Abstract
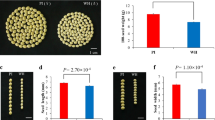
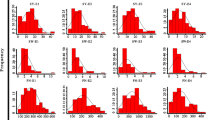
Seed-size/weight traits, controlled by multiple genes in soybean, play an important role in determining seed yield. However, the molecular mechanisms controlling the seed size and weight in soybean remain unclear. In Arabidopsis, P450/CYP78A gene family has been proved extremely relevant to seed size (such as AtCYP78A5, AtCYP78A6 and AtCYP78A9). We found that a soybean GmCYP78A10 gene underwent artificial selection during soybean breeding. The GmCYP78A10a allele mainly distributed in wild soybean (Glycine soja), but has been eliminated in the cultivars during early stage of soybean breeding, while the GmCYP78A10b allele has been accumulated and become the predominant allele in cultivated soybean (G. max). ANOVA analysis showed that the mean seed weight, seed width and seed thickness of soybean varieties with GmCYP78A10b allele was significantly heavier/bigger than those with GmCYP78A10a allele (P < 0.01). The allele could explain 7.2 % variation in seed weight. The pod number of the soybeans with GmCYP78A10b allele significantly decreased compared to those with GmCYP78A10a allele (P < 0.01, R2 = 5.8 %), while other agronomic traits including seed weight/plant were not significantly affected by these two alleles. We speculated that during the early stage of soybean breeding, breeders selected big seed carrying GmCYP78A10b allele, but lowered pod number simultaneously. Overall, the selection did not cause the significantly change in soybean seed yield. Our results suggests that the soybean GmCYP78A10 gene may have a similar function to those genes belonging to P450/CYP78A subfamily in Arabidopsis and provides new information for the genetic control of seed size in soybean.





Similar content being viewed by others
References
Adamski NM, Anastasiou E, Eriksson S, O’Neill CM, Lenhard M (2009) Local maternal control of seed size by KLUH/CYP78A5-dependent growth signaling. Proc Natl Acad Sci USA 106(47):20115–20120
Cannon SB, May GD, Jackson SA (2009) Three sequenced legume genomes and many crop species: rich opportunities for translational genomics. Plant Physiol 151(3):970–977
Cervantes-Martinez I, Sandhu D, Xu M, Ortiz-Perez E, Kato KK, Horner HT, Palmer RG (2009) The male sterility locus ms3 is present in a fertility controlling gene cluster in soybean. J Hered 100(5):565–570
Chauhan BS, Johnson DE (2010) The role of seed ecology in improving weed management strategies in the tropics. Adv Agron 105:221–262
Costa JH, Mota EF, Cambursano MV, Lauxmann MA, de Oliveira LMN, Lima MDS, Orellano EG, de Melo DF (2010) Stress-induced co-expression of two alternative oxidase (VuAox1 and 2b) genes in Vigna unguiculata. J Plant Physiol 167(7):561–570
Davis VM, Gibson KD, Bauman TT, Weller SC, Johnson WG (2009) Influence of weed management practices and crop rotation on glyphosate-resistant horseweed (Conyza canadensis) population dynamics and crop yield-years III and IV. Weed Sci 57(4):417–426
Eriksson S, Stransfeld L, Adamski NM, Breuninger H, Lenhard M (2010) KLUH/CYP78A5-dependent growth signaling coordinates floral organ growth in Arabidopsis. Curr Biol CB 20(6):527–532
Fan C, Xing Y, Mao H, Lu T, Han B, Xu C, Li X, Zhang Q (2006) GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet 112(6):1164–1171
Fang WJ, Wang ZB, Cui RF, Li J, Li YH (2012) Maternal control of seed size by EOD3/CYP78A6 in Arabidopsis thaliana. Plant J 70(6):929–939
