Abstract
Flowering is a critical event in the life cycle of plants and is regulated by a combination of endogenous controls and environmental cues. In the present work, we provide clear genetic evidence that GASA5, a GASA family gene in Arabidopsis (Arabidopsis thaliana), is involved in controlling flowering time and stem growth. GASA5 expression was present in all tissues of Arabidopsis plants, as detected by RT-PCR, and robust GUS staining was observed in the shoot apex of 8-day-old seedlings and inflorescence meristems during reproductive development. Phenotypic analysis showed that a GASA5 null mutant (gasa5-1) flowered earlier than wild type with a faster stem growth rate under both long-day (LD) and short-day (SD) photoperiods. In contrast, transgenic plants overexpressing GASA5 demonstrated delayed flowering, with a slower stem growth rate compared to wild-type plants. However, neither the GASA5 null mutants nor the GASA5 overexpressing plants revealed obvious differences in flowering time upon treatment with gibberellic acid (GA3), indicating that GASA5 is involved in gibberellin (GA)-promoted flowering. GAI (GA INSENSITIVE), one of the five DELLAs in Arabidopsis, was more highly expressed in GASA5-overexpressing plants, but it was lower in gasa5-1. Further transcript profiling analysis suggested that GASA5 delayed flowering by enhancing FLOWERING LOCUS C (FLC) expression and repressing the expression of key flowering-time genes, FLOWERING LOCUS T (FT) and LEAFY (LFY). Our results suggest that GASA5 is a negative regulator of GA-induced flowering and stem growth.









Similar content being viewed by others
Abbreviations
- CO:
-
CONSTANS
- CRPs:
-
Cysteine-rich peptides
- FLC:
-
FLOWERING LOCUS C
- FT:
-
FLOWERING LOCUS T
- GASA:
-
Gibberellic stimulated in Arabidopsis
- GA:
-
Gibberellin
- GA3 :
-
Gibberellic acid 3
- GAI:
-
GA insensitive
- GFP:
-
Green fluorescent protein
- GUS:
-
β-Glucuronidase
- LD:
-
Long days
- LFY:
-
LEAFY
- PAC:
-
Paclobutrazol
- RGA:
-
Repressor of the ga1-3 mutant
- SD:
-
Short days
- SOC1:
-
SUPPRESSOR OF OVEREXPRESSION OF CO 1
References
Anderson CM, Wagner TA, Perret M, He ZH, He D, Kohorn BD (2001) WAKs: cell wall-associated kinases linking the cytoplasm to the extracellular matrix. Plant Mol Biol 47:197–206. doi:10.1023/A:1010691701578
Ariizumi T, Murase K, Sun TP, Steber CM (2008) Proteolysis-independent downregulation of DELLA repression in Arabidopsis by the gibberellin receptor GIBBERELLIN INSENSITIVE DWARF1. Plant Cell 20:2447–2559. doi:10.1105/tpc.108.058487
Aubert D, Chevillard M, Dorne AM, Arlaud G, Herzog M (1998) Expression patterns of GASA genes in Arabidopsis thaliana: the GASA4 gene is up-regulated by gibberellins in meristematic regions. Plant Mol Biol 36:871–883. doi:10.1023/A:1005938624418
Ben-Nissan G, Weiss D (1996) The petunia homologue of tomato gast1: transcript accumulation coincides with gibberellin-induced corolla cell elongation. Plant Mol Biol 32:1067–1074. doi:10.1007/BF00041390
Ben-Nissan G, Lee JY, Borohov A, Weiss D (2004) GIP, a Petunia hybrida GA-induced cysteine-rich protein: a possible role in shoot elongation and transition to flowering. Plant J 37:229–238
Berrocal-Lobo M, Segura A, Moreno M, Lopez G, Garcia-Olmedo F, Molina A (2002) Snakin-2, an antimicrobial peptide from potato whose gene is locally induced by wounding and responds to pathogen infection. Plant Physiol 128:951–961. doi:10.1104/pp.010685
