Abstract
Context
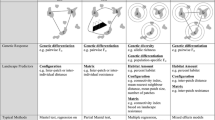
Landscape structure shapes the genetic structure of populations by delimiting spatial patterns of dispersal and reproduction across generations. Thus, descriptions of human-altered landscapes can be used to predict demographic and evolutionary outcomes of populations. Effectively measuring landscape structure to predict genetic structure requires that we understand the relative importance of distinct components of landscape structure (e.g., habitat amount and configuration) in creating spatial patterns of genetic variation.
Objectives
We thus developed an individual-based simulation model to test predictions about the relative importance of habitat amount and configuration in producing genetic structure. We also investigated the independent relationships between components of landscape structure and the population dynamics that underlie genetic effects.
Methods
We ran experiments in which we allowed gene flow and population size to vary as emergent outcomes of the interactions between hypothetical populations and heterogeneous landscapes.
Results
We found that the amount of habitat in a landscape is a much better predictor of genetic structure than is habitat configuration. This pattern holds across a range of landscapes and dispersal distances and behaviors. When habitat is non-contiguous (i.e., fragmented), habitat amount mediates production of genetic differentiation by regulating both the size and isolation of habitat patches, which in turn regulate population size and gene flow.
Conclusions
These results suggest that habitat amount, a simple measure that is easy to calculate, may often be the best metric for predicting population genetic structure and that when possible, measures of habitat amount and population size should be incorporated into landscape genetic studies.







Similar content being viewed by others
References
Akcakaya HR, Atwood JL (1997) A habitat based metapopulation model of the California gnatcatcher. Conserv Biol 11(2):422–434
Balkenhol N, Gugerli F, Cushman SA, Waits LP, Coulon A, Arntzen JW, Holderegger R, Wagner HH (2009) Identifying future research needs in landscape genetics: where to from here? Landscape Ecol 24(4):455–463
Balkenhol N, Pardini R, Cornelius C, Fernandes F, Sommer S (2013) Landscape-level comparison of genetic diversity and differentiation in a small mammal inhabiting different fragmented landscapes of the Brazilian Atlantic Forest. Conserv Genet 14(2):355–367
Beale CM, Lennon JJ, Yearsley JM, Brewer MJ, Elston DA (2010) Regression analysis of spatial data. Ecol Lett 13(2):246–264
Betts MG, Forbes GJ, Diamond AW (2007) Thresholds in songbird occurrence in relation to landscape structure. Conserv Biol 21(4):1046–1058
Betts MG, Fahrig L, Hadley AS, Halstead KE, Bowman J, Robinson WD, Wiens JA, Lindenmayer DB (2014) A species-centered approach for uncovering generalities in organism responses to habitat loss and fragmentation. Ecography 37(6):517–527
Bowcock AM, Ruiz-Linares A, Tomfohrde J, Minch E, Kidd JR, Cavalli-Sforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Nature 368:455–457
Bowler DE, Benton TG (2005) Causes and consequences of animal dispersal strategies: relating individual behaviour to spatial dynamics. Biol Rev 80(2):205–225
Bruggeman DJ, Wiegand T, Fernandez N (2010) The relative effects of habitat loss and fragmentation on population genetic variation in the red-cockaded woodpecker (Picoides borealis). Mol Ecol 19(17):3679–3691
Bunn AG, Urban DL, Keitt TH (2000) Landscape connectivity: a conservation application of graph theory. J Environ Manag 59(4):265–278
Burkey TV (1997) Metapopulation extinction in fragmented landscapes: using bacteria and protozoa communities as model ecosystems. Am Nat 150(5):568–591
Bustamante J, Seoane J (2004) Predicting the distribution of four species of raptors (Aves: Accipitridae) in southern Spain: statistical models work better than existing maps. J Biogeogr 31:295–306
