Abstract
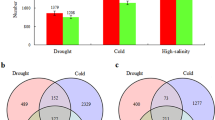
Rapeseed production is limited by abiotic stresses such as drought, salinity, and low temperature. Evidences suggest that common stress response genes are shared by multiple stresses. To study how rapeseed responds to abiotic stresses at transcriptional level and identify genes that regulate multiple abiotic stress tolerance, we investigated transcriptional dynamics of the rapeseed treated by abscisic acid (ABA), salt, dehydration, and cold stresses at two different time points, respectively. A total of 30,908 differentially expressed genes (DEGs) under 4 abiotic stresses were identified. There were 2568 upregulated and 4376 downregulated DEGs (2-fold change) commonly shared by four stresses. Analysis of the DEGs identified significantly enriched gene ontology biological processes under multiple stress conditions. The commonly shared DEGs included 225 upregulated and 294 downregulated transcription factors belonging to 35 and 40 different families, respectively. The representative mostly upregulated and downregulated DEGs at each time point of abiotic stress treatment were presented. We identified a list of core abiotic stress genes commonly regulated by four stresses, which mainly included ERD15, RAB18, LEA14, and transcription factors belonging to ERF, bZIP, and MYBR1 families. The findings of shared abiotic stress responsive genes may help develop strategies for breeding rapeseed varieties with improved tolerance to multiple abiotic stresses.





Similar content being viewed by others
References
Abdeen A, Schnell J, Miki B (2010) Transcriptome analysis reveals absence of unintended effects in drought-tolerant transgenic plants overexpressing the transcription factor ABF3. BMC Genomics 11:69. https://doi.org/10.1186/1471-2164-11-69
Abuqamar S, Ajeb S, Sham A, Enan MR, Iratni R (2013) A mutation in the expansin-like A2 gene enhances resistance to necrotrophic fungi and hypersensitivity to abiotic stress in Arabidopsis thaliana. Mol Plant Pathol 14(8):813–827. https://doi.org/10.1111/mpp.12049
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402. https://doi.org/10.1093/nar/25.17.338
Alves MS, Reis PAB, Dadalto SP, Faria JAQA, Fontes EPB, Fietto LG (2011) A novel transcription factor, ERD15 (early responsive to dehydration 15), connects endoplasmic reticulum stress with an osmotic stress-induced cell death signal. J Biol Chem 286(22):20020–20030. https://doi.org/10.1074/jbc.M111.233494
An J, Sun M, van Velzen R, Ji C, Zheng Z, Limpens E, Bisseling T, Deng X, Xiao S, Pan Z (2018) Comparative transcriptome analysis of Poncirus trifoliata identifies a core set of genes involved in arbuscular mycorrhizal symbiosis. J Exp Bot 69(21):5255–5264. https://doi.org/10.1093/jxb/ery283
Ariel FD, Manavella PA, Dezar CA, Chan RL (2007) The true story of the HD-zip family. Trends Plant Sci 12(9):419–426. https://doi.org/10.1016/j.tplants.2007.08.003
Babula-Skowronska D, Ludwikow A, Ciesla A, Olejnik A, Cegielska-Taras T, Bartkowiak-Broda I, Sadowski J (2015) Involvement of genes encoding ABI1 protein phosphatases in the response of Brassica napus L. to drought stress. Plant Mol Biol 88(4–5):445–457. https://doi.org/10.1007/s11103-015-0334-x
Banaei-Asl F, Bandehagh A, Uliaei ED, Farajzadeh D, Sakata K, Mustafa G, Komatsu S (2015) Proteomic analysis of canola root inoculated with bacteria under salt stress. J Proteome 124:88–111. https://doi.org/10.1016/j.jprot.2015.04.009
Banaei-Asl F, Farajzadeh D, Bandehagh A, Komatsu S (2016) Comprehensive proteomic analysis of canola leaf inoculated with a plant growth-promoting bacterium, Pseudomonas fluorescens, under salt stress. Biochim Biophys Acta 1864(9):1222–1236. https://doi.org/10.1016/j.bbapap.2016.04.013
