Abstract
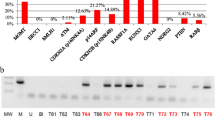
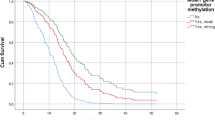
Epigenetic alterations alone or in combination with genetic mechanisms play a key role in brain tumorigenesis. Glioblastoma is one of the most common, lethal and poor clinical outcome primary brain tumors with extraordinarily miscellaneous epigenetic alterations profile. The aim of this study was to investigate new potential prognostic epigenetic markers such as AREG, HOXA11, hMLH1, NDRG2, NTPX2 and Tes genes promoter methylation, frequency and value for patients outcome. We examined the promoter methylation status using methylation-specific polymerase chain reaction in 100 glioblastoma tissue samples. The value for clinical outcome was calculated using Kaplan–Meier estimation with log-rank test. DNA promoter methylation was frequent event appearing more than 45 % for gene. AREG and HOXA11 methylation status was significantly associated with patient age. HOXA11 showed the tendency to be associated with patient outcome in glioblastomas. AREG gene promoter methylation showed significant correlation with poor patient outcome. AREG methylation remained significantly associated with patient survival in a Cox multivariate model including MGMT promoter methylation status. This study of new epigenetic targets has shown considerably high level of analyzed genes promoter methylation variability in glioblastoma tissue. AREG gene might be valuable marker for glioblastoma patient survival prognosis, however further analysis is needed to clarify the independence and appropriateness of the marker.


Similar content being viewed by others
References
Krex D, Klink B, Hartmann C, von Deimling A, Pietsch T, Simon M, Sabel M, Steinbach JP, Heese O, Reifenberger G, Weller M, Schackert G, German Glioma Network (2007) Long-term survival with glioblastoma multiforme. Brain 130:2596–2606
Martinez R, Martin-Subero JI, Rohde V, Kirsch M, Alaminos M, Fernandez AF (2009) A microarray-based DNA methylation study of glioblastoma multiforme. Epigenetics 4:255–264
Rahnenfuhrer J, Beerenwinkel N, Schulz WA, Hartmann C, von Deimling A, Wullich B, Lengauer T (2005) Estimating cancer survival and clinical outcome based on genetic tumor progression scores. Bioinformatics 10:2438–2446
Poplawski T, Tomaszewska K, Galicki M, Morawiec Z, Blasiak J (2008) Promoter methylation of cancer-related genes in gastric carcinoma. Exp Oncol 30:112–116
Etcheverry A, Aubry M, de Tayrac M, Vauleon E, Boniface R, Guenot F, Saikali S, Hamlat A, Riffaud L, Menei P, Quillien V, Mosser J (2010) DNA methylation in glioblastoma: impact on gene expression and clinical outcome. BMC Genomics 11:701
Laffaire J, Everhard S, Idbaih A, Criniere E, Marie Y, de Reynies A, Schiappa R, Mokhtari K, Hoang-Xuan K, Sanson M, Delattre JY, Thillet J, Ducray F (2011) Methylation profiling identifies 2 groups of gliomas according to their tumorigenesis. Neuro Oncol 13:84–98
Noushmehr H, Weisenberger DJ, Diefes K, Phillips HS, Pujara K, Berman BP, Pan F, Pelloski CE, Sulman EP, Bhat KP, Verhaak RGW, Hoadley KA, Hayes DN, Perou CM, Schmidt HK, Ding L, Wilson RK, Van Den Berg D, Shen H, Bengtsson H, Neuvial P, Cope LM, Buckley J, Herman JG, Baylin SB, Laird PW, Aldape K (2010) Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17:510–522
Mueller W, Nutt CL, Ehrich M, Riemenschneider MJ, von Deimling A, van den Boom D, Louis DN (2007) Downregulation of RUNX3 and TES by hypermethylation in glioblastoma. Oncogene 26:583–593
Gomori E, Pal J, Meszaros I, Doczi T, Matolcsy A (2007) Epigenetic inactivation of the hMLH1 gene in progression of gliomas. Diagn Mol Pathol 16:104–107