Fan H, Wen ZX, Wang CE, Wang F, Xin GN, Zhao TJ, Gai JY (2013) Association analysis between agronomic-processing traits and ssr markers and genetic dissection of specific accessions in Chinese wild soybean population. Acta Agron Sin 39(05):775–788 (in Chinese)
Feldmann KA (2001) Cytochrome P450s as genes for crop improvement. Curr Opin Plant Biol 4(2):162–167
Fu XP, Deng JJ, Yang HX, Masuda T, Goto F, Yoshihara T, Zhao GH (2010) A novel EP-involved pathway for iron release from soya bean seed ferritin. Biochem J 427:313–321
Garcia D, Saingery V, Chambrier P, Mayer U, Jurgens G, Berger F (2003) Arabidopsis haiku mutants reveal new controls of seed size by endosperm. Plant Physiol 131(4):1661–1670
Hao D, Cheng H, Yin Z, Cui S, Zhang D, Wang H, Yu D (2012) Identification of single nucleotide polymorphisms and haplotypes associated with yield and yield components in soybean (Glycine max) landraces across multiple environments. Theor Appl Genet 124:447–458
Helliwell CA, Peacock WJ, Dennis ES (2002) Isolation and functional characterization of cytochrome P450s in gibberellin biosynthesis pathway. Methods Enzymol 357:381–388
Huang F, Chi YJ, Gai JY, Yu DY (2009) Identification of transcription factors predominantly expressed in soybean flowers and characterization of GmSEP1 encoding a SEPALLATA1-like protein. Gene 438(1–2):40–48
Hull AK, Vij R, Celenza JL (2000) Arabidopsis cytochrome P450s that catalyze the first step of tryptophan-dependent indole-3-acetic acid biosynthesis. Proc Natl Acad Sci USA 97(5):2379–2384
Ishimoto M, Rahman SM, Hanafy MS, Khalafalla MM, El-Shemy HA, Nakamoto Y, Kita Y, Takanashi K, Matsuda F, Murano Y, Funabashi T, Miyagawa H, Wakasa K (2010) Evaluation of amino acid content and nutritional quality of transgenic soybean seeds with high-level tryptophan accumulation. Mol Breed 25(2):313–326
Kang HW, Cho YG, Yoon UH (1998) A rapid DNA extraction method for RFLP and PCR analysis from a single dry seed. Plant Mol Biol Rep 16:1–9
Korir PC, Zhang J, Wu K, Zhao T, Gai J (2013) Association mapping combined with linkage analysis for aluminum tolerance among soybean cultivars released in Yellow and Changjiang River Valleys in China. Theor Appl Genet 126(6):1659–1675
Kurauchi T, Matsumoto T, Taneda A, Sano T, Senda M (2009) Endogenous short interfering RNAs of chalcone synthase genes associated with inhibition of seed coat pigmentation in soybean. Breed Sci 59(4):419–426
Li YH, Zheng LY, Corke F, Smith C, Bevan MW (2008) Control of final seed and organ size by the DA1 gene family in Arabidopsis thaliana. Gene Dev 22(10):1331–1336
Li YH, Li W, Zhang C, Yang L, Chang RZ, Gaut BS, Qiu LJ (2010) Genetic diversity in domesticated soybean (Glycine max) and its wild progenitor (Glycine soja) for simple sequence repeat and single-nucleotide polymorphism loci. New Phytol 188:242–253
Li YH, Zhao SC, Ma JX, Li D, Yan L, Li J, Qi XT, Guo XS, Zhang L, He WM, Chang RZ, Liang QS, Guo Y, Ye C, Wang XB, Tao Y, Guan RX, Wang JY, Liu YL, Jin LG, Zhang XQ, Liu ZX, Zhang LJ, Chen J, Wang KJ, Nielsen R, Li RQ, Chen PY, Li WB, Reif JC, Purugganan M, Wang J, Zhang MC, Wang J, Qiu LJ (2013) Molecular footprints of domestication and improvement in soybean revealed by whole genome re-sequencing. BMC Genom 14:579. doi:10.1186/1471-2164-14-579
Liu B, Fujita T, Yan ZH, Sakamoto S, Xu D, Abe J (2007) QTL mapping of domestication-related traits in soybean (Glycine max). Ann Bot 100(5):1027–1038
