Abstract
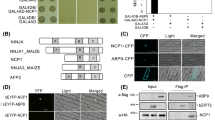
The transcription factor ABA-Insensitive5 (ABI5) is a key regulator of ABA signaling and stress response in Arabidopsis seeds and seedlings. Potential ABI5-interacting proteins were identified by a yeast two-hybrid screen; the most prevalent interactors were a family of four highly conserved plant-specific proteins with no domains of known function, but homology to a previously characterized ABI Five Binding Protein (AFP). This study compares expression and function of the family members. The AFPs are induced by ABA and/or dehydrating stresses in young seedlings, but the developmental timing of their induction differs. Mutations in AFP1 or AFP2 result in increased sensitivity to ABA and salt, whereas afp4 mutants are mildly ABA-resistant. AFP2, like AFP1, acts epistatically to ABI5. Reduced germination or seedling growth of the mutants under stress correlates with a higher level of ABI5 protein when compared to wild-type seedlings, but it is not clear whether this is a cause or effect of the reduced growth. Although both ABI5 and the AFPs are ABA-induced, the ABI5:AFP ratio increases at high ABA concentrations, maintaining growth inhibition under severe stress. An AFP2:GFP fusion, which complements the afp2 mutation, is nuclear-localized in seedlings exposed to stress, but becomes delocalized before being degraded following removal of stress. The AFPs may also interact to varying extents with many ABI5-related bZIP transcription factors. This study suggests that germination and seedling growth are regulated by antagonistic interactions among at least two functionally redundant families, the AFPs and the ABI5-related proteins, providing a mechanism to fine-tune seedling stress responses.











Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- ABI:
-
ABA-insensitive
- AFP:
-
ABI Five Binding Protein
- ABF:
-
ABRE binding factor
- ABRE:
-
ABA-responsive element
- AD:
-
GAL4 activation domain
- AtEm :
-
Arabidopsis early methionine-labelled
- AREB:
-
ABA response element binding factor
- AtDPBF:
-
Arabidopsis thaliana Dc3 promoter binding factor
- BD:
-
GAL4 DNA-binding domain
- bZIP:
-
Basic leucine zipper
- EEL:
-
Enhanced em level
- GM:
-
Germination media
- GUS:
-
Beta-glucuronidase
- LEA:
-
Late embryogenesis abundant
- Min:
-
Minimal media
- RAB:
-
Responsive to ABA
- TMAC2:
-
Two or more ABREs-containing gene 2
References
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Aguilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Arenas-Huertero F, Arroyo A, Zhou L, Sheen J, León P (2000) Analysis of Arabidopsis glucose insensitive mutants, gin5 and gin6, reveals a central role of the plant hormone ABA in the regulation of plant vegetative development by sugar. Genes Dev 14:2085–2096
Bensmihen S, Rippa S, Lambert G, Jublot D, Pautot V, Granier F, Giraudat J, Parcy F (2002) The homologous ABI5 and EEL transcription factors function antagonistically to fine-tune gene expression during late embryogenesis. Plant Cell 14:1391–1403
Bensmihen S, Giraudat J, Parcy F (2005) Characterization of three homologous basic leucine zipper transcription factors (bZIP) of the ABI5 family during Arabidopsis thaliana embryo maturation. J Exp Bot 56:597–603
Brocard I, Lynch T, Finkelstein R (2002) Regulation and role of the Arabidopsis ABA-insensitive (ABI)5 gene in ABA, sugar and stress response. Plant Physiol 129:1533–1543
Carles C, Bies-Ethève N, Aspart L, Leon-Kloosterziel KM, Koornneef M, Echeverria M, Delseny M (2002) Regulation of Arabidopsis thaliana Em genes: role of ABI5. Plant J 30:373–383
Chae MJ, Lee JS, Nam MH, Cho K, Hong JY, Yi SA, Suh SC, Yoon IS (2007) A rice dehydration-inducible SNF1-related protein kinase 2 phosphorylates an abscisic acid responsive element-binding factor and associates with ABA signaling. Plant Mol Biol 63:151–169
Choi H, Hong J, Ha J, Kang J, Kim S (2000) ABFs, a family of ABA-responsive element binding factors. J Biol Chem 275:1723–1730
