Abstract
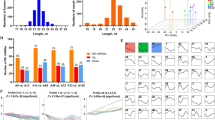
MicroRNAs (miRNAs) are endogenously expressed RNAs consisting of 20–24 nucleotides. These molecules are thought to repress protein translation by binding to target mRNAs. However, biological functions have not been assigned to most of the 175 porcine miRNAs registered in miRBase (release 15.0). In an effort to uncover miR-103 important in pigs, we examined the integrative tissue expression profile and gene ontology (GO) term enrichment of predicted target genes to determine the global biological functions of miR-103. Our results demonstrated that miR-103 is involved in various biological processes including brain development, lipid metabolism, adipocyte differentiation, hematopoiesis, and immunity. Moreover, we also experimentally verified effects of miR-103 in porcine preadipocytes. miR-103 levels increased in differentiating adipocytes, and inhibition of miR-103 effectively inhibited preadipocyte differentiation. In addition, mRNA levels of the putative miR-103 target RAI14 were higher in miR-103 inhibitor-treated adipocytes. These results demonstrate that miR-103 is involved in porcine preadipocyte differentiation and may act through the putative target gene RAI14. In a word, our data provide new insights into the global biological role of miR-103.







Similar content being viewed by others
References
Aravin AA, Lagos-Quintana M, Yalcin A et al (2003) The small RNA profile during Drosophila melanogaster development. Dev Cell 5(2):337–350
Esau C, Davis S, Murray SF et al (2006) miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3(2):87–98
He L, He X, Lowe SW et al (2007) MicroRNAs join the p53 network—another piece in the tumour-suppression puzzle. Nat Rev Cancer 7(11):819–822
Chen CZ, Li L, Lodish HF et al (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303(5654):83–86
Callis TE, Chen JF, Wang DZ (2007) MicroRNAs in skeletal and cardiac muscle development. DNA Cell Biol 26(4):219–225
Rodriguez A, Griffiths-Jones S, Ashurst JL et al (2004) Identification of mammalian microRNA host genes and transcription units. Genome Res 14(10A):1902–1910
Kim J, Krichevsky A, Grad Y et al (2004) Identification of many microRNAs that copurify with polyribosomes in mammalian neurons. Proc Natl Acad Sci USA 101(1):360–365
Coutinho LL, Matukumalli LK, Sonstegard TS et al (2007) Discovery and profiling of bovine microRNAs from immune-related and embryonic tissues. Physiol Genomics 29(1):35–43
Landgraf P, Rusu M, Sheridan R et al (2007) A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129(7):1401–1414
Miska EA, Alvarez-Saavedra E, Townsend M et al (2004) Microarray analysis of microRNA expression in the developing mammalian brain. Genome Biol 5(9):R68
Choong ML, Yang HH, McNiece I (2007) MicroRNA expression profiling during human cord blood-derived CD34 cell erythropoiesis. Exp Hematol 35(4):551–564
Liu SP, Fu RH, Yu HH et al (2009) MicroRNAs regulation modulated self-renewal and lineage differentiation of stem cells. Cell Transplant 18(9):1039–1045
Roldo C, Missiaglia E, Hagan JP et al (2006) MicroRNA expression abnormalities in pancreatic endocrine and acinar tumors are associated with distinctive pathologic features and clinical behavior. J Clin Oncol 24(29):4677–4684
Kulshreshtha R, Ferracin M, Wojcik SE et al (2007) A microRNA signature of hypoxia. Mol Cell Biol 27(5):1859–1867
Marsit CJ, Eddy K, Kelsey KT (2006) MicroRNA responses to cellular stress. Cancer Res 66(22):10843–10848
Iliopoulos D, Malizos KN, Oikonomou P et al (2008) Integrative microRNA and proteomic approaches identify novel osteoarthritis genes and their collaborative metabolic and inflammatory networks. PLoS One 3(11):e3740
Yang GH, Wang F, Yu J et al (2009) MicroRNAs are involved in erythroid differentiation control. J Cell Biochem 107(3):548–556
Xie H, Lim B, Lodish HF (2009) MicroRNAs induced during adipogenesis that accelerate fat cell development are downregulated in obesity. Diabetes 58(5):1050–1057
Guo Y, Chen Z, Zhang L et al (2008) Distinctive microRNA profiles relating to patient survival in esophageal squamous cell carcinoma. Cancer Res 68(1):26–33
Wernersson R, Schierup MH, Jørgensen FG et al (2005) Pigs in sequence space: a 0.66X coverage pig genome survey based on shotgun sequencing. BMC Genomics 6(1):70