Lu L, Yan W, Xue W, Shao D, Xing Y (2012) Evolution and association analysis of Ghd7 in rice. PLoS One 7(5):e34021
Neelakandan AK, Song ZH, Wang JQ, Richards MH, Wu XL, Valliyodan B, Nguyen HT, Nes WD (2009) Cloning, functional expression and phylogenetic analysis of plant sterol 24C-methyltransferases involved in sitosterol biosynthesis. Phytochemistry 70(17–18):1982–1998
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Pan XX, Tang YY, Li MR, Wu GJ, Jiang HW (2009) Isoforms of GBSSI and SSII in four legumes and their phylogenetic relationship to their orthologs from other angiosperms. J Mol Evol 69(6):625–634
Qiu LJ, Xing LL, Guo Y, Wang J, Jackson SA, Chang RZ (2013) A platform for soybean molecular breeding: the utilization of core collections for food security. Plant Mol Biol 83(1–2):41–50
Schruff MC, Spielman M, Tiwari S, Adams S, Fenby N, Scott RJ (2006) The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 133(2):251–261
Simard MJ, Panneton B, Longchamps L, Lemieux C, Legere A, Leroux GD (2009) Validation of a management program based on a weed cover threshold model: effects on herbicide use and weed populations. Weed Sci 57(2):187–193
Song XE, Li YH, Chang RZ, Guo PY, Qiu LJ (2010) Population structure and genetic diversity of mini core collection of cultivated soybean (Glycine max (L.) Merr.) in China. Sci Agric Sin 43:2209–2219
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Wang JW, Schwab R, Czech B, Mica E, Weigel D (2008) Dual effects of miR156-targeted SPL genes and CYP78A5/KLUH on plastochron length and organ size in Arabidopsis thaliana. Plant Cell 20(5):1231–1243
Wang LX, Guan Y, Guan RX, Li YH, Ma YS, Dong ZM, Liu X, Zhang HY, Zhang YQ, Liu ZX, Chang RZ, Xu HM, Li LH, Lin FY, Luan WJ, Yan Z, Ning XC, Zhu L, Cui YH, Piao R, Liu Y, Chen PY, Qiu LJ (2006) Establishment of Chinese soybean Glycine max core collections with agronomic traits and SSR markers. Euphytica 151(2):215–223
Wang KJ, Li XH (2012) Phylogenetic relationships, interspecific hybridization and origin of some rare characters of wild soybean in the subgenus Glycine soja in China. Genet Resour Crop Evol 59(8):1673–1685
Weng J, Gu S, Wan X, Gao H, Guo T, Su N, Lei C, Zhang X, Cheng Z, Guo X, Wang J, Jiang L, Zhai H, Wan J (2008) Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res 18(12):1199–1209
Yin ZT, Meng FF, Song HN, Wang XL, Xu XM, Yu DY (2010) Expression quantitative trait loci analysis of two genes encoding rubisco activase in soybean. Plant Physiol 152(3):1625–1637
Zhao W, Park EJ, Chung JW, Park YJ, Chung IM, Ahn JK, Kim GH (2009) Association analysis of the amino acid contents in rice. J Integr Plant Biol 51(12):1126–1137
Zou XA, Li WJ, Zeng W, Chu J, Zhuang YP, Zhang SL (2011) An assessment of seed quality on erythromycin production by recombinant Saccharopolyspora erythraea strain. Bioresour Technol 102(3):3360–3365
Acknowledgments
This research was supported by the State Key Basic Research and Development Plan of China (973) (No. 2010CB125900), the National Natural Science Foundation of China (Nos. 31271753 & 31201226) and Nature Science Foundation of Anhui province (No. 1308085QC49), National Crop Germplasm Platform (2012–004).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, X., Li, Y., Zhang, H. et al. Evolution and association analysis of GmCYP78A10 gene with seed size/weight and pod number in soybean. Mol Biol Rep 42, 489–496 (2015). https://doi.org/10.1007/s11033-014-3792-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-014-3792-3