Abstract
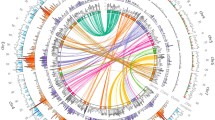
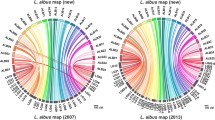
This study reports on the construction of the first genetic maps of subterranean clover (Trifolium subterraneum L.), a diploid, inbreeding annual pasture legume, and alignment of its linkage groups with those of red clover (T. pratense L.) and Medicago truncatula Gaertn. Transferability of red and white clover (T. repens L.) simple sequence repeat (SSR) markers to subterranean clover was observed. A total of 343 SSR loci were mapped into eight subterranean clover linkage groups, with 6–31 loci per linkage group and 27 loci with similar locations between two distinct F 2 mapping populations. Phenotypic data obtained for flowering time, content of three isoflavonoids (formononetin, genistein and biochanin A), hardseededness, leaf markings, calyx pigmentation and hairiness of stems were analyzed, together with genotypic data. Genomic intervals influencing each trait were assigned to one to three chromosome regions, accounting for 5.5–59.8% of the phenotypic variance. Syntenic relationships were observed among subterranean clover, red clover and Medicago truncatula genomes. Comparisons of loci shared between the three species indicated that at least two chromosomal regions have undergone duplications in the subterranean clover genome. Candidate genes for isoflavone content were identified using M. truncatula as a reference genome. Synteny-based segmentation observed in Brassicaceae chromosomes helped to account for the apparent segmental-based relationship between the clover genomes, particularly within the subterranean clover lines. The proposed segmental nature of clover genome could account for the extensive variation observed between the parental genotypes, while not preventing production of fertile intercrosses.



Similar content being viewed by others
References
Barrett B, Griffiths A, Schreiber M, Ellison N, Mercer C, Bouton J, Ong B, Forster J, Sawbridge T, Spangenberg G, Bryan G, Woodfield D (2004) A microsatellite map of white clover. Theor Appl Genet 109(3):596–608
Cannon SB, Sterck L, Rombauts S, Sato S, Cheung F, Gouzy J, Wang X, Mudge J, Vasdewani J, Schiex T, Spannagl M, Monaghan E, Nicholson C, Humphray SJ, Schoof H, Mayer KFX, Rogers J, Quetier F, Oldroyd GE, Debelle F, Cook DR, Retzel EF, Roe BA, Town CD, Tabata S, Van de Peer Y, Young ND (2006) Legume genome evolution viewed through the Medicago truncatula and Lotus japonicus genomes. Proc Natl Acad Sci USA 103(40):14959–14964. doi:10.1073/pnas.0603228103
Chandley AC, McBeath S, Speed RM, Yorston L, Hargreave TB (1987) Pericentric inversion in human chromosome 1 and the risk for male sterility. Br Med J 24(6):325
Choi H-K, Mun J-H, Kim D-J, Zhu H, Baek J-M, Mudge J, Roe B, Ellis N, Doyle J, Kiss GB, Young ND, Cook DR (2004) Estimating genome conservation between crop and model legume species. Proc Natl Acad Sci USA 101(43):15289–15294. doi:10.1073/pnas.0402251101
Collins WJ, Cox RI (1984) Oestrogenic activity in forage legumes. In: Barnes RF, Ball PR, Brougham RW, Marten GC, Minson DJ (eds) Forage legumes for energy-efficient animal production, Palmerston North, New Zealand. USDA-ARS: Springfield Virginia, pp 268–276
Faga B (2007) Installing and configuring CMap. Current protocols in bioinformatics. Wiley, London. doi:10.1002/0471250953.bi0908s17
Francis CM, Gladstones JS (1983) Exploitation of genetic resources in the selection and breeding of new cultivars of Trifolium subterraneum L. (sensu lato). In: Mclvor JG, Bray RA (eds) Genetic resources of forage plants. CSIRO, Melbourne, pp 2–14
Francis CM, Hume ID (1971) The relationship between lignification and flavonoid production in subterranean clover. Aust J Agric Res 24:1–5
Francis CM, Millington AJ (1965) Varietal variation in the isoflavone content of subterranean clover: Its estimation by a microtechnique. Aust J Agric Res 16(4):557–564. doi:10.1071/AR9650557
Fridman E, Zamir D (2003) Functional divergence of a syntenic invertase gene family in tomato, potato, and Arabidopsis. Plant Physiol 131(2):603
George J, Sawbridge TI, Cogan NOI, Gendall AR, Smith KF, Spangenberg GC, Forster JW (2008) Comparison of genome structure between white clover and Medicago truncatula supports homoeologous group nomenclature based on conserved synteny. Genome 51:905–911
Isobe S, Klimenko I, Ivashuta S, Gau M, Kozlov NN (2003) First RFLP linkage map of red clover (Trifolium pratense L.) based on cDNA probes and its transferability to other red clover germplasm. Theor Appl Genet 108:105–112