Tepel M, Roerig P, Wolter M, Gutmann DH, Perry A, Reifenberger G, Riemenschneider MJ (2008) Frequent promoter hypermethylation and transcriptional downregulation of the NDRG2 gene at 14q11.2 in primary glioblastoma. Int J Cancer 123:2080–2086
Louis D, Ohgaki H, Wiestler O, Cavenee W, Burger P, Jouvet A, Scheithauer B, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114:97–109
Ali AH, Kondo K, Namura T, Senba Y, Takizawa H, Nakagawa Y, Toba H, Kenzaki K, Sakiyama S, Tangoku A (2011) Aberrant DNA methylation of some tumor suppressor genes in lung cancers from workers with chromate exposure. Mol Carcinog 50:89–99
Piepoli A, Cotugno R, Merla G, Gentile A, Augello B, Quitadamo M, Merla A, Panza A, Carella M, Maglietta R, D’Addabbo A, Ancona N, Fusilli S, Perri F, Andriulli A (2009) Promoter methylation correlates with reduced NDRG2 expression in advanced colon tumour. BMC Med Genomics 2:11
Ma H, Weng D, Chen Y, Huang W, Pan K, Wang H, Sun J, Wang Q, Zhou Z, Wang H, Xia J (2010) Extensive analysis of D7S486 in primary gastric cancer supports TESTIN as a candidate tumor suppressor gene. Mol Cancer 9:190
Li LC, Dahiya R (2002) MethPrimer: designing primers for methylation PCRs. Bioinformatics 18:1427–1431
Esteller M, Hamilton SR, Burger PC, Baylin SB, Herman JG (1999) Inactivation of the DNA repair gene O6-methylguanine-DNA methyltransferase by promoter hypermethylation is a common event in primary human neoplasia. Cancer Res 59:793–797
Carlson MR, Pope WB, Horvath S, Braunstein JG, Nghiemphu P, Tso CL, Mellinghoff I, Lai A, Liau LM, Mischel PS, Dong J, Nelson SF, Cloughesy TF (2007) Relationship between survival and edema in malignant gliomas: role of vascular endothelial growth factor and neuronal pentraxin 2. Clin Cancer Res 13:2592–2598
Lamber EP, Horwitz AA, Parvin JD (2010) BRCA1 represses amphiregulin gene expression. Cancer Res 70:996–1005
Shoyab M, McDonald VL, Bradley JG, Todaro GJ (1988) Amphiregulin: a bifunctional growth-modulating glycoprotein produced by the phorbol 12-myristate 13-acetate-treated human breast adenocarcinoma cell line MCF-7. Proc Natl Acad Sci USA 85:6528–6532
Mallakin A, Sugiyama T, Kai F, Taneja P, Kendig RD, Frazier DP, Maglic D, Matise LA, Willingham MC, Inoue K (2010) The Arf-inducing transcription factor Dmp1 encodes a transcriptional activator of amphiregulin, thrombospondin-1, JunB and Egr1. Int J Cancer 126:1403–1416
Busser B, Sancey L, Josserand V, Niang C, Favrot MC, Coll JL, Hurbin A (2010) Amphiregulin promotes BAX Inhibition and resistance to gefitinib in non-small-cell lung cancers. Mol Ther 18:528–535
Duhem-Tonnelle V, Bieche I, Vacher S, Loyens A, Maurage C-A, Collier F, Baroncini M, Blond S, Prevot V, Sharif A (2010) Differential distribution of erbB receptors in human glioblastoma multiforme: expression of erbB3 in CD133-positive putative cancer stem cells. J Neuropathol Exp Neurol 69:606–622
Yamashita S, Tsujino Y, Moriguchi K, Tatematsu M, Ushijima T (2006) Chemical genomic screening for methylation-silenced genes in gastric cancer cell lines using 5-aza-2′-deoxycytidine treatment and oligonucleotide microarray. Cancer Sci 97:64–71
Shih IM, Chen L, Wang CC, Gu J, Davidson B, Cope L, Kurman RJ, Xuan J, Wang TL (2010) Distinct DNA methylation profiles in ovarian serous neoplasms and their implications in ovarian carcinogenesis. Am J Obstet Gynecol 203:584
Liang G, Gonzales FA, Jones PA, Orntoft TF, Thykjaer T (2002) Analysis of gene induction in human fibroblasts and bladder cancer cells exposed to the methylation inhibitor 5-aza-2′-deoxycytidine. Cancer Res 62:961–966
Cillo C, Cantile M, Faiella A, Boncinelli E (2001) Homeobox genes in normal and malignant cells. J Cell Physiol 188:161–169