Bandehagh A, Salekdeh GH, Toorchi M, Mohammadi A, Komatsu S (2011) Comparative proteomic analysis of canola leaves under salinity stress. Proteomics 11(10):1965–1975. https://doi.org/10.1002/pmic.201000564
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Capella M, Ribone PA, Arce AL, Chan RL (2015) Arabidopsis thaliana HomeoBox 1 (AtHB1), a Homedomain-leucine zipper I (HD-Zip I) transcription factor, is regulated by phytochrome-interacting factor 1 to promote hypocotyl elongation. New Phytol 207(3):669–682. https://doi.org/10.1111/nph.13401
Carabelli M, Possenti M, Sessa G, Ruzza V, Morelli G, Ruberti I (2018) Arabidopsis HD-zip II proteins regulate the exit from proliferation during leaf development in canopy shade. J Exp Bot 69(22):5419–5431. https://doi.org/10.1093/jxb/ery331
Coolen S, Proietti S, Hickman R, Davila Olivas NH, Huang PP, Van Verk MC, Van Pelt JA, Wittenberg AH, De Vos M, Prins M, Van Loon JJ, Aarts MG, Dicke M, Pieterse CM, Van Wees SC (2016) Transcriptome dynamics of Arabidopsis during sequential biotic and abiotic stresses. Plant J 86(3):249–267. https://doi.org/10.1111/tpj.13167
Du C, Hu K, Xian S, Liu C, Fan J, Tu J, Fu T (2016) Dynamic transcriptome analysis reveals AP2/ERF transcription factors responsible for cold stress in rapeseed (Brassica napus L.). Mol Gen Genomics 291(3):1053–1067. https://doi.org/10.1007/s00438-015-1161-0
Duan M, Zhang R, Zhu F, Zhang Z, Gou L, Wen J, Dong J, Wang T (2017) A lipid-anchored NAC transcription factor is translocated into the nucleus and activates glyoxalase I expression during drought stress. Plant Cell 29(7):1748–1772. https://doi.org/10.1105/tpc.17.00044
Espartero J, Sanchez-Aguayo I, Pardo JM (1995) Molecular characterization of glyoxalase-I from a higher plant; upregulation by stress. Plant Mol Biol 29(6):1223–1233
Fiebelkorn D, Rahman MJTCJ (2016) Development of a protocol for frost-tolerance evaluation in rapeseed/canola (Brassica napus L.). 4(2):147–152. https://doi.org/10.1016/j.cj.2015.11.004
Georgii E, Jin M, Zhao J, Kanawati B, Schmitt-Kopplin P, Albert A, Winkler JB, Schaffner AR (2017) Relationships between drought, heat and air humidity responses revealed by transcriptome-metabolome co-analysis. BMC Plant Biol 17(1):120. https://doi.org/10.1186/s12870-017-1062-y
Ghosh A, Pareek A, Sopory SK, Singla-Pareek SL (2014) A glutathione responsive rice glyoxalase II, OsGLYII-2, functions in salinity adaptation by maintaining better photosynthesis efficiency and anti-oxidant pool. Plant J 80(1):93–105. https://doi.org/10.1111/tpj.12621
Gou JY, Yu XH, Liu CJ (2009) A hydroxycinnamoyltransferase responsible for synthesizing suberin aromatics in Arabidopsis. Proc Natl Acad Sci U S A 106(44):18855–18860. https://doi.org/10.1073/pnas.0905555106
Guimaraes LA, Mota APZ, Araujo ACG, de Alencar Figueiredo LF, Pereira BM, de Passos Saraiva MA, Silva RB, Danchin EGJ, Guimaraes PM, Brasileiro ACMJPMB (2017) Genome-wide analysis of expansin superfamily in wild Arachis discloses a stress-responsive expansin-like B gene. Plant Mol Biol 94(1):79–96. https://doi.org/10.1007/s11103-017-0594-8
Hara M, Kondo M, Kato T (2013) A KS-type dehydrin and its related domains reduce Cu-promoted radical generation and the histidine residues contribute to the radical-reducing activities. J Exp Bot 64(6):1615–1624. https://doi.org/10.1093/jxb/ert016
He X, Zeng J, Cao F, Ahmed IM, Zhang G, Vincze E, Wu F (2015) HvEXPB7, a novel beta-expansin gene revealed by the root hair transcriptome of Tibetan wild barley, improves root hair growth under drought stress. J Exp Bot 66(22):7405–7419. https://doi.org/10.1093/jxb/erv436