Chapman DS, Dytham C, Oxford GS (2007) Modelling population redistribution in a leaf beetle: an evaluation of alternative dispersal functions. J Anim Ecol 76(1):36–44
Conradt L, Bodsworth EJ, Roper TJ, Thomas CD (2000) Non-random dispersal in the butterfly Maniola jurtina: implications for metapopulation models. Proc R Soc B 267(1452):1505–1510
Coulon A, Fitzpatrick JW, Bowman R, Lovette IJ (2010) Effects of habitat fragmentation on effective dispersal of Florida scrub-jays. Conserv Biol 24(4):1080–1088
Coulon A, Fitzpatrick JW, Bowman R, Lovette IJ (2012) Mind the gap: genetic distance increases with habitat gap size in Florida scrub jays. Biol Lett 8(4):582–585
Cushman SA, McKelvey KS, Hayden J, Schwartz MK (2006) Gene flow in complex landscapes: testing multiple hypotheses with causal modeling. Am Nat 168(4):486–499
Cushman SA, Shirk A, Landguth EL (2012) Separating the effects of habitat area, fragmentation and matrix resistance on genetic differentiation in complex landscapes. Landscape Ecol 27(3):369–380
Cushman SA, Shirk AJ, Landguth EL (2013) Landscape genetics and limiting factors. Conserv Genet 14:263–274
Ezard THG, Travis JMJ (2006) The impact of habitat loss and fragmentation on genetic drift and fixation time. Oikos 114(2):367–375
Fahrig L (1997) Relative effects of habitat loss and fragmentation on population extinction. J Wildl Manag 61(3):603–610
Fahrig L (1998) When does fragmentation of breeding habitat affect population survival? Ecol Model 105(2–3):273–292
Fahrig L (2003) Effects of habitat fragmentation on biodiversity. Annu Rev Ecol Evol Syst 34:487–515
Fahrig L (2013) Rethinking patch size and isolation effects: the habitat amount hypothesis. J Biogeogr 40(9):1649–1663
Flather CH, Bevers M (2002) Patchy reaction-diffusion and population abundance: the relative importance of habitat amount and arrangement. Am Nat 159(1):40–56
Fletcher RJ Jr (2006) Emergent properties of conspecific attraction in fragmented landscapes. Am Nat 168(2):207–219
Funk WC, Blouin MS, Corn PS, Maxell BA, Pilliod DS, Amish S, Allendorf FW (2005) Population structure of Columbia spotted frogs (Rana luteiventris) is strongly affected by the landscape. Mol Ecol 14(2):483–496
Graves TA, Wasserman TN, Ribeiro MC, Landguth EL, Spear SF, Balkenhol N, Higgins CB, Fortin MJ, Cushman SA, Waits LP (2012) The influence of landscape characteristics and home-range size on the quantification of landscape-genetics relationships. Landscape Ecol 27(2):253–266
Grimm V, Railsback SF (2005) Individual-based modeling and ecology. Princeton University Press, Princeton
Gustafson EJ (1998) Quantifying landscape spatial pattern: what is the state of the art? Ecosystems 1(2):143–156
Gustafson EJ, Parker GR (1994) Using an index of habitat patch proximity for landscape design. Landsc Urban Plan 29(2–3):117–130
Haines-Young R, Chopping M (1996) Quantifying landscape structure: a review of landscape indices and their application to forested landscapes. Prog Phys Geogr 20(4):418–445
Holderegger R, Wagner HH (2008) Landscape genetics. Bioscience 58(3):199–207
Honnay O, Coart E, Butaye J, Adriaens D, Van Glabeke S, Roldan-Ruiz I (2006) Low impact of present and historical landscape configuration on the genetics of fragmented Anthyllis vulneraria populations. Biol Conserv 127(4):411–419
Hutchison DW, Templeton AR (1999) Correlation of pairwise genetic and geographic distance measures: inferring the relative influences of gene flow and drift on the distribution of genetic variability. Evolution 53(6):1898–1914
Jackson HB, Fahrig L (2012) What size is a biologically relevant landscape? Landscape Ecol 27(7):929–941
Jackson ND, Fahrig L (2014) Landscape context affects genetic diversity at a much larger spatial extent than population abundance. Ecology 95(4):871–881
