Abstract
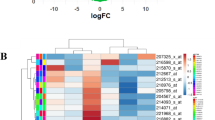
Cross-talk between competitive endogenous RNAs (ceRNAs) may play a critical role in revealing potential mechanisms of tumor development and physiology. Glioblastoma is the most common type of malignant primary brain tumor, and the mechanisms of tumor genesis and development in glioblastoma are unclear. Here, to investigate the role of non-coding RNAs and the ceRNA network in glioblastoma, we performed paired-end RNA sequencing and microarray analyses to obtain the expression profiles of mRNAs, lncRNAs, circRNAs and miRNAs. We identified that the expression of 501 lncRNAs, 1999 mRNAs, 2038 circRNAs and 143 miRNAs were often altered between glioblastoma and matched normal brain tissue. Gene ontology and Kyoto Encyclopedia of Genes and Genomes pathway analyses were performed on these differentially expressed mRNAs and miRNA-mediated target genes of lncRNAs and circRNAs. Furthermore, we used a multi-step computational framework and several bioinformatics methods to construct a ceRNA network combining mRNAs, miRNAs, lncRNAs and circRNA, based on co-expression analysis between the differentially expressed RNAs. We identified that plenty of lncRNAs, CircRNAs and their downstream target genes in the ceRNA network are related to glutamatergic synapse, suggesting that glutamate metabolism is involved in glioma biological functions. Our results will accelerate the understanding of tumorigenesis, cancer progression and even therapeutic targeting in glioblastoma.





Similar content being viewed by others
References
Venkatesh T, Suresh PS, Tsutsumi R (2015) Non-coding RNAs: functions and applications in endocrine-related cancer. Mol Cell Endocrinol 416(1):1–21
Sumazin P et al (2011) An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Can Res 147(2):370–381
Chen Y, Li C, Tan C, Liu X (2016) Circular RNAs: a new frontier in the study of human diseases. J Med Genet 53(6):359–365
Li JH, Liu S, Zhou H, Qu LH, Yang JH (2013) Starbase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale clip-seq data. Nucleic Acids Res 42(D1): 92–97
Ju YP, Lee JE, Park et al (2014) Roles of long non-coding RNAs on tumorigenesis and glioma development. Brain Tumor Res Treat (1):1–6
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) CeRNA hypothesis: the rosetta stone of a hidden RNA language? Cell 146(3):353–358
Riquelme I, Ili C, Roa JC, Brebi P (2016) Long non-coding RNAs in gastric cancer: mechanisms and potential applications. Oncotarget. https://doi.org/10.18632/oncotarget.9396
Tay Y, Rinn J, Pandolfi PP (2014) The multilayered complexity of ceRNA crosstalk and competition. Nature 505(7483):344–352
Thomson DW, Dinger ME (2016) Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet 17(5):272 –283
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J et al (2015) Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res 25(8):981
Shao T, Wu A, Chen J, Chen H, Lu J, Bai J et al (2015) Identification of module biomarkers from the dysregulated ceRNA-ceRNA interaction network in lung adenocarcinoma. Mol Biosyst 11(11):3048–3058
Dou C, Cao Z, Yang B, Ding N, Hou T, Luo F et al (2016) Changing expression profiles of lncRNAs, mRNAs, circRNAs and miRNAs during osteoclastogenesis. Sci Rep 6:21499
Wang PF, Liu N, Song HW, Yao K, Jiang T, Li SW et al (2016) Idh-1r132h mutation status in diffuse glioma patients: implications for classification. Oncotarget 7(21):31393–31400
Zou P, Xu H, Chen P, Yan Q, Zhao L, Zhao P et al (2013) Idh1/idh2 mutations define the prognosis and molecular profiles of patients with gliomas: a meta-analysis. PLoS ONE 8(7):e68782–e68782
Li Y, Wang Z, Wang Y, Zhao Z, Zhang J, Lu J et al (2016) Identification and characterization of lncRNA mediated transcriptional dysregulation dictates lncRNA roles in glioblastoma. Oncotarget 19(29):45027–45041