Whitcomb BP, Mutch DG, Herzog TJ, Rader JS, Gibb RK, Goodfellow PJ (2003) Frequent HOXA11 and THBS2 promoter methylation, and a methylator phenotype in endometrial adenocarcinoma. Clin Cancer Res 9:2277–2287
Fiegl H, Windbichler G, Mueller-Holzner E, Goebel G, Lechner M, Jacobs IJ, Widschwendter M (2008) HOXA11 DNA methylationtion–a novel prognostic biomarker in ovarian cancer. Int J Cancer 123:725–729
Murat A, Migliavacca E, Gorlia T, Lambiv WL, Shay T, Hamou MF, de Tribolet N, Regli L, Wick W, Kouwenhoven MCM, Hainfellner JA, Heppner FL, Dietrich PY, Zimmer Y, Cairncross JG, Janzer RC, Domany E, Delorenzi M, Stupp R, Hegi ME (2008) Stem cell related “self-renewal” signature and high epidermal growth factor receptor expression associated with resistance to concomitant chemoradiotherapy in glioblastoma. J Clin Oncol 26:3015–3024
Di Vinci A, Casciano I, Marasco E, Banelli B, Ravetti G, Borzi L, Brigati C, Forlani A, Dorcaratto A, Allemanni G, Zona G, Spaziante R, Gohlke H, Gardin G, Merlo D, Mantovani V, Romani M (2012) Quantitative methylation analysis of HOXA3,7,9 and 10 genes in glioma: association with tumor WHO grade and clinical outcome. J Cancer Res Clin Oncol 138:35–47
Qu X, Zhai Y, Wei H, Zhang C, Xing G, Yu Y, He F (2002) Characterization and expression of three novel differentiation-related genes belong to the human NDRG gene family. Mol Cell Biochem 229:35–44
Deng Y, Yao L, Chau L, Ng SS, Peng Y, Liu X, Au WS, Wang J, Li F, Ji S, Han H, Nie X, Li Q, Kung HF, Leung SY, Lin MC (2003) N-Myc downstream-regulated gene 2 (NDRG2) inhibits glioblastoma cell proliferation. Int J Cancer 106:342–347
Feng L, Xie Y, Zhang H, Wu Y (2011) Down-regulation of NDRG2 gene expression in human colorectal cancer involves promoter methylation and microRNA-650. Biochem Biophys Res Commun 406:534–538
Furuta H, Kondo Y, Nakahata S, Hamasaki M, Sakoda S, Morishita K (2010) NDRG2 is a candidate tumor-suppressor for oral squamous-cell carcinoma. Biochem Biophys Res Commun 391:1785–1791
Lee DC, Kang YK, Kim WH, Jang YJ, Kim DJ, Park IY, Sohn BH, Sohn HA, Lee HG, Lim JS, Kim JW, Song EY, Kim DM, Lee MN, Oh GT, Kim SJ, Park KC, Yoo HS, Choi JY, Yeom YI (2008) Functional and clinical evidence for NDRG2 as a candidate suppressor of liver cancer metastasis. Cancer Res 68:4210–4220
Lusis EA, Watson MA, Chicoine MR, Lyman M, Roerig P, Reifenberger G, Gutmann DH, Perry A (2005) Integrative genomic analysis identifies NDRG2 as a candidate tumor suppressor gene frequently inactivated in clinically aggressive meningioma. Cancer Res 65:7121–7126
Li L, Wang J, Shen X, Wang L, Li X, Liu Y, Shi M, Zhao G, Deng Y (2011) Expression and prognostic value of NDRG2 in human astrocytomas. J Neurol Sci 308:77–82
Li W, Chu D, Chu X, Meng F, Wei D, Li H, Sun B (2011) Decreased expression of NDRG2 is related to poor overall survival in patients with glioma. J Clin Neurosci 18:1534–1537
Martinez R, Esteller M (2010) The DNA methylome of glioblastoma multiforme. Neurobiol Dis 39:40–46
Acknowledgments
This research was funded by a Grant (No. LIG-17/2010) from the Research Council of Lithuania. We kindly thank former Master student of VMU Ms. Dalia Gedmintaitė for her technical assistance in hMLH1 and Tes gene methylation investigation.
Disclosure
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Skiriutė, D., Vaitkienė, P., Ašmonienė, V. et al. Promoter methylation of AREG, HOXA11, hMLH1, NDRG2, NPTX2 and Tes genes in glioblastoma. J Neurooncol 113, 441–449 (2013). https://doi.org/10.1007/s11060-013-1133-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-013-1133-3