Isobe S, Kolliker R, Hisano H, Sasamoto S, Wada T, Klimenko I, Okumura K, Tabata S (2009) Construction of a consensus linkage map for red clover (Trifolium pratense L.). BMC Plant Biol 9(1):57
Johnson ME, Viggiano L, Bailey JA, Abdul-Rauf M, Goodwin G, Rocchi M, Eichler EE (2001) Positive selection of a gene family during the emergence of humans and African apes. Nature 413(6855):514–519
Jones ES, Hughes LJ, Drayton MC, Abberton MT, Michaelson-Yeates TPT, Bowen C, Forster JW (2003) An SSR and AFLP molecular marker-based genetic map of white clover (Trifolium repens L.). Plant Sci 165 (3):531–539
Kölliker R, Jones ES, Drayton MC, Dupal MP, Forster JW (2001) Development and characterisation of simple sequence repeat (SSR) markers for white clover (Trifolium repens L.). TAG Theor Appl Genet 102(2):416–424
Kölliker R, Enkeri J, Widmer F (2006) Characterization of novel microsatellite loci for red clover (Trifolium pratense L.) from enriched genomic libraries. Mol Ecol Notes 6:50–53
Kosambi DD (1944) The estimation of map distance from recombination values. Ann Eugen 12(3):172–175
Kosman E, Leonard KJ (2005) Similarity coefficients for molecular markers in studies of genetic relationships between individuals for haploid, diploid, and polyploid species. Mol Ecol 14(2):415–424
Li L, Modolo LV, Escamilla-Trevino LL, Achnine L, Dixon RA, Wang X (2007) Crystal structure of Medicago truncatula UGT85H2—insights into the structural basis of a multifunctional (iso)flavonoid glycosyltransferase. J Mol Biol 370(5):951–963
Liu C-J, Huhman D, Sumner LW, Dixon RA (2003) Regiospecific hydroxylation of isoflavones by cytochrome P450 81E enzymes from Medicago truncatula. The Plant J 36(4):471–484. doi:10.1046/j.1365-313X.2003.01893.x
Manly FF, Cudmore RH, Meer JM (2001) Map Manager QTX, cross-platform software for genetic mapping. Mamm Genome 12:930–932
Mefford HC, Eichler EE (2009) Duplication hotspots, rare genomic disorders, and common disease. Curr Opin Genet Dev 19(3):196–204
Morley FHW (1961) Subterranean clover. In: Norman AG (ed) Advances in agronomy, vol 13. Elsevier, Burlington, pp 125–163
Nichols PGH (2004) Evolution in sown mixtures of subterranean clover (Trifolium subterraneum L.). University of Western Australia, Perth
Nichols PGH, Collins WJ, Barbetti MJ (1996) Registered cultivars of subterranean clover—their characteristics, origin and identification., vol Bulletin No. 4327. Agriculture Western Australia, South Perth
Nichols PGH, Loi A, Nutt BJ, Evans PM, Craig AD, Pengelly BC, Dear BS, Lloyd DL, Revell CK, Nair RM, Ewing MA, Howieson JG, Auricht GA, Howie JH, Sandral GA, Carr SJ, de Koning CT, Hackney BF, Crocker GJ, Snowball R, Hughes SJ, Hall EJ, Foster KJ, Skinner PW, Barbetti MJ, You MP (2007) New annual and short-lived perennial pasture legumes for Australian agriculture—15 years of revolution. Field Crops Res 104(1–3):10–23. doi:10.1016/j.fcr.2007.03.016
Nichols P, Cocks P, Francis C (2009) Evolution over 16 years in a bulk-hybrid population of subterranean clover (Trifolium subterraneum L.) at two contrasting sites in south-western Australia. Euphytica 169(1):31–48
Ouyang S, Zhu W, Hamilton J, Lin H, Campbell M, Childs K, Thibaud-Nissen F, Malek RL, Lee Y, Zheng L (2006) The TIGR rice genome annotation resource: improvements and new features. Nucleic Acids Res 35(suppl 1):D883
Parkin IAP, Gulden SM, Sharpe AG, Lukens L, Trick M, Osborn TC, Lydiate DJ (2005) Segmental structure of the Brassica napus genome based on comparative analysis with Arabidopsis thaliana. Genetics. doi:10.1534/genetics.105.042093
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457(7229):551–556
Quinlivan BJ (1961) The effect of constant and fluctuating temperatures on the permeability of the hard seeds of some legume species. Aust J Agric Res 12(6):1009–1022. doi:10.1071/AR9611009
Salisbury PA, Halloran GM (1983) Inheritance of rate of breakdown of seed coat impermeability in subterranean clover (Trifolium subterraneum L.). Euphytica 32(3):807–819
Salisbury PA, Aitken Y, Halloran GM (1987) Genetic control of flowering time and its component processes in subterranean clover (Trifolium subterraneum L.). Euphytica 36(3):887–902
Sato S, Isobe S, Asamizu E, Ohmido N, Kataoka R, Nakamura Y, Kaneko T, Sakurai N, Okumura K, Klimenko I, Sasamoto S, Wada T, Watanabe A, Kohara M, Fujishiro T, Tabata S (2005) Comprehensive structural analysis of the genome of red clover (Trifolium pratense L.). DNA Res 12:301–364
Slattery HD (1981) Physiological and genetic control of seed coat impermeability in subterranean clover (Trifolium subterraneum L.). University of Western Australia, Perth
