Abstract
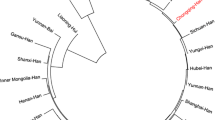
In the present study, we investigated the diversity distributions of allelic frequencies of 15 short tandem repeats (STRs) loci in a sample of Chinese Hui ethnic group in the Ningxia Hui Autonomous Region. The allelic frequencies of the 15 STR loci (D8S1179, D21S11, D7S820, CSF1PO, D3S1358, TH01, D13S317, D16S539, D2S1338, D19S433, vWA, TPOX, D18S51, D5S818 and FGA) were obtained from 2975 unrelated healthy Hui individuals. The STR genotyping data of all the samples were generated by DNA extraction, multiple amplification, GeneScan and genotype analysis. The genetic distances among different populations were calculated by using Nei’s method and a phylogenetic tree was constructed based on the allelic frequencies of the same 15 STR loci using the neighbor-joining method. A total of 185 alleles were observed in the Hui population, with the corresponding allelic frequencies ranging from 0.0002 to 0.5322. Chi-Square tests showed that all STR loci were in Hardy–Weinberg equilibrium. The forensic statistical parameters of all the loci showed high values. The population data in this study were compared with the previously published population data from other ethnics or areas. The Hui population showed significant differences from the Minnan Han, Uigur, Ewenki, Yi, Tibetan, Maonan and Malay ethnic minority groups in some loci, and from the South Morocco population and the Moroccan population in all the loci. Our results are valuable for human individual identification and paternity testing in the Chinese Hui population and are expected to enrich the genetic information resources of Chinese populations.


Similar content being viewed by others
References
Li SX (2001) National Properties of Hui Language. J Ningxia Teach Univ 22:61–64 (in Chinese)
Jin XJ (1989) Language research of Hui nationality. China Natly 2:36–37 (in Chinese)
Ma ZJ (2001) Islamic law’s influence on the Huis’ moral concept and convention. J Ningxia Univ (Soc Sci Ed) 23:95–97 (in Chinese)
Li BP (2009) Dietary customs and cultural boundaries of Hui nationality. J Ningxia Teach Univ (Soc Sci Ed) 30:81–85 (in Chinese)
Edwards A, Civitello A, Hammond HA, Caskey CT (1991) DNA typing and genetic mapping with trimeric and tetrameric tandem repeats. Am J Hum Genet 49:746–756
Ritter N (2007) Missing persons and unidentified remains: the nation’s silent mass disaster. NIJ Journal 256:7
Hammond HA, Jin L, Zhong Y, Caskey CT, Chakraborty R (1994) Evaluation of 13 short tandem repeat loci for use in personal identification applications. Am J Hum Genet 55:175–189
Zhu BF, Wang ZY, Yang CH, Li XS, Zhu J, Yang G, Huang P, Liu Y (2005) Y-chromosomal STR haplotypes in Chinese Uigur ethnic group. Int J Legal Med 119:306–309
Zhu BF, Shen CM, Wu QJ, Deng YJ (2008) Population data of 15 STR loci of Chinese Yi ethnic minority group. Leg Med (Tokyo) 10:220–224
Deng Y, Zhu B, Yu X, Li Y, Fang J, Xiong X, Mu H, Huang Y, Shi X (2007) Genetic polymorphisms of 15 STR loci of Chinese Dongxiang and Salar ethnic minority living in Qinghai province of China. Leg Med (Tokyo) 9:38–42
Zhu B, Yan J, Shen C, Li T, Li Y, Yu X, Xiong X, Mu H, Huang Y, Deng Y (2008) Population genetic analysis of 15 STR loci of Chinese Tu ethnic minority group. Forensic Sci Int 174:255–258
Yan J, Shen C, Li Y, Yu X, Xiong X, Mu H, Huang Y, Deng Y, Yu J (2007) Genetic analysis of 15 STR loci on Chinese Tibetan in Qinghai province. Forensic Sci Int 169:e3–e6
Walsh PS, Metzger DA, Higuchi R (1991) Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10:506–513
Bär W, Brinkmann B, Budowle B, Carracedo A, Gill P, Lincoln P, Mayr W, Olaisen B (1997) DNA recommendations. Further report of the DNA commission of the ISFH regarding the use of short tandem repeat systems. Int J Leg Med 110:175–176
Nei M (1987) Molecular Evolutionary Genetics. Columbia University Press, New York
Ohno Y, Sebetan IM, Akaishi S (1982) A simple method for calculating the probability of excluding paternity with any number of codominant alleles. Forensic Sci Int 19:93–98
Nei M, Roychoudhury AK (1974) Sampling variances of heterozygosity and genetic distance. Genetics 76:379–390
Chen L, He Y, Li S (2006) Genetic analysis of 15 STR loci of Chinese Ewenki ethnic population. J Forensic Sci 51:1408–1409
Zhu J, Li J, Guo Y, Liu K, Zhu B, Liu Y (2005) Population data of 15 STR in Chinese Han population from north of Guangdong. J Forensic Sci 50:1510–1511
Hu SP, Wu DQ, Xu XH, Liu JW, Li B (2005) Genetic profile of 15 STR loci in the Min Nan population in Southeast China. Forensic Sci Int 152:263–265
Zhu B, Wang Z, Wu Q, Sedike ME, Zhu J, Huang P, Xu Y, Liu Y (2005) Genetic analysis of 15 STR loci of Chinese Uigur ethnic population. J Forensic Sci 50:1235–1236
Xu L, Li SF, Deng QY, Zhou LN, Gong JC, Xu R (2007) Polymorphism of 15 STR loci in Mao Nan minority of Guangxi. J Guangxi Med Univ 24:180–183 (in Chinese)
Ossmani HE, Talbi J, Bouchrif B, Chafik A (2009) Allele frequencies of 15 autosomal STR loci in the southern Morocco population with phylogenetic structure among worldwide populations. Leg Med (Tokyo) 11:155–158
Seah LH, Jeevan NH, Othman MI, Jaya P, Ooi YS, Wong PC, Kee SS (2003) STR Data for the AmpFlSTR Identifiler loci in three ethnic groups (Malay, Chinese, Indian) of the Malaysian population. Forensic Sci Int 138:134–137
Bouabdellah M, Ouenzar F, Aboukhalid R, Elmzibri M, Squalli D, Amzazi S (2008) STR data for the 15 AmpFlSTR Identifiler loci in the Moroccan population. Forensic Science Int 1:306–308
Acknowledgments
All authors would like to thank Mr. Zhanhai Wang for his hard work on collecting the blood samples. This project was supported by National Natural Science Foundation of China (NSFC, No. 30700470) and the Item of Science Technology Foundation of Shaanxi Province, People’s Republic of China (No. 2008K08-02).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Deng, Yj., Zhu, Bf., Shen, Cm. et al. Genetic polymorphism analysis of 15 STR loci in Chinese Hui ethnic group residing in Qinghai province of China. Mol Biol Rep 38, 2315–2322 (2011). https://doi.org/10.1007/s11033-010-0364-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-010-0364-z