Choi HI, Park HJ, Park JH, Kim S, Im MY, Seo HH, Kim YW, Hwang I, Kim SY (2005) Arabidopsis calcium-dependent protein kinase AtCPK32 interacts with ABF4, a transcriptional regulator of abscisic acid-responsive gene expression, and modulates its activity. Plant Physiol 139:1750–1761
Clough S, Bent A (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Cutler S, Ehrhardt D, Griffitts J, Somerville C (2000) Random GFP::cDNA fusions enable visualization of subcellular structures in cells of Arabidopsis at a high frequency. Proc Natl Acad Sci USA 97:3718–3723
De Smet I, Signora L, Beeckman T, Inze D, Foyer C, Zhang H (2003) An abscisic acid-sensitive checkpoint in lateral root development in Arabidopsis. Plant J 33:543–555
Finkelstein RR (1994) Mutations at two new Arabidopsis ABA response loci are similar to the abi3 mutations. Plant J 5:765–771
Finkelstein R, Lynch T (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12:599–609
Finkelstein R, Gampala S, Rock C (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14:S15–S45
Finkelstein R, Gampala SSL, Lynch TJ, Thomas TL, Rock CD (2005) Redundant and distinct functions of the ABA response loci ABA-INSENSITIVE(ABI)5 and ABRE-BINDING FACTOR (ABF)3. Plant Mol Biol 59:253–267
Fujii H, Verslues PE, Zhu JK (2007) Identification of two protein kinases required for abscisic acid regulation of seed germination, root growth, and gene expression in Arabidopsis. Plant Cell 19:485–494
Furihata T, Maruyama K, Fujita Y, Umezawa T, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2006) Abscisic acid-dependent multisite phosphorylation regulates the activity of a transcription activator AREB1. Proc Natl Acad Sci USA 103:1988–1993
Haughn G, Somerville C (1986) Sulfonylurea-resistant mutants of Arabidopsis thaliana. Mol Gen Genet 204:430–434
Hirayama T, Shinozaki K (2007) Perception and transduction of abscisic acid signals: keys to the function of the versatile plant hormone ABA. Trends Plant Sci 12:343–351
Huang MD, Wu WL (2007) Overexpression of TMAC2, a novel negative regulator of abscisic acid and salinity responses, has pleiotropic effects in Arabidopsis thaliana. Plant Mol Biol 63:557–569
Jakoby M, Weisshaar B, Droege-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7:106–111
Jefferson R, Kavanagh T, Bevan M (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jiao Y, Ma L, Strickland E, Deng XW (2005) Conservation and divergence of light-regulated genome expression patterns during seedling development in Rice and Arabidopsis. Plant Cell 17:3239–3256
Johnson RR, Wagner RL, Verhey SD, Walker-Simmons MK (2002) The abscisic acid-responsive kinase PKABA1 interacts with a seed-specific abscisic acid response element-binding factor, TaABF, and phosphorylates TaABF peptide sequences. Plant Physiol 130:837–846
Kang J-Y, Choi H-I, Im M-Y, Kim SY (2002) Arabidopsis basic leucine zipper proteins that mediate stress-responsive abscisic acid signaling. Plant Cell 14:343–357
Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50:347–363
Kim J, Harter K, Theologis A (1997) Protein–protein interactions among the Aux/IAA proteins. Proc Natl Acad Sci USA 94:11786–11791
Kim S, Ma J, Perret P, Li Z, Thomas T (2002) Arabidopsis ABI5 subfamily members have distinct DNA binding and transcriptional activities. Plant Physiol 130:688–697
Kim S, Kang J-Y, Cho D-I, Park JH, Kim SY (2004) ABF2, an ABRE-binding bZIP factor, is an essential component of glucose signaling and its overexpression affects multiple stress tolerance. Plant J 40:75–87
Kobayashi Y, Murata M, Minami H, Yamamoto S, Kagaya Y, Hobo T, Yamamoto A, Hattori T (2005) Abscisic acid-activated SNRK2 protein kinases function in the gene-regulation pathway of ABA signal transduction by phosphorylating ABA response element-binding factors. Plant J 44:939–949
Koornneef M, Jorna M, Brinkhorst-van der Swan D, Karssen C (1982) The isolation of abscisic acid (ABA)-deficient mutants by selection of induced revertants in non-germinating gibberellin sensitive lines of Arabidopsis thaliana (L.) Heynh. Theor Appl Genet 61:385–393
