Abstract
The chemical space covered by compounds involved in metabolic reactions was compared with that of a random dataset of purchasable compounds by chemoinformatics techniques. The comparison was based on 3D structure, 2D structure, or descriptors of global properties, by means of self-organizing maps, random forests, and classification trees. The overlap between metabolites and non-metabolites was observed to be the least in the space defined by the global descriptors, the most discriminatory features being the number of OH groups, presence of aromatic systems, and molecular weight. Discrimination between the two datasets was achieved with accuracy up to 97%. Models were built to produce a metabolite-likeness parameter. A relationship between metabolite-likeness and ready biodegradability was observed.
Similar content being viewed by others
Abbreviations
- CPG NN:
-
Counterpropagation neural network
- CV:
-
Cross-validation
- MITI:
-
Japanese Ministry of International Trade and Industry
- OOB:
-
Out-of-bag
- RDF:
-
Radial distribution function
- RF:
-
Random forest
- SOM:
-
Self-organizing map
References
Kulkarni SA, Zhu J, Blechinger S (2005) In silico techniques for the study and prediction of xenobiotic metabolism: a review. Xenobiotica 35:955–973, and references therein
Ellis LBM, Roe D, Wackett LP (2006) The University of Minnesota biocatalysis/biodegradation database: the first decade. Nucleic Acids Res 34:D517–D521
Holliday GL, Bartlett GJ, Almonacid DE, O’Boyle NM, Murray-Rust P, Thornton JM, Mitchell JBO (2005) MACiE: a database of enzyme reaction mechanisms. Bioinformatics 21:4315–4316
Reitz M, Sacher O, Tarkhov A, Trümbach D, Gasteiger J (2004) Enabling the exploration of biochemical pathways. Org Biomol Chem 2:3226–3237
Hattori M, Okuno Y, Goto S, Kanehisa M (2003) Development of a chemical structure comparison method for integrated analysis of chemical and genomic information in the metabolic pathways. J Am Chem Soc 125:11853–11865
Nobeli I, Ponstingl H, Krissinel EB, Thornton JM (2003) A structure-based anatomy of E. coli metabolome. J Mol Biol 334:697–719
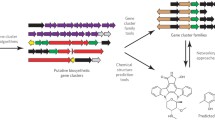
Latino DARS, Aires-de-Sousa J (2006) Genome-scale classification of metabolic reactions: a chemoinformatics approach. Angew Chem Int Ed 45:2066–2069
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280
Kanehisa M, Goto S, Hattori M, Aoki-Kinoshita KF, Itoh M, Kawashima S, Katayama T, Araki M, Hirakawa M (2006) From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Res 34:D354–D357
Irwin JJ, Shoichet BK (2005) ZINC—a free database of commercially available compounds for virtual screening. J Chem Inf Comput Sci 45:177–182
Henkel T, Brunne RM, Muller H, Reichel F (1999) Statistical investigation into the structural complementarity of natural products and synthetic compounds. Angew Chem Int Ed Engl 38:643–647
Stahura FL, Jeffrey WG, Xue L, Bajorath J (2000) Distinguishing between natural products and synthetic molecules by descriptor Shannon entropy analysis and binary QSAR calculations. J Chem Inf Comput Sci 40:1245–1252
Lee M-L, Schneider G (2001) Scaffold architecture and pharmacophoric properties of natural products and trade drugs: application in the design of natural product-based combinatorial libraries. J Comb Chem 3:284–289
Feher M, Schmidt JM (2003) Property distributions: differences between drugs, natural products, and molecules from combinatorial chemistry. J Chem Inf Comput Sci 43:218–227
Koch MA, Schuffenhauer A, Scheck M, Wetzel S, Casaulta M, Odermatt A, Ertl P, Waldmann H (2005) Charting biologically relevant chemical space: a structural classification of natural products (SCONP). Proc Natl Acad Sci USA 102:17272–17277
Karakoc E, Sahinalp SC, Cherkasov A (2006) Comparative QSAR and fragments distribution analysis of drugs, druglikes, metabolic substances, and antimicrobial compounds. J Chem Inf Model 46:2167–2182
Cherkasov A (2006) Can ‹bacterial-metabolite-likeness’ model improve odds of ‹in silico’ antibiotic discovery? J Chem Inf Model 46:1214–1222
Kohonen T (1989) Self-organization and associative memory, 3rd edn. Springer, Berlin Heidelberg New York
Breiman L (2001) Random forests. Mach Learn 45:5–32
Breiman L, Friedman JH, Olshen RA, Stone CJ (2000) Classification and regression trees. Chapman & Hall/CRC, Boca Raton
Hemmer MC, Steinhauer V, Gasteiger J (1999) The prediction of the 3D structure of organic molecules from their infrared spectra. Vib Spectrosc 19:151–164
Organisation for Economic Co-operation and Development. http://www.oecd.org
PETRA can be tested on the website http://www2.chemie.uni-erlangen.de and is developed by Molecular Networks GmbH (Erlangen, Germany). http://www.mol-net.de
National Institute of Technology and Evaluation, Japan. http://www.safe.nite.go.jp/english/db.html
CORINA software is available from Molecular Networks GmbH (Erlangen, Germany). http://www.mol-net.de
Aires-de-Sousa J (2002) Java tools for neural networks. Chemometrics Intell Lab Syst 61:167–173
The JATOON applets are available at http://www.dq.fct.unl.pt/staff/jas/jatoon
Svetnik V, Liaw A, Tong C, Culberson JC, Sheridan RP, Feuston BP (2003) Random forest: a classification and regression tool for compound classification and QSAR modeling. J Chem Inf Comput Sci 43:1947–1958
R Development Core Team (2004) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0. http://www.R-project.org
Fortran original by Breiman L, Cutler A, R port by Liaw A, Wiener M (2004)
Zupan J, Gasteiger J (1999) Neural networks in chemistry and drug design, 2nd edn. Wiley-VCH, Weinheim
Dobson CM (2004) Chemical space and biology. Nature 432:824–828
Acknowledgments
S. Gupta acknowledges Fundação para a Ciência e Tecnologia (Lisbon, Portugal) for the postdoctoral grant SFRH/BPD/14475/2003 co-funded by the POCI 2010 EU program. Molecular Networks GmbH (Erlangen, Germany) is acknowledged for access to PETRA and CORINA software packages. The authors thank Dr Robert Boethling for assistance with the MITI data.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
The importance of global molecular descriptors obtained from RF for the discrimination between metabolites and non-metabolites. (a) Mean decrease in accuracy. (b) Gini criterion. A1 number of atoms, A2 number of bonds, A3 molecular weight, A4 number of aromatic atoms, A5 mean molecular polarizability, A6 number of NH groups, A7 number of NH2 groups, A8 number of non-hydrogen atoms, A9 number of OH groups, A10 number of O atoms, A11 number of N atoms, A12 number of F atoms, A13 number of P atoms, A14 number of S atoms, A15 number of Cl atoms, A16 number of Br atoms, A17 number of I atoms, A18 total number of halogen atoms, A19 minimum partial atomic charge, A20 maximum partial atomic charge, A21 minimum charge on H, A22 maximum charge on H, A23 aromatic delocalization energy, A24 ring strain energy, A25 maximum length of C chain
Rights and permissions
About this article
Cite this article
Gupta, S., Aires-de-Sousa, J. Comparing the chemical spaces of metabolites and available chemicals: models of metabolite-likeness. Mol Divers 11, 23–36 (2007). https://doi.org/10.1007/s11030-006-9054-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-006-9054-0