Blázquez MA, Weigel D (2000) Integration of floral inductive signals in Arabidopsis. Nature 404:889–892. doi:10.1038/35009125
Blázquez MA, Soowal LN, Lee I, Weigel D (1997) LEAFY expression and flower initiation in Arabidopsis. Development 124:3835–3844
Blázquez MA, Green R, Nilsson O, Sussman MR, Weigel D (1998) Gibberellins promote flowering of arabidopsis by activating the LEAFY promoter. Plant Cell 10:791–800
Boss PK, Bastow RM, Mylne JS, Dean C (2004) Multiple pathways in the decision to flower: enabling, promoting, and resetting. Plant Cell 16(Suppl):S18–S31. doi:10.1105/tpc.015958
Cao D, Cheng H, Wu W, Soo HM, Peng J (2006) Gibberellin mobilizes distinct DELLA-dependent transcriptomes to regulate seed germination and floral development in Arabidopsis. Plant Physiol 142:509–525. doi:10.1104/pp.106.082289
Cheng H, Qin L, Lee S, Fu X, Richards DE, Cao D, Luo D, Harberd NP, Peng J (2004) Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 131:1055–1064. doi:10.1242/dev.00992
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. doi:10.1046/j.1365-313x.1998.00343.x
de la Fuente JI, Amaya I, Castillejo C, Sanchez-Sevilla JF, Quesada MA, Botella MA, Valpuesta V (2006) The strawberry gene FaGAST affects plant growth through inhibition of cell elongation. J Exp Bot 57:2401–2411. doi:10.1093/jxb/erj213
Dill A, Sun T (2001) Synergistic derepression of gibberellin signaling by removing RGA and GAI function in Arabidopsis thaliana. Genetics 159:777–785
Eriksson S, Bohlenius H, Moritz T, Nilsson O (2006) GA4 is the active gibberellin in the regulation of LEAFY transcription and Arabidopsis floral initiation. Plant Cell 18:2172–2181. doi:10.1105/tpc.106.042317
Fleck B, Harberd NP (2002) Evidence that the Arabidopsis nuclear gibberellin signalling protein GAI is not destabilised by gibberellin. Plant J 32:935–947. doi:10.1046/j.1365-313X.2002.01478.x
Fu X, Richards DE, Ait-Ali T, Hynes LW, Ougham H, Peng J, Harberd NP (2002) Gibberellin-mediated proteasome-dependent degradation of the barley DELLA protein SLN1 repressor. Plant Cell 14:3191–3200. doi:10.1105/tpc.006197
Furukawa T, Sakaguchi N, Shimada H (2006) Two OsGASR genes, rice GAST homologue genes that are abundant in proliferating tissues, show different expression patterns in developing panicles. Genes Genet Syst 81:171–180. doi:10.1266/ggs.81.171
Gocal GF, Sheldon CC, Gubler F, Moritz T, Bagnall DJ, MacMillan CP, Li SF, Parish RW, Dennis ES, Weigel D, King RW (2001) GAMYB-like genes, flowering, and gibberellin signaling in Arabidopsis. Plant Physiol 127:1682–1693. doi:10.1104/pp.010442
Harberd NP (2003) Botany. Relieving DELLA restraint. Science 299:1853–1854. doi:10.1126/science.1083217
Hartweck LM, Olszewski NE (2006) Rice GIBBERELLIN INSENSITIVE DWARF1 is a gibberellin receptor that illuminates and raises questions about GA signaling. Plant Cell 18:278–282. doi:10.1105/tpc.105.039958
Herzog M, Dorne AM, Grellet F (1995) GASA, a gibberellin-regulated gene family from Arabidopsis thaliana related to the tomato GAST1 gene. Plant Mol Biol 27:743–752. doi:10.1007/BF00020227
Hussain A, Cao D, Cheng H, Wen Z, Peng J (2005) Identification of the conserved serine/threonine residues important for gibberellin-sensitivity of Arabidopsis RGL2 protein. Plant J 44:88–99. doi:10.1111/j.1365-313X.2005.02512.x
Itoh H, Matsuoka M, Steber CM (2003) A role for the ubiquitin-26S-proteasome pathway in gibberellin signaling. Trends Plant Sci 8:492–497. doi:10.1016/j.tplants.2003.08.002