Han L, Zhang K, Shi Z, Zhang J et al (2012) LncRNA profile of glioblastoma reveals the potential role of lncRNAs, in contributing to glioblastoma pathogenesis. Int J Oncol 40(6):2004–2012
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A et al (2013) String v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 41(Database issue):808–815
Kanehisa M, Araki M, Goto S et al (2008) Kegg for linking genomes to life and the environment. Nucleic Acids Res 36(1):d480–d484
Wagle N, Berger MF, Davis MJ, Blumenstiel B, Defelice M, Pochanard P et al (2012) High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing. Cancer Discov 2(1):82–93
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):1–10
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with bowtie 2. Nat Methods 9(4):357–359
Mao BX, Cai T, Olyarchuk J, Wei (2005) L: Automated genome annotation and pathway identification using the KEGG orthology (KO) as a controlled vocabulary. Bioinformatics 21(19):3787–3793
Mortazavi A, Williams BA, Mccue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat Methods 5(7):621–628
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28(5):511–515
Trapnell C, Pachter L, Salzberg SI (2009) Discovering splice junctions with RNA-seq. Bioinformatics 25(9):1105–1111
Jin M, Zhang T, Liu C, Badeaux M et al (2014) MiRNA-128 suppresses prostate cancer by inhibiting BMI-1 to inhibit tumor-initiating cells. Cancer Res 74(15):4183–4195
Cheng CJ, Bahal R, Babar IA et al (2015) MicroRNA silencing for cancer therapy targeted to the tumour microenvironment. Nature 518(7537):107–110
Tay Y, Kats L, Salmena L, Weiss D et al (2011) Coding-independent regulation of the tumor suppressor pten by competing endogenous mRNAs. Cell 147(2):344–357
Yan Y, Zhang L, Jiang Y, Xu T, Mei Q, Wang H et al (2015) LncRNA and mRNA interaction study based on transcriptome profiles reveals potential core genes in the pathogenesis of human glioblastoma multiforme. J Cancer Res Clin Oncol 141(5):827–838
Johnstone KA, Dubose AJ, Futtner CR, Elmore MD, Brannan CI, Resnick JL (2006) A human imprinting centre demonstrates conserved acquisition but diverged maintenance of imprinting in a mouse model for angelman syndrome imprinting defects. Hum Mol Genet 15(3):393–404
Sutherland HF, Wadey R, Mckie JM, Taylor C, Atif U, Johnstone KA et al (1996) Identification of a novel transcript disrupted by a balanced translocation associated with digeorge syndrome. Am J Hum Genet 59(1):23–31
Willingham AT, Schultz PG (2005) A strategy for probing the function of noncoding RNAs finds a repressor of NFAT. Science 309(5740):1570–1573
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK et al (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495(7441):384–388
Zhao ZJ, Shen J (2015) Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA Biol 4(3):501–589
Xu H, Guo S, Li W, Yu P (2015) The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci Rep 5(1):385–393
Le TD, Zhang J, Liu L, Li J (2016) Computational methods for identifying miRNA sponge interactions. Brief Bioinform 18(4):577–590
Cao Y, Wang P, Ning S, Xiao W, Xiao B, Li X (2016) Identification of prognostic biomarkers in glioblastoma using a long non-coding RNA-mediated, competitive endogenous RNA network. Oncotarget 5(27):41737–41747
Kiang MY, Zhang XQ, Leung KK (2015) Long non-coding RNAs: the key players in glioma pathogenesis. Cancers 7(3):1406–1424
Funding
This study was funded by Sichuan province science and technology support Plan Nos. 2012S20152 and 0040205301911.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We declare that we have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yuan, Y., Jiaoming, L., Xiang, W. et al. Analyzing the interactions of mRNAs, miRNAs, lncRNAs and circRNAs to predict competing endogenous RNA networks in glioblastoma. J Neurooncol 137, 493–502 (2018). https://doi.org/10.1007/s11060-018-2757-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-018-2757-0