Abstract
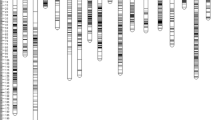
Sex ratio and shell-thickness type are among the main components determining yield in oil palm. An integrated linkage map of oil palm was constructed based on 208 offspring derived from a cross between two tenera palms differing in inherited sex ratio. The map consisted of 210 genomic simple sequence repeats (SSRs), 28 expressed sequence tag SSRs, 185 amplified fragment length polymorphism markers, and the Sh locus, which controls shell-thickness phenotype, distributed across 16 linkage groups covering 1,931 cM, with an average marker distance of 4.6 cM. Quantitative trait locus (QTL) analysis identified eight QTLs across six linkage groups associated with sex ratio and related traits. These QTLs explained 8.1–13.1 % of the total phenotypic variance. The QTL for sex ratio on linkage group 8 overlapped with a QTL for number of male inflorescences. In most cases a specific QTL allele combination was responsible for genotype class mean differences, suggesting that most QTLs in heterozygous oil palm are likely to be segregating for multiple alleles with different degrees of dominance. In addition, two new SSRs were shown to flank the major Sh locus controlling the fruit variety type in oil palm.
Similar content being viewed by others
References
Adam H, Jouannic S, Morcillo F, Richaud F, Duval Y, Tregear J (2006) MADS box genes in oil palm (Elaeis guineensis): patterns in the evolution of the SQUAMOSA, DEFICIENS, GLOBOSA, AGAMOUS, and SEPALLATA. J Mol Evol 62:15–31
Adam H, Jouannic S, Morcillo F, Verdeil JL, Duval Y, Tregear JW (2007a) Determination of flower structure in Elaeis guineensis: do palms use the same homeotic genes as other species? Ann Bot 100:1–12
Adam H, Jouannic S, Orieux Y, Morcillo F, Richaud F, Duval Y, Tregear JW (2007b) Functional characterization of MADS box genes involved in the determination of oil palm flower structure. J Exp Bot 58:1245–1259
Barrett B, Baird I, Woodfield D (2005) A QTL analysis of white clover seed production. Crop Sci 45:1844–1850
Beirnaert A, Vanderweyen R (1941) Contribution à l’étude génétique et biométrique des variétés d’Elaeis guineensis Jacq. Publications de l’Institut National pour l’Etude Agronomique du Congo Belge. Série scientifique no 27
Benbouza H, Jacquemin JM, Baudoin T, Mergeai G (2006) Optimization of a reliable, fast, cheap and sensitive silver staining method to detect SSR markers in polyacrylamide gels. Biotechnol Agron Soc Environ 10:77–81
Billotte N, Marseillac N, Risterucci AM, Adon B, Brottier P, Baurens FC, Singh R, Herrán A, BillotC A, Amblard P, Durand-Gasselin T, Courtois B, Asmono D, Cheah SC, Rohde W, Ritter E, Charrier A (2005) Microsatellite-based high density linkage map in oil palm (Elaeis guineensis Jacq.). Theor Appl Genet 110:754–765
Billotte N, Jourjon MF, Marseillac N, Berger A, Flori A, Asmady H, Adon B, Singh R, Nouy B, Potier F, Cheah SC, Rohde W, Ritter E, Courtois B, Charrier A, Mangin B (2010) QTL detection by multi-parent linkage mapping in oil palm (Elaeis guineensis Jacq.). Theor Appl Genet 120:1673–1687
Bishop DT, Cannings C, Skolnick M, Williamson JA (1983) The number of polymorphic clones required to map the human genome. In: Weir BS (ed) Statistical analysis of DNA sequence data. Marcel and Dekker, New York, pp 181–200
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Corley RHV (1977) Oil palm yield components and yield cycles. In: Earp DA, Newall W (eds) International developments in oil palm. Incorporated Society of Planters, Kuala Lumpur, Malaysia, pp 116–129
Corley RHV, Tinker PB (2003) In: Corley RHV, Tinker PB (eds) The oil palm, 4th edn. Wiley-Blackwell, Oxford
Gawel N, Jarret R (1991) A modified CTAB DNA extraction protocol for Musa and Ipomea. Plant Mol Biol Rep 9:262–266