Kanuch P, Jarcuska B, Schlosserova D, Sliacka A, Paule L, Kristin A (2012) Landscape configuration determines gene flow and phenotype in a flightless forest-edge ground-dwelling bush-cricket Pholidoptera griseoaptera. Evol Ecol 26(6):1331–1343
Keyghobadi N (2007) The genetic implications of habitat fragmentation for animals. Can J Zool 85(10):1049–1064
Kimura M, Ohta T (1969) The average number of generations until fixation of a mutant gene in a finite population. Genetics 61(3):763–771
Kot M, Lewis MA, van den Driessche P (1996) Dispersal data and the spread of invading organisms. Ecology 77:2027–2042
Kozakiewicz M (1995) Resource tracking in space and time. In: Hansson L, Fahrig L, Merriam G (eds) Mosaic landscapes and ecological processes. Chapman & Hall, London, pp 136–148
Landguth EL, Cushman SA, Schwartz MK, McKelvey KS, Murphy M, Luikart G (2010) Quantifying the lag time to detect barriers in landscape genetics. Mol Ecol 19(19):4179–4191
Landguth EL, Fedy BC, Oyler-McCance SJ, Garey GL, Emel SL, Mumma M, Wagner HH, Fortin MJ, Cushman SA (2012) Effects of sample size, number of markers, and allelic richness on the detection of spatial genetic pattern. Mol Ecol Resour 12(2):276–284
Lange R, Diekoetter T, Schiffmann LA, Wolters V, Durka W (2012) Matrix quality and habitat configuration interactively determine functional connectivity in a widespread bush cricket at a small spatial scale. Landscape Ecol 27(3):381–392
Legendre P, Fortin MJ (2010) Comparison of the Mantel test and alternative approaches for detecting complex multivariate relationships in the spatial analysis of genetic data. Mol Ecol Resour 10(5):831–844
Legendre P, Dale MRT, Fortin MJ, Gurevitch J, Hohn M, Myers D (2002) The consequences of spatial structure for the design and analysis of ecological field surveys. Ecography 25(5):601–615
Li HB, Wu JG (2004) Use and misuse of landscape indices. Landscape Ecol 19(4):389–399
Lowe WH (2010) Explaining long-distance dispersal: effects of dispersal distance on survival and growth in a stream salamander. Ecology 91(10):3008–3015
Mallet J (2001) Gene flow. In: Woiwod IP, Reynolds DR, Thomas CD (eds) Insect movement: mechanisms and consequences. CAB International, Wallingford, pp 337–360
Mapelli FJ, Mora MS, Mirol PM, Kittlein MJ (2012) Population structure and landscape genetics in the endangered subterranean rodent Ctenomys porteousi. Conserv Genet 13(1):165–181
McGarigal K, Cushman SA, Neel MC, Ene E (2002) FRAGSTATS: spatial pattern analysis program for categorical maps. University of Massachusetts, Amherst. http://www.umass.edu/landeco/research/fragstats/fragstats.html
McRae BH (2006) Isolation by resistance. Evolution 60(8):1551–1561
McRae BH, Beier P (2007) Circuit theory predicts gene flow in plant and animal populations. Proc Natl Acad Sci USA 104(50):19885–19890
Millette KL, Keyghobadi N (2015) The relative influence of habitat amount and configuration on genetic structure across multiple spatial scales. Ecol Evol 5(1):73–86
Murphy MA, Evans JS, Cushman SA, Storfer A (2008) Representing genetic variation as continuous surfaces: an approach for identifying spatial dependency in landscape genetic studies. Ecography 31(6):685–697
Murrell DJ, Travis JMJ, Dytham C (2002) The evolution of dispersal distance in spatially-structured populations. Oikos 97(2):229–236
Nathan R (2006) Long-distance dispersal of plants. Science 313(5788):786–788
Neel MC, McGarigal K, Cushman SA (2004) Behavior of class-level landscape metrics across gradients of class aggregation and area. Landscape Ecol 19(4):435–455
Nichols RA, Hewitt GM (1994) The genetic consequences of long-distance dispersal during colonization. Heredity 72:312–317
Okubo A (1980) Diffusion and ecological problems: mathematical models. Springer, New York
Peakall R, Ruibal M, Lindenmayer DB (2003) Spatial autocorrelation analysis offers new insights into gene flow in the Austrailian bush rat Rattus fuscipes. Evolution 57(5):1182–1195