Jaradat MR, Feurtado JA, Huang D, Lu Y, Cutler AJ (2013) Multiple roles of the transcription factor AtMYBR1/AtMYB44 in ABA signaling, stress responses, and leaf senescence. BMC Plant Biol 13:192. https://doi.org/10.1186/1471-2229-13-192
Jian H, Wang J, Wang T, Wei L, Li J, Liu L (2016) Identification of rapeseed MicroRNAs involved in early stage seed germination under salt and drought stresses. Front Plant Sci 7:658. https://doi.org/10.3389/fpls.2016.00658
Kariola T, Brader G, Helenius E, Li J, Heino P, Palva ET (2006) Early responsive to dehydration 15, a negative regulator of abscisic acid responses in Arabidopsis. Plant Physiol 142(4):1559–1573. https://doi.org/10.1104/pp.106.086223
Kende H, Bradford K, Brummell D, Cho HT, Cosgrove D, Fleming A, Gehring C, Lee Y, McQueen-Mason S, Rose J, Voesenek LA (2004) Nomenclature for members of the expansin superfamily of genes and proteins. Plant Mol Biol 55(3):311–314. https://doi.org/10.1007/s11103-004-0158-6
Kerr TCC, Abdel-Mageed H, Aleman L, Lee J, Payton P, Cryer D, Allen RD (2018) Ectopic expression of two AREB/ABF orthologs increases drought tolerance in cotton (Gossypium hirsutum). Plant Cell Environ 41(5):898–907. https://doi.org/10.1111/pce.12906
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12(4):357–360. https://doi.org/10.1038/nmeth.3317
Kim HJ, Cho HS, Pak JH, Kwon T, Lee JH, Kim DH, Lee DH, Kim CG, Chung YS (2018) Confirmation of drought tolerance of ectopically expressed AtABF3 gene in soybean. Mol Cell 41(5):413–422. https://doi.org/10.14348/molcells.2018.2254
Kimura M, Yamamoto YY, Seki M, Sakurai T, Sato M, Abe T, Yoshida S, Manabe K, Shinozaki K, Matsui M (2003) Identification of Arabidopsis genes regulated by high light-stress using cDNA microarray. Photochem Photobiol 77(2):226–233
Kiyosue T, Yamaguchi-Shinozaki K, Shinozaki K (1994) Cloning of cDNAs for genes that are early-responsive to dehydration stress (ERDs) in Arabidopsis thaliana L.: identification of three ERDs as HSP cognate genes. Plant Mol Biol 25(5):791–798
Kumagai T, Ito S, Nakamichi N, Niwa Y, Murakami M, Yamashino T, Mizuno T (2008) The common function of a novel subfamily of B-Box zinc finger proteins with reference to circadian-associated events in Arabidopsis thaliana. Biosci Biotechnol Biochem 72(6):1539–1549. https://doi.org/10.1271/bbb.80041
Labra M, Grassi F, Imazio S, Di Fabio T, Citterio S, Sgorbati S, Agradi E (2004) Genetic and DNA-methylation changes induced by potassium dichromate in Brassica napus L. Chemosphere 54(8):1049–1058. https://doi.org/10.1016/j.chemosphere.2003.10.024
Lau KH, del Rosario Herrera M, Crisovan E, Wu S, Fei Z, Khan MA, Buell CR, Gemenet DCJPD (2018) Transcriptomic analysis of sweet potato under dehydration stress identifies candidate genes for drought tolerance. 2(10):e00092
Lee S-C, Lim M-H, Kim JA, Lee S-I, Kim JS, Jin M, Kwon S-J, Mun J-H, Kim Y-K, Kim HUJM (2008) Transcriptome analysis in Brassica rapa under the abiotic stresses using Brassica 24K oligo microarray. Mol Cells 26(6)
Li F, Xing S, Guo Q, Zhao M, Zhang J, Gao Q, Wang G, Wang W (2011) Drought tolerance through over-expression of the expansin gene TaEXPB23 in transgenic tobacco. J Plant Physiol 168(9):960–966. https://doi.org/10.1016/j.jplph.2010.11.023
Li Q, Yin M, Li Y, Fan C, Yang Q, Wu J, Zhang C, Wang H, Zhou Y (2015) Expression of Brassica napus TTG2, a regulator of trichome development, increases plant sensitivity to salt stress by suppressing the expression of auxin biosynthesis genes. J Exp Bot 66(19):5821–5836. https://doi.org/10.1093/jxb/erv287