Jacobsen SE, Olszewski NE (1993) Mutations at the SPINDLY locus of Arabidopsis alter gibberellin signal transduction. Plant Cell 5:887–896
Jiang C, Fu X (2007) GA action: turning on de-DELLA repressing signaling. Curr Opin Plant Biol 10:461–465. doi:10.1016/j.pbi.2007.08.011
Kardailsky I, Shukla VK, Ahn JH, Dagenais N, Christensen SK, Nguyen JT, Chory J, Harrison MJ, Weigel D (1999) Activation tagging of the floral inducer FT. Science 286:1962–1965. doi:10.1126/science.286.5446.1962
King KE, Moritz T, Harberd NP (2001) Gibberellins are not required for normal stem growth in Arabidopsis thaliana in the absence of GAI and RGA. Genetics 159:767–776
Kobayashi Y, Kaya H, Goto K, Iwabuchi M, Araki T (1999) A pair of related genes with antagonistic roles in mediating flowering signals. Science 286:1960–1962. doi:10.1126/science.286.5446.1960
Kohorn BD (2000) Plasma membrane—cell wall contacts. Plant Physiol 124:31–38. doi:10.1104/pp.124.1.31
Koornneef M, Hanhart CJ, van der Veen JH (1991) A genetic and physiological analysis of late flowering mutants in Arabidopsis thaliana. Mol Gen Genet 229:57–66. doi:10.1007/BF00264213
Kotilainen M, Helariutta Y, Mehto M, Pollanen E, Albert VA, Elomaa P, Teeri TH (1999) GEG participates in the regulation of cell and organ shape during corolla and carpel development in gerbera hybrida. Plant Cell 11:1093–1104
Lee H, Suh SS, Park E, Cho E, Ahn JH, Kim SG, Lee JS, Kwon YM, Lee I (2000) The AGAMOUS-LIKE 20 MADS domain protein integrates floral inductive pathways in Arabidopsis. Genes Dev 14:2366–2376. doi:10.1101/gad.813600
Lim MH, Kim J, Kim YS, Chung KS, Seo YH, Lee I, Kim J, Hong CB, Kim HJ, Park CM (2004) A new Arabidopsis gene, FLK, encodes an RNA binding protein with K homology motifs and regulates flowering time via FLOWERING LOCUS C. Plant Cell 16:731–740. doi:10.1105/tpc.019331
Michaels SD, Amasino RM (1999) FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell 11:949–956
Mockler TC, Yu X, Shalitin D, Parikh D, Michael TP, Liou J, Huang J, Smith Z, Alonso JM, Ecker JR, Chory J, Lin C (2004) Regulation of flowering time in Arabidopsis by K homology domain proteins. Proc Natl Acad Sci USA 101:12759–12764. doi:10.1073/pnas.0404552101
Mouradov A, Cremer F, Coupland G (2002) Control of flowering time: interacting pathways as a basis for diversity. Plant Cell 14(Suppl):S111–S130
Oh E, Yamaguchi S, Hu JH, Yusuke J, Jung B, Paik I, Lee HS, Sun TP, Kamiya Y, Choi G (2007) PIL5, a phytochrome-interacting bHLH protein, regulates gibberellin responsiveness by binding directly to the GAI and RGA promoters in Arabidopsis seeds. Plant Cell 19:1192–1208. doi:10.1105/tpc.107.050153
Parcy F (2005) Flowering: a time for integration. Int J Dev Biol 49:585–593. doi:10.1387/ijdb.041930fp
Peng J, Carol P, Richards DE, King KE, Cowling RJ, Murphy GP, Harberd NP (1997) The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev 11:3194–3205. doi:10.1101/gad.11.23.3194
Putterill J, Robson F, Lee K, Simon R, Coupland G (1995) The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell 80:847–857. doi:10.1016/0092-8674(95)90288-0
Reeves PH, Coupland G (2001) Analysis of flowering time control in Arabidopsis by comparison of double and triple mutants. Plant Physiol 126:1085–1091. doi:10.1104/pp.126.3.1085
Rogers SO, Bendich AJ (1985) Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol 5:69–76. doi:10.1007/BF00020088