Krek A, Grun D, Poy MN et al (2005) Combinatorial microRNA target predictions. Nat Genet 37:495–500
Lewis BP, Shih IH, Jones-Rhoades MW et al (2003) Prediction of mammalian microRNA targets. Cell 115:787–798
Boyle EI, Weng S, Gollub J et al (2004) GO:TermFinder–open source software for accessing gene ontology information and finding significantly enriched gene ontology terms associated with a list of genes. Bioinformatics 20(18):3710–3715
Huixia Li, Luo Xiao, Liu Rongxin et al (2008) Roles of Wnt/_-catenin signaling in adipogenic differentiation potential of adipose-derived mesenchymal stem cells. Mol Cell Endocrinol 291:116–124
Leibrock J, Lottspeich F, Hohn A et al (1989) Molecular cloning and expression of brain-derived neurotrophic factor. Nature 341(6238):149–152
Yokoyama M, Nishi Y, Miyamoto Y et al (1996) Molecular cloning of a human neuroD from a neuroblastoma cell line specifically expressed in the fetal brain and adult cerebellum. Brain Res Mol Brain Res 42(1):135–139
Brunham LR, Singaraja RR, Duong M et al (2009) Tissue-specific roles of ABCA1 influence susceptibility to atherosclerosis. Arterioscler Thromb Vasc Biol 29(4):548–554
Cao Y, Traer E, Zimmerman GA et al (1998) Cloning, expression, and chromosomal localization of human long-chain fatty acid-CoA ligase 4 (FACL4). Genomics 49(2):327–330
Nöhammer C, El-Shabrawi Y, Schauer S et al (2000) cDNA cloning and analysis of tissue-specific expression of mouse peroxisomal straight-chain acyl-CoA oxidase. Eur J Biochem 267(4):1254–1260
Inglis-Broadgate SL, Ocaka L, Banerjee R et al (2005) Isolation and characterization of murine Cds (CDP-diacylglycerol synthase) 1 and 2. Gene 356:19–31
Cesta MF (2006) Normal structure, function, and histology of the spleen. Toxicol Pathol 34(5):455–465
Crosby WH (1959) Normal functions of the spleen relative to red blood cells: a review. Blood 14(4):399–408
Balakirev MY, Tcherniuk SO, Jaquinod M et al (2003) Otubains: a new family of cysteine proteases in the ubiquitin pathway. EMBO Rep 4(5):517–522
Neilson JR, Zheng GX, Burge CB et al (2007) Dynamic regulation of miRNA expression in ordered stages of cellular development. Genes Dev 21(5):578–589
Sonkoly E, Stahle M, Pivarcsi A (2008) MicroRNAs and immunity: novel players in the regulation of normal immune function and inflammation. Semin Cancer Bio 18(2):131–140
Cho IS, Kim J, Seo HY et al (2010) Cloning and characterization of microRNAs from porcine skeletal muscle and adipose tissue. Mol Biol Rep 37(7):3567–3574
Liu G, Fang Y, Zhang H et al (2010) Computational identification and microarray-based validation of microRNAs in Oryctolagus cuniculus. Mol Biol Rep 37(7):3575–3581
Sheng X, Song X, Yu Y et al. (2010) Characterization of microRNAs from sheep (Ovis aries) using computational and experimental analyses. Mol Biol Rep, PubMed PMID: 20140706
Huang Y, Zou Q, Tang SM et al (2010) Computational identification and characteristics of novel microRNAs from the silkworm (Bombyx mori L.). Mol Biol Rep 37(7):3171–3176
Chen K, Rajewsky N (2006) Deep conservation of microRNA target relationships and 3′UTR motifs in vertebrates, flies, and nematodes. Cold Spring Harb Symp Quant Biol 71:149–156
Wilfred BR, Wang WX, Nelson PT (2007) Energizing miRNA research: a review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways. Mol Genet Metab 91(3):209–217
Brandebourg TD, Hu CY (2005) Regulation of differentiating pig preadipocytes by retinoic acid. J Anim Sci 83(1):98–107
Suryawan A, Hu CY (1997) Effect of retinoic acid on differentiation of cultured pig preadipocytes. J Anim Sci 75(1):112–117
Acknowledgments
We are grateful to members of our laboratories for critical reading of the manuscript and helpful discussion. This work was supported by National Natural Science Foundation of China (NO.31072014) and the National High Technology Research and Development Program of China (863 Program) (No. 2006AA10Z138).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, G., Wu, Z., Li, X. et al. Biological role of MicroRNA-103 based on expression profile and target genes analysis in pigs. Mol Biol Rep 38, 4777–4786 (2011). https://doi.org/10.1007/s11033-010-0615-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0615-z