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Liu X, Baird W (2004) Identification of a novel gene, HaABRC, from Helianthus annus (Asteraceae) that is upregulated in response to drought, salinity, and abscisic acid. Am J Bot 91:184–191
Lopez-Molina L, Chua N-H (2000) A null mutation in a bZIP factor confers ABA-insensitivity in Arabidopsis thaliana. Plant Cell Physiol 41:541–547
Lopez-Molina L, Mongrand S, Chua N-H (2001) A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc Natl Acad Sci USA 98:4782–4787
Lopez-Molina L, Mongrand S, McLachlin DT, Chait BT, Chua N-H (2002) ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J 32:317–328
Lopez-Molina L, Mongrand S, Kinoshita N, Chua N-H (2003) AFP is a novel negative regulator of ABA signaling that promotes ABI5 protein degradation. Genes Dev 17:410–418
Lu C, Han M-H, Guevara-Garcia A, Fedoroff N (2002) Mitogen-activated protein kinase signaling in postgermination arrest of development by abscisic acid. Proc Natl Acad Sci USA 99:15812–15817
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Nakamura S, Lynch T, Finkelstein R (2001) Physical interactions between ABA response loci of Arabidopsis. Plant J 26:627–635
Notredame C, Higgins DG, Heringa J (2000) T-coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217
Sambrook J, Fritsch E, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Samson F, Brunaud V, Balzergue S, Dubreucq B, Lepiniec L, Pelletier G, Caboche M, Lecharny A (2002) FLAGdb/FST: a database of mapped flanking insertion sites (FSTs) of Arabidopsis thaliana T-DNA transformants. Nucleic Acids Res 30:94–97
Smalle J, Kurepa J, Yang P, Emborg TJ, Babiychuk E, Kushnir S, Vierstra RD (2003) The pleiotropic role of the 26S proteasome subunit RPN10 in Arabidopsis growth and development supports a substrate-specific function in abscisic acid signaling. Plant Cell 15:965–980
Stone SL, Williams LA, Farmer LM, Vierstra RD, Callis J (2006) KEEP ON GOING, a RING E3 ligase essential for Arabidopsis growth and development, is involved in abscisic acid signaling. Plant Cell 18:3415–3428
Toufighi K, Brady SM, Austin R, Ly E, Provart NJ (2005) The botany array resource: e-Northerns, expression angling, and promoter analyses. Plant J 43:153–163
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97:11632–11637
Von Arnim AG, Deng X-W (1994) Light inactivation of Arabidopsis photomorphogenic repressor COP1 involves a cell-specific regulation of its nucleocytoplasmic partitioning. Cell 79:1035–1045
Weatherwax SC, Williams SA, Tingay S, Tobin EM (1998) The phytochrome response of the Lemna gibba NPR1 gene is mediated primarily through changes in abscisic acid levels. Plant Physiol 116:1299–1305
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Ann Rev Plant Biol 57:781–803
Zhang Y, Yang C, Li Y, Zheng N, Chen H, Zhao Q, Gao T, Guo H, Xie Q (2007) SDIR1 is a RING Finger E3 ligase that positively regulates stress-responsive abscisic acid signaling in Arabidopsis. Plant Cell 19:1912–1929
Zhu JK (2002) Salt and drought stress signal transduction in plants. Ann Rev Plant Biol 53:247–273
Acknowledgements
We thank the ABRC team at Ohio State University for efficient distribution of the T-DNA insertion lines from the SALK SIGnAL collection and the French Génoplante Program for the FLAG lines. We thank L. Lopez-Molina for the anti-AFP antibody. This work was supported by United States Department of Agriculture Grant #2003-35304-13201 to RRF and a fellowship from Sigma Delta Epsilon, Women in Science to MEG.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Garcia, M.E., Lynch, T., Peeters, J. et al. A small plant-specific protein family of ABI five binding proteins (AFPs) regulates stress response in germinating Arabidopsis seeds and seedlings. Plant Mol Biol 67, 643–658 (2008). https://doi.org/10.1007/s11103-008-9344-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-008-9344-2