Rojo E, Sharma VK, Kovaleva V, Raikhel NV, Fletcher JC (2002) CLV3 is localized to the extracellular space, where it activates the Arabidopsis CLAVATA stem cell signaling pathway. Plant Cell 14:969–977. doi:10.1105/tpc.002196
Rosso MG, Li Y, Strizhov N, Reiss B, Dekker K, Weisshaar B (2003) An Arabidopsis thaliana T-DNA mutagenized population (GABI-Kat) for flanking sequence tag-based reverse genetics. Plant Mol Biol 53:247–259. doi:10.1023/B:PLAN.0000009297.37235.4a
Roxrud I, Lid SE, Fletcher JC, Schmidt ED, Opsahl-Sorteberg HG (2007) GASA4, one of the 14-member Arabidopsis GASA family of small polypeptides, regulates flowering and seed development. Plant Cell Physiol 48:471–483. doi:10.1093/pcp/pcm016
Samach A, Onouchi H, Gold SE, Ditta GS, Schwarz-Sommer Z, Yanofsky MF, Coupland G (2000) Distinct roles of CONSTANS target genes in reproductive development of Arabidopsis. Science 288:1613–1616. doi:10.1126/science.288.5471.1613
Sanchez-Fernandez R, Ardiles-Diaz W, Van Montagu M, Inze D, May MJ (1998) Gene note. Cloning of a novel Arabidopsis thaliana RGA-like gene, a putative member of the VHIID-domain transcription factor family. J Exp Bot 49:1609–1610. doi:10.1093/jexbot/49.326.1609
Searle I, He Y, Turck F, Vincent C, Fornara F, Krober S, Amasino RA, Coupland G (2006) The transcription factor FLC confers a flowering response to vernalization by repressing meristem competence and systemic signaling in Arabidopsis. Genes Dev 20:898–912. doi:10.1101/gad.373506
Segura A, Moreno M, Madueno F, Molina A, Garcia-Olmedo F (1999) Snakin-1, a peptide from potato that is active against plant pathogens. Mol Plant Microbe Interact 12:16–23. doi:10.1094/MPMI.1999.12.1.16
Shi L, Gast RT, Gopalraj M, Olszewski NE (1992) Characterization of a shoot-specific, GA3- and ABA-regulated gene from tomato. Plant J 2:153–159
Silverstein KA, Moskal WA Jr, Wu HC, Underwood BA, Graham MA, Town CD, VandenBosch KA (2007) Small cysteine-rich peptides resembling antimicrobial peptides have been under-predicted in plants. Plant J 51:262–280. doi:10.1111/j.1365-313X.2007.03136.x
Silverstone AL, Mak PY, Martinez EC, Sun TP (1997) The new RGA locus encodes a negative regulator of gibberellin response in Arabidopsis thaliana. Genetics 146:1087–1099
Silverstone AL, Ciampaglio CN, Sun T (1998) The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway. Plant Cell 10:155–169
Silverstone AL, Jung HS, Dill A, Kawaide H, Kamiya Y, Sun TP (2001) Repressing a repressor: gibberellin-induced rapid reduction of the RGA protein in Arabidopsis. Plant Cell 13:1555–1566
Thomas SG, Sun TP (2004) Update on gibberellin signaling. A tale of the tall and the short. Plant Physiol 135:668–676. doi:10.1104/pp.104.040279
Thomas SG, Phillips AL, Hedden P (1999) Molecular cloning and functional expression of gibberellin 2-oxidases, multifunctional enzymes involved in gibberellin deactivation. Proc Natl Acad Sci USA 96:4698–4703. doi:10.1073/pnas.96.8.4698
Tyler L, Thomas SG, Hu J, Dill A, Alonso JM, Ecker JR, Sun TP (2004) DELLA proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol 135:1008–1019. doi:10.1104/pp.104.039578
Ueguchi-Tanaka M, Ashikari M, Nakajima M, Itoh H, Katoh E, Kobayashi M, Chow TY, Hsing YI, Kitano H, Yamaguchi I, Matsuoka M (2005) GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 437:693–698. doi:10.1038/nature04028
Ueguchi-Tanaka M, Hirano K, Hasegawa Y, Kitano H, Matsuoka M (2008) Release of the repressive activity of rice DELLA protein SLR1 by gibberellin does not require SLR1 degradation in the gid2 mutant. Plant Cell 20:2437–2446. doi:10.1105/tpc.108.061648