Li W, Wang J, Sun Q, Li W, Yu Y, Zhao M, Meng Z (2017) Expression analysis of genes encoding double B-box zinc finger proteins in maize. Funct Integr Genomics 17(6):653–666. https://doi.org/10.1007/s10142-017-0562-z
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7):923–930. https://doi.org/10.1093/bioinformatics/btt656
Liu A, Xiao Z, Li MW, Wong FL, Yung WS, Ku YS, Wang Q, Wang X, Xie M, Yim AK, Chan TF, Lam HM (2018) Transcriptomic reprogramming in soybean seedlings under salt stress. Plant Cell Environ 42(1):98–114. https://doi.org/10.1111/pce.13186
Long L, Yang WW, Liao P, Guo YW, Kumar A, Gao W (2019) Transcriptome analysis reveals differentially expressed ERF transcription factors associated with salt response in cotton. Plant Sci 281:72–81. https://doi.org/10.1016/j.plantsci.2019.01.012
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Lu P, Kang M, Jiang X, Dai F, Gao J, Zhang C (2013) RhEXPA4, a rose expansin gene, modulates leaf growth and confers drought and salt tolerance to Arabidopsis. Planta 237(6):1547–1559. https://doi.org/10.1007/s00425-013-1867-3
Lv A, Su L, Liu X, Xing Q, Huang B, An Y, Zhou P (2018) Characterization of dehydrin protein, CdDHN4-L and CdDHN4-S, and their differential protective roles against abiotic stress in vitro. BMC Plant Biol 18(1):299. https://doi.org/10.1186/s12870-018-1511-2
Ma N, Hu C, Wan L, Hu Q, Xiong J, Zhang C (2017) Strigolactones improve plant growth, photosynthesis, and alleviate oxidative stress under salinity in rapeseed (Brassica napus L.) by regulating gene expression. Front Plant Sci 8:1671. https://doi.org/10.3389/fpls.2017.01671
Miao ZQ, Zhao PX, Mao J, Yu L, Yuan Y, Tang H, Liu ZB, Xiang C (2018) Arabidopsis HB52 mediates the crosstalk between ethylene and auxin by transcriptionally modulating PIN2, WAG1, and WAG2 during primary root elongation. Plant Cell. https://doi.org/10.1105/tpc.18.00584
Naranjo MA, Forment J, Roldan M, Serrano R, Vicente O (2006) Overexpression of Arabidopsis thaliana LTL1, a salt-induced gene encoding a GDSL-motif lipase, increases salt tolerance in yeast and transgenic plants. Plant Cell Environ 29(10):1890–1900. https://doi.org/10.1111/j.1365-3040.2006.01565.x
Oh SJ, Song SI, Kim YS, Jang HJ, Kim SY, Kim M, Kim YK, Nahm BH, Kim JK (2005) Arabidopsis CBF3/DREB1A and ABF3 in transgenic rice increased tolerance to abiotic stress without stunting growth. Plant Physiol 138(1):341–351. https://doi.org/10.1104/pp.104.059147
Peng D, Zhou B, Jiang Y, Tan X, Yuan D, Zhang L (2018) Enhancing freezing tolerance of Brassica napus L. by overexpression of a stearoyl-acyl carrier protein desaturase gene (SAD) from Sapium sebiferum (L.) Roxb. Plant Sci 272:32–41. https://doi.org/10.1016/j.plantsci.2018.03.028
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18(5):1292–1309. https://doi.org/10.1105/tpc.105.035881
Sangtarash MH, Qaderi M, Chinnappa C, Reid DM (2009) Differential sensitivity of canola (Brassica napus) seedlings to ultraviolet-B radiation, water stress and abscisic acid. Environ Exp Bot 66. https://doi.org/10.1016/j.envexpbot.2009.03.004
Savitch LV, Allard G, Seki M, Robert LS, Tinker NA, Huner NP, Shinozaki K, Singh J (2005) The effect of overexpression of two Brassica CBF/DREB1-like transcription factors on photosynthetic capacity and freezing tolerance in Brassica napus. Plant Cell Physiol 46(9):1525–1539. https://doi.org/10.1093/pcp/pci165
Scrucca L, Fop M, Murphy TB, Raftery AE (2016) mclust 5: Clustering, Classification and Density Estimation Using Gaussian Finite Mixture Models. R J 8(1):289–317