Pope SE, Fahrig L, Merriam NG (2000) Landscape complementation and metapopulation effects on leopard frog populations. Ecology 81(9):2498–2508
R Development Core Team (2012) R: a language and environment for statistical computing. R Foundation for Statistical Computing. Vienna, Austria. http://www.R-project.org
Reed JM, Dobson AP (1993) Behavioural constraints and conservation biology—conspecific attraction and recruitment. Trends Ecol Evol 8(7):253–256
Riley SPD, Pollinger JP, Sauvajot RM, York EC, Bromley C, Fuller TK, Wayne RK (2006) A southern California freeway is a physical and social barrier to gene flow in carnivores. Mol Ecol 15(7):1733–1741
Saupe D (1988) Algorithms for random fractals. In: Peitgen H-O, Saupe D (eds) The science of fractal images. Springer-Verlag, New York, pp 71–113
Schmidt T, Arens P, Smulders MJM, Billeter R, Liira J, Augenstein I, Durka W (2009) Effects of landscape structure on genetic diversity of Geum urbanum L. populations in agricultural landscapes. Flora 204(7):549–559
Schumaker NH (1996) Using landscape indices to predict habitat connectivity. Ecology 77(4):1210–1225
Schwartz M, McKelvey K (2009) Why sampling scheme matters: the effect of sampling scheme on landscape genetic results. Conserv Genet 10(2):441–452
Shirk AJ, Cushman SA (2011) sGD: software for estimating spatially explicit indices of genetic diversity. Mol Ecol Resour 11(5):922–934
Shirk AJ, Cushman SA (2014) Spatially-explicit estimation of Wright’s neighborhood size in continuous populations. Front Ecol Evol 2(62):1–12
Slatkin M (1987) Gene flow and the geographic structure of natural populations. Science 236(4803):787–792
Stow AJ, Sunnucks P, Briscoe DA, Gardner MG (2001) The impact of habitat fragmentation on dispersal of Cunningham’s skink (Egernia cunninghami): evidence from allelic and genotypic analyses of microsatellites. Mol Ecol 10(4):867–878
Telles MPD, Dobrovolski R, Souza KDE, Lima JD, Collevatti RG, Soares TN, Chaves LJ, Diniz JAF (2014) Disentangling landscape effects on population genetic structure of a Neotropical savanna tree. Nat Conserv 12(1):65–70
Templeton AR, Shaw K, Routman E, Davis SK (1990) The genetic consequences of habitat fragmentation. Ann Mo Bot Gard 77(1):13–27
Turchin P (1998) Quantitative analysis of movement: measuring and modeling population redistribution in animals and plants. Sinauer Associates Inc, Sunderland
van Strien MJ, Keller D, Holderegger R, Ghazoul J, Kienast F, Bolliger J (2014) Landscape genetics as a tool for conservation planning: predicting the effects of landscape change on gene flow. Ecol Appl 24(2):327–339
Varvio SL, Chakraborty R, Nei M (1986) Genetic variation in subdivided populations and conservation genetics. Heredity 57:189–198
Weckworth BV, Musiani M, DeCesare NJ, McDevitt AD, Hebblewhite M, Mariani S (2013) Preferred habitat and effective population size drive landscape genetic patterns in an endangered species. Proc R Soc B 280(1769):20131756
With KA, King AW (1999) Extinction thresholds for species in fractal landscapes. Conserv Biol 13(2):314–326
Wright S (1931) Evolution in Mendelian populations. Genetics 16:97–159
Wright S (1969) Evolution and the genetics of populations, vol. 2. The theory of gene frequencies. University of Chicago Press, Chicago
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418
Acknowledgments
We thank five reviewers for very helpful comments on a previous version of this paper. Funding was provided by a Natural Sciences and Engineering Research Council of Canada (NSERC) grant to L. Fahrig.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jackson, N.D., Fahrig, L. Habitat amount, not habitat configuration, best predicts population genetic structure in fragmented landscapes. Landscape Ecol 31, 951–968 (2016). https://doi.org/10.1007/s10980-015-0313-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-015-0313-2