Tan BH, Collins WJ (1987) Multi-allelic nature of the locus controlling leaf marking in subterranean clover. Aust J Agric Res 38(3):547–558. doi:10.1071/AR9870547
Taylor GB (2005) Hardseededness in Mediterranean annual pasture legumes in Australia: a review. Aust J Agric Res 56(7):645–662
Varshney R, Bertioli D, Moretzsohn M, Vadez V, Krishnamurthy L, Aruna R, Nigam S, Moss B, Seetha K, Ravi K, He G, Knapp S, Hoisington D (2009) The first SSR-based genetic linkage map for cultivated groundnut (Arachis hypogaea L.). Theor Appl Genet 118(4):729–739
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93(1):77–78
Wang X (2011) Structure, function, and engineering of enzymes in isoflavonoid biosynthesis. Func Integr Genomics 11(1):13–22. doi:10.1007/s10142-010-0197-9
Wang J, Drayton MC, George J, Cogan NOI, Baillie RC, Hand ML, Kearney G, Trigg P, Erb S, Wilkinson T, Bannan N, Forster JW, Smith KF (2010) QTL analysis of salt stress tolerance in white clover (Trifolium repens L.). TAG Theor Appl Genet 120:607–619
Young ND, Cannon SB, Sato S, Kim D, Cook DR, Town CD, Roe BA, Tabata S (2005) Sequencing the Genespaces of Medicago truncatula and Lotus japonicus. Plant Physiol 137(4):1174–1181. doi:10.1104/pp.104.057034
Zeng ZB (1993) Theoretical basis for separation of multiple linked gene effects in mapping quantitative trait loci. Proc Natl Acad Sci USA 90:10972–10976
Zeng ZB (1994) Precision mapping of quantitative trait loci. Genetics 136:1457–1468
Zhang L, Lu HHS, Chung W, Yang J, Li WH (2005) Patterns of segmental duplication in the human genome. Mol Biol Evol 22(1):135
Zhang Y, Sledge M, Bouton J (2007) Genome mapping of white clover (Trifolium repens L.) and comparative analysis within the Trifolieae using cross-species SSR markers. Theor Appl Genet 114(8):1367–1378
Acknowledgments
We are grateful to Hiroshi Hisano (KDRI, currently at Samuel Noble Foundation in USA), Rosemarie Lugg and Nader Aryamanesh (from UWA) for their help in conducting some experiments and data collection. Financial and other support from the Australian Research Council (ARC), Department of Agriculture and Food Western Australia (DAFWA) and Kazusa DNA Research Institute (KDRI) is gratefully acknowledged. Troy Faithful was the recipient of a DAFWA studentship award. Financial support was also obtained from a bequest to UWA by the late Frank Ford.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11032_2011_9612_MOESM2_ESM.pdf
Fig. S1 Genetic linkage maps of T. subterraneum. The order of LGs has been chosen on the basis of syntenic markers between these maps and the red clover map. QTLs identified for studied traits, as shown in Table 2, are indicated by bars on right-hand side of the linkage groups. Numbers on the left of each group are Kosambi map distances (PDF 22 kb)
11032_2011_9612_MOESM3_ESM.pdf
Fig. S2 Comparative map showing matching of T. subterraneum linkage groups (LGa–LGg) with those of T. pratense (LG1–LG7). T. subterraneum population 92S05 is shown on the left and 92S80 is on the right for each T. pratense LG. Red lines compare positions of shared markers (highlighted in red) between T. pratense and T. subterraneum LGs. Numbers in brackets indicate the number of mapped loci for each LG. Note: the LGe for T. subterraneum population 92S80 did not align with LG5 of T. pratense (PDF 76 kb)
11032_2011_9612_MOESM4_ESM.pdf
Fig. S3 Comparison of SSR marker loci for linkage groups LGf and LGc of T. subterraneum populations 92S05 and 92S80, respectively, and simplified T. pratense linkage group LG6, illustrating synteny between the three LGs. Markers in common with two or more chromosomes are indicated by horizontal bars. Solid rectangles represent the same chromosome segments, with arrows representing the direction of synteny. Numbers represent distance in centimorgans. Note: the entire linkage map of T. pratense LG6 is not shown due to the very high density of molecular markers (PDF 18 kb)
11032_2011_9612_MOESM5_ESM.pdf
Fig. S4 Hypothetical inter-specific shared segments containing isoflavone content genes. Units are cM in clover LGs and Mbp in Medicago truncatula chromosomes (PDF 30 kb)
Rights and permissions
About this article
Cite this article
Ghamkhar, K., Isobe, S., Nichols, P.G.H. et al. The first genetic maps for subterranean clover (Trifolium subterraneum L.) and comparative genomics with T. pratense L. and Medicago truncatula Gaertn. to identify new molecular markers for breeding. Mol Breeding 30, 213–226 (2012). https://doi.org/10.1007/s11032-011-9612-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-011-9612-8