Wagner TA, Kohorn BD (2001) Wall-associated kinases are expressed throughout plant development and are required for cell expansion. Plant Cell 13:303–318
Wigoda N, Ben-Nissan G, Granot D, Schwartz A, Weiss D (2006) The gibberellin-induced, cysteine-rich protein GIP2 from Petunia hybrida exhibits in planta antioxidant activity. Plant J 48:796–805. doi:10.1111/j.1365-313X.2006.02917.x
Xu YL, Gage DA, Zeevaart JA (1997) Gibberellins and stem growth in Arabidopsis thaliana. Effects of photoperiod on expression of the GA4 and GA5 loci. Plant Physiol 114:1471–1476. doi:10.1104/pp.114.4.1471
Yu H, Ito T, Zhao Y, Peng J, Kumar P, Meyerowitz EM (2004) Floral homeotic genes are targets of gibberellin signaling in flower development. Proc Natl Acad Sci USA 101:7827–7832. doi:10.1073/pnas.0402377101
Zentella R, Zhang Z-L, Park M, Thomas SG, Endo A, Murase K, Fleet CM, Jikumaru Y, Nambara E, Kamiya Y, Sun T-P (2007) Global analysis of DELLA direct targets in early gibberellin signaling in Arabidopsis. Plant Cell 19:3037–3057. doi:10.1105/tpc.107.054999
Acknowledgements
We thank Professor Fu Xiang-dong at the Institute of Genetics and Developmental Biology, Chinese Academy of Sciences for kindly providing the rga-24 and gai-t6 seeds and Professor Huang Tao at Xiamen University in China for the ft-10 seeds. This research was supported by grants from the Nature Science Foundation of China (No. 30570165, U0731006) and the National Key Technology R & D Program in China (No. 2007BAD59B06).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below are the electronic supplementary materials.
Fig. S1
GASA5 protein structure and the relationship between GASA5 protein and other GASA homologues. (A) Structure of the GASA5 protein. Gray box represents the GASA domain, and white box represents the cleavable signal peptide (1-27 residues). Prediction of signal peptide in GASA5 protein used the web software PSORT (http://psort.nibb.ac.jp/form.html). (B) Multiple sequence alignment of the GASA5 with several GASA family proteins. Identical residues were labeled with asterisk at the bottom of the sequence. (C) Unrooted phylogenetic tree shows the relationship between the GASA5 protein and other GASA homologues. GASA protein sequences and accession numbers were obtained from public databases (http://www.ncbi.nlm.nih.gov/, http://www.arabidopsis.org). Multiple sequence alignments and phylogenetic tree were constructed by ClustalW (V.1.83) using neighbour-joining (TIFF 433 kb)
Fig. S2
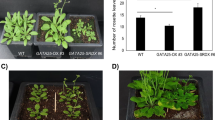
Phenotypes of GASA5-overexpressing plants compared with transgenic plants lacking GASA5 (TIFF 667 kb)
Fig. S3
Phenotypes of gasa5-1 plants after vernalization for three weeks under LD (TIFF 651 kb)
Fig. S4
Phenotypes of GASA5 mutants after being sprayed with 100 µM GA3 under LD (TIFF 757 kb)
Fig. S5
The expression levels of gibberellin 20-oxidase 1 gene (GA20OX1), gibberellin 3-oxidase 1 gene (GA3OX1) and gibberellin 2-oxidase gene 1 (GA2OX1) were detected by RT-PCR. The RNA used was extracted from the inflorescence stem harvested at eight days after bolting under LD (TIFF 241 kb)
Rights and permissions
About this article
Cite this article
Zhang, S., Yang, C., Peng, J. et al. GASA5, a regulator of flowering time and stem growth in Arabidopsis thaliana . Plant Mol Biol 69, 745–759 (2009). https://doi.org/10.1007/s11103-009-9452-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-009-9452-7