Abstract
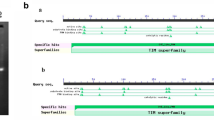
Aldo–keto reductase (AKR) is an enzyme superfamily whose members are involved in the metabolism of aldehydes/ketones. The AKR4 subfamily C (AKR4C) is a group of aldo–keto reductases that are found in plants. Some AKR4C(s) in dicot plants are capable of metabolizing reactive aldehydes whereas, such activities have not been reported for AKR4C(s) from monocot species. In this study, we have screened Indica rice genome for genes with significant homology to dicot AKR4C(s) and identified a cluster of putative AKR4C(s) located on the Indica rice chromosome I. The genes including OsI_04426, OsI_04428 and OsI_04429 were successfully cloned and sequenced by qRT-PCR from leaves of Thai Jasmine rice (KDML105). OsI_04428, later named AKR4C14, was chosen for further studies because it shares highest homology to the dicot AKR4C(s). The bacterially expressed recombinant protein of AKR4C14 was successfully produced as a MBP fusion protein and his-tagged protein. The recombinant AKR4C14 were capable of metabolizing sugars and reactive aldehydes i.e. methylglyoxal, a toxic by-product of the glycolysis pathway, glutaraldehyde, and trans-2-hexenal, a natural reactive 2-alkenal. AKR4C14 was highly expressed in green tissues, i.e. leaf sheets and stems, whereas flowers and roots had a significantly lower level of expression. These findings indicated that monocot AKR4C(s) can metabolize reactive aldehydes like the dicot AKR4C(s) and possibly play a role in detoxification mechanism of reactive aldehydes.




Similar content being viewed by others
Abbreviations
- AKR:
-
Aldo-keto reductase
- MBP:
-
Maltose-binding protein
- HNE:
-
trans-4-hydroxy-2-nonenal
- CDS:
-
Coding sequence
- DTT:
-
Dithiothreitol
- IPTG:
-
Isopropyl-β-D-thio-galactoside
- EDTA:
-
Ethylenediaminetetraacetic acid
- GST:
-
Glutathione S-transferase
References
Aida K, Tawata M, Ikegishi Y, Onaya T (1999) Endocrinology 140(2):609–617
Bagnasco SM, Uchida S, Balaban RS, Kador PF, Burg MB (1987) Proc Natl Acad Sci USA 84(6):1718–1720
Bartels D, Engelhardt K, Roncarati R, Schneider K, Rotter M, Salamini F (1991) EMBO J 10(5):1037–1043
Bladier C, Carrier P, Chagvardieff P (1994) Plant Physiol 106(3):941–947
Burge C, Karlin S (1997) J Molecular Biol 268(1):78–94. doi:10.1006/jmbi.1997.0951
Chehab EW, Raman G, Walley JW, Perea JV, Banu G, Theg S, Dehesh K (2006) Plant Physiol 141(1):121–134. doi:10.1104/pp.106.078592
de Sousa SM, Rosselli LK, Kiyota E, da Silva JC, Souza GH, Peroni LA, Stach-Machado DR, Eberlin MN, Souza AP, Koch KE, Arruda P, Torriani IL, Yunes JA (2009) Plant Physiol Biochem 47(2):98–104
Desmond JC, Mountford JC, Drayson MT, Walker EA, Hewison M, Ride JP, Luong QT, Hayden RE, Vanin EF, Bunce CM (2003) Cancer Res 63(2):505–512
Di Luccio E, Elling RA, Wilson DK (2006) Biochem J 400(1):105–114
Ellis EM, Judah DJ, Neal GE, Hayes JD (1993) Proc Natl Acad Sci USA 90(21):10350–10354
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein Identification and Analysis Tools on the ExPASyServer. In: Walker JM (ed) The Proteomics Protocols Handbook. Humana Press, Totowa, NJ
Gavidia I, Perez-Bermudez P, Seitz HU (2002) Eur J Biochem 269(12):2842–2850
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Adv Bioinform 2008:420747. doi:10.1155/2008/420747
Hyndman D, Bauman DR, Heredia VV, Penning TM (2003) Chem Biol Interact 143–144:621–631
Ireland LS, Harrison DJ, Neal GE, Hayes JD (1998) Biochem J 332(Pt 1):21–34
Ishikura S, Horie K, Sanai M, Matsumoto K, Hara A (2005) Biol Pharm Bull 28(6):1075–1078
Jez JM, Bennett MJ, Schlegel BP, Lewis M, Penning TM (1997) Biochem J 326(Pt 3):625–636
Jez JM, Flynn TG, Penning TM (1997) Biochem Pharmacol 54(6):639–647
Kapust RB, Waugh DS (1999) Protein Sci 8(8):1668–1674
Kolb NS, Hunsaker LA, Vander Jagt DL (1994) Mol Pharmacol 45(4):797–801
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Bioinformatics 23(21):2947–2948. doi:10.1093/bioinformatics/btm404
Lee SP, Chen TH (1993) Plant Physiol 101(3):1089–1096
Li B, Foley ME (1995) Plant Mol Biol 29(4):823–831
Mano J, Miyatake F, Hiraoka E, Tamoi M (2009) Planta 230(4):639–648. doi:10.1007/s00425-009-0964-9
Mundree SG, Whittaker A, Thomson JA, Farrant JM (2000) Planta 211(5):693–700
Oberschall A, Deak M, Torok K, Sass L, Vass I, Kovacs I, Feher A, Dudits D, Horvath GV (2000) Plant J 24(4):437–446
Penning TM, Burczynski ME, Jez JM, Hung CF, Lin HK, Ma H, Moore M, Palackal N, Ratnam K (2000) Biochem J 351(Pt 1):67–77
Petrash JM (2004) Cell Mol Life Sci 61(7–8):737–749
Roncarati R, Salamini F, Bartels D (