Grattapaglia D, Sederoff R (1994) Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-testcross: mapping strategy and RAPD markers. Genetics 137:1121–1137
Hackett CA, Broadfoot LB (2003) Effects of genotyping errors, missing values and segregation distortion in molecular marker data on the construction of linkage maps. Heredity 90:33–38
Huang X, Madan A (1999) CAP3: a DNA sequence assembly program. Genome Res 9:868–877
Hulbert SH, Ilott TW, Legg EJ, Lincoln SE, Lander ES, Michelmore RW (1988) Genetic analysis of the fungus, Bremia lactucae, using restriction fragment length polymorphisms. Genetics 120:947–958
Kaufmann K, Muino JM, Jauregui R, Airoldi CA, Smaczniak C, Krajewski P, Angenent GC (2009) Target genes of the MADS transcription factor SEPALLATA3: integration of developmental and hormonal pathways in the Arabidopsis flower. PLoS Biol 7:854–875
Maria M, Clyde MM, Cheah SC (1995) Cytological analysis of Elaeis guineensis (tenera) chromosomes. Elaeis 7:122–134
Martins WS, Lucas DC, Neves KF, Bertioli DJ (2009) WebSat—a web software for microsatellite marker development. Bioinformation 3:282–283
Mason TG, Lewin CJ (1925) Growth and correlation in the oil-palm (Elaeis guineensis). Ann Appl Biol 12:410–421
Mayes S, Jack PL, Marshall DF, Corley RHV (1997) Construction of a RFLP genetic linkage map for oil palm (Elaeis guineensis Jacq.). Genome 40:116–122
Moretzsohn MC, Nunes CDM, Ferreira ME, Grattapaglia D (2000) RAPD linkage mapping of the shell thickness locus in oil palm (Elaeis guineensis Jacq.). Theor Appl Genet 100:63–70
Paterson AH, Damon S, Hewitt JD, Zamir D, Rabinowitch HD, Lincoln SE, Lander ES, Tanksley SD (1991) Mendelian factors underlying quantitative traits in tomato: comparison across species, generations, and environments. Genetics 127:181–197
Rance KA, Mayes S, Price Z, Jack PL, Corley RHV (2001) Quantitative trait loci for yield components in oil palm (Elaeis guineensis Jacq). Theor Appl Genet 103:1302–1310
Rival A, Beule T, Barre P, Hamon S, Duval Y, Noirot M (1997) Comparative flow cytometric estimation of nuclear DNA content in oil palm (Elaeis guineensis Jacq.) tissue cultures and seed-derived plants. Plant Cell Rep 16:884–887
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Seng TY, Mohamed Saad SH, Chin CW, Ting NC, Harminder Singh RS, Zaman FQ, Tan SG, Syed Alwee SSR (2011) Genetic linkage map of a high yielding FELDA Deli × Yangambi oil palm cross. PLoS ONE 6(11):e26593
Singh R, Tan SG, Panandam JM, Rahman RA, Ooi LCL, Low ETL, Sharma M, Jansen J, Cheah SC (2009) Mapping quantitative trait loci (QTLs) for fatty acid composition in an interspecific cross of oil palm. BMC Plant Biol 9:114
Tanksley SD, McCouch SR (1997) Seed banks and molecular maps: unlocking genetic potential from the wild. Science 277:1063–1066
Van Ooijen J, Voorrips R (2001) JoinMap v. 3, software in the calculation of genetic linkage maps. Plant Research International, Wageningen, The Netherlands
Van Ooijen JW, Boer MP, Jansen RC, Maliepaard C (2002) MapQTL 4.0, software for the calculation of QTL positions on genetic maps. Plant Research International, Wageningen, The Netherlands
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 11:4407–4414
Acknowledgments
This study was supported by grants from the National Centre for Genetic Engineering and Biotechnology (BIOTEC), Thailand.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ukoskit, K., Chanroj, V., Bhusudsawang, G. et al. Oil palm (Elaeis guineensis Jacq.) linkage map, and quantitative trait locus analysis for sex ratio and related traits. Mol Breeding 33, 415–424 (2014). https://doi.org/10.1007/s11032-013-9959-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-013-9959-0