Sham A, Al-Azzawi A, Al-Ameri S, Al-Mahmoud B, Awwad F, Al-Rawashdeh A, Iratni R, AbuQamar S (2014) Transcriptome analysis reveals genes commonly induced by Botrytis cinerea infection, cold, drought and oxidative stresses in Arabidopsis. PLoS One 9(11):e113718. https://doi.org/10.1371/journal.pone.0113718
Song K, Kim HC, Shin S, Kim KH, Moon JC, Kim JY, Lee BM (2017) Transcriptome analysis of flowering time genes under drought stress in maize leaves. Front Plant Sci 8:267. https://doi.org/10.3389/fpls.2017.00267
Vojta P, Kokas F, Husickova A, Gruz J, Bergougnoux V, Marchetti CF, Jiskrova E, Jezilova E, Mik V, Ikeda Y, Galuszka P (2016) Whole transcriptome analysis of transgenic barley with altered cytokinin homeostasis and increased tolerance to drought stress. N Biotechnol 33(5 Pt B):676–691. https://doi.org/10.1016/j.nbt.2016.01.010
Wang P, Yang C, Chen H, Song C, Zhang X, Wang D (2017) Transcriptomic basis for drought-resistance in Brassica napus L. Sci Rep 7:40532. https://doi.org/10.1038/srep40532
Wang P, Yang C, Chen H, Luo L, Leng Q, Li S et al (2018) Exploring transcription factors reveals crucial members and regulatory networks involved in different abiotic stresses in Brassica napus L. BMC Plant Biol 18(1):202. https://doi.org/10.1186/s12870-018-1417-z
Wu C, Ma C, Pan Y, Gong S, Zhao C, Chen S, Li H (2013) Sugar beet M14 glyoxalase I gene can enhance plant tolerance to abiotic stresses. J Plant Res 126(3):415–425. https://doi.org/10.1007/s10265-012-0532-4
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10(2):88–94. https://doi.org/10.1016/j.tplants.2004.12.012
Yeats TH, Huang W, Chatterjee S, Viart HM, Clausen MH, Stark RE, Rose JK (2014) Tomato cutin deficient 1 (CD1) and putative orthologs comprise an ancient family of cutin synthase-like (CUS) proteins that are conserved among land plants. Plant J 77(5):667–675. https://doi.org/10.1111/tpj.12422
Yin X, Cui Y, Wang M, Xia X (2017) Overexpression of a novel MYB-related transcription factor, OsMYBR1, confers improved drought tolerance and decreased ABA sensitivity in rice. Biochem Biophys Res Commun 490(4):1355–1361. https://doi.org/10.1016/j.bbrc.2017.07.029b
Yu D, Zhang L, Zhao K, Niu R, Zhai H, Zhang J (2017) VaERD15, a transcription factor gene associated with cold-tolerance in Chinese wild Vitis amurensis. Front Plant Sci 8:297. https://doi.org/10.3389/fpls.2017.00297
Zhang J, Mason AS, Wu J, Liu S, Zhang X, Luo T, Redden R, Batley J, Hu L, Yan G (2015) Identification of putative candidate genes for water stress tolerance in canola (Brassica napus). Front Plant Sci 6:1058. https://doi.org/10.3389/fpls.2015.01058
Zhou S, Sauve R, Thannhauser TW (2009) Proteome changes induced by aluminium stress in tomato roots. J Exp Bot 60(6):1849–1857. https://doi.org/10.1093/jxb/erp065
Zhu G, Li W, Zhang F, Guo W (2018) RNA-seq analysis reveals alternative splicing under salt stress in cotton, Gossypium davidsonii. BMC Genomics 19(1):73. https://doi.org/10.1186/s12864-018-4449-8
Zhu K-M, Xu S, Li K-X, Chen S, Zafar S, Cao W, Wang Z, Ding L-N, Yang Y-H, Li Y-MJMB (2019) Transcriptome analysis of the irregular shape of shoot apical meristem in dt (dou tou) mutant of Brassica napus L. Mol Breed 39(3):39. https://doi.org/10.1007/s11032-019-0943-1
Funding
This work was funded by Major Scientific and Technological Projects of Xinjiang Production and Construction Corps of China (2018AA005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, Y., Ali, U., Zhang, G. et al. Transcriptome analysis reveals genes commonly responding to multiple abiotic stresses in rapeseed. Mol Breeding 39, 158 (2019). https://doi.org/10.1007/s11032-019-1052-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-019-1052-x