (2009) Histone H2B deubiquitination is required for transcriptional activation of FLOWERING LOCUS C and for proper control of flowering in Arabidopsis. Plant Physiol 149:1196–1204
Searle N (2003) Physiology of flowering. Annu Rev Plant Biol 16:97–118
Sheldon CC, Burn JE, Perez P et al (1999) The FLF MADS box gene: a repressor of flowering in Arabidopsis regulated by vernalization and methylation. Plant Cell 11:445–458
Shi H, Ishitani M, Kim C et al (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci USA 97:6896–6901
Smalle J, Vierstra RD (2004) The ubiquitin 26 S proteasome proteolytic pathway. Annu Rev Plant Biol 55:555–590
Somerville C, Ogren W (1981) Photorespiration-deficient Mutants of Arabidopsis thaliana lacking mitochondrial serine transhydroxymethylase activity. Plant Physiol 67:666–671
Sridhar V, Kapoor A, Zhang K et al (2007) Control of DNA methylation and heterochromatic silencing by histone H2B deubiquitination. Nature 447:735–U18
Steinhorst L, Kudla J (2013) Calcium: a central regulator of pollen germination and tube growth. BBA 1833:1573–1581
Suarezlopez P, Wheatley K, Robson F et al (2001) CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 410:1116–1120
Sullivan ML, Callis J, Vierstra RD (1990) High performance liquid chromatography resolution of ubiquitin pathway enzymes from wheat germ. Plant Physiol 94:710–716
Taylor LP, Hepler PK (2003) Pollengermination and tube growth. Annu Rev Plant Biol 48:461–491
Thomma B, Nurnberger T, Joosten M (2011) Of PAMPs and effectors: the blurred PTI-ETI dichotomy. Plant Cell 23:4–15
Turck F, Fornara F, Coupland G (2008) Regulation and identity of florigen: FLOWERING LOCUS T moves center stage. Annu Rev Plant Biol 59:573–594
Ulrich H (2012) Ubiquitin and SUMO in DNA repair at a glance. J Cell Sci 125:249–254
Urade R (2009) The endoplasmic reticulum stress signaling pathways in plants. Biofactors 35:326–331
Verma R, Aravind L, Oania R et al (2002) Role of Rpn11 metalloprotease in deubiquitination and degradation by the 26 S proteasome. Science 298:611–615
Vierstra R (1996) Proteolysis in plants: mechanisms and functions. Plant Mol Biol 32:275–302
Voll L, Jamai A, Renne P et al (2006) The photorespiratory Arabidopsisshm1 mutant is deficient in SHM1. Plant Physiol 140:59–66
Wang F, Liu P, Zhang Q et al (2012) Phosphorylation and ubiquitination of dynamin-related proteins (AtDRP3A/3B) synergically regulate mitochondrial proliferation during mitosis. Plant J 72:43–56
Wilkinson KD (1997) Regulation of ubiquitin-dependent processes by deubiquitinating enzymes. FASEB J 11:1245–1256
Wilkinson KD (2000) Ubiquitination and deubiquitination: targeting of proteins for degradation by the proteasome. Semin Cell Dev Biol 11:141–148
Xia T, Li N, Dumenil J et al (2013) The ubiquitin receptor DA1 interacts with the E3 ubiquitin ligase DA2 to regulate seed and organ size in Arabidopsis. Plant Cell 25:3347–3359
Xu Y, Jin W, Li N et al (2016) UBIQUITIN-SPECIFIC PROTEASE 14 interacts with ULTRAVIOLET-B INSENSITIVE 4 to regulate endoreduplication and cell and organ growth in Arabidopsis. Plant Cell 28:1200–1214
Yan N, Doelling J, Falbel T et al (2000) The ubiquitin-specific protease family from arabidopsis. AtUBP1 and 2 are required for the resistance to the amino acid analog canavanine. Plant Physiol 124:1828–1843
Zelazny E, Barberon M, Curie C, Vert G (2011) Ubiquitination of transporters at the forefront of plant nutrition. Plant Signal Behav 6:1597–1599
Zhang Y (2003) Transcriptional regulation by histone ubiquitination and deubiquitination. Gene Dev 17:2733–2740
Zhang X, Garreton V, Chua N (2005) The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Gene Dev 19:1532–1543
Zhao J, Zhou H, Zhang M et al (2016) Ubiquitin-specific protease24 negatively regulates abscisic acid signalling in Arabidopsis thaliana. Plant Cell Environ 39:427–440
Zhou H, Zhao J, Yang Y et al (2012) UBIQUITIN-SPECIFIC PROTEASE16 modulates salt tolerance in Arabidopsis by regulating Na+/H+antiport activity and serine hydroxymethyltransferase stability. Plant Cell 24:5106–5122
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Zielke N, Querings S, Rottig C et al (2008) The anaphase-promoting complex/cyclosome (APC/C) is required for rereplication control in endoreplication cycles. Genes Dev 22:1690–1703
Zipfel C (2008) Pattern-recognition receptors in plant innate immunity. Curr Opin Immunol 20:10–16
Acknowledgements
We sincerely apologize to those authors for not being able to cite their works in this review due to space limitation. We thank Dr. Yan Guo from China Agricultural University for critical reading of the manuscript and stimulating discussions. This work was supported by the National Natural Science Foundation of China (31600201 to H.Z.).
Author information
Authors and Affiliations
Contributions
All the authors contributed to the discussion. JC illustrated the artwork. HZ wrote the article.
Corresponding author
Additional information
Accession numbers: Sequence data of AtUBPs related to this review can be found in the Arabidopsis Genome Initiative or GenBank/EMBL databases under the following accession numbers: At2g32780 (AtUBP1), At1g04860 (AtUBP2), At4g39910 (AtUBP3), At2g22310 (AtUBP4), At2g40930 (AtUBP5), At1g51710 (AtUBP6), At3g21280 (AtUBP7), At5g22030 (AtUBP8), At4g10570 (AtUBP9), At4g10590 (AtUBP10), At1g32850 (AtUBP11), At5g06600 (AtUBP12), At3g11910 (AtUBP13), At3g20630 (AtUBP14), At1g17110 (AtUBP15), At4g24560 (AtUBP16), At5g65450 (AtUBP17), At4g31670 (AtUBP18), At2g24640 (AtUBP19), At4g17895 (AtUBP20), At5g46740 (AtUBP21), At5g10790 (AtUBP22), At5g57990 (AtUBP23), At4g30890 (AtUBP24), At3g14400 (AtUBP25), At3g49600 (AtUBP26), and At4g39370 (AtUBP27).
Rights and permissions
About this article
Cite this article
Zhou, H., Zhao, J., Cai, J. et al. UBIQUITIN-SPECIFIC PROTEASES function in plant development and stress responses. Plant Mol Biol 94, 565–576 (2017). https://doi.org/10.1007/s11103-017-0633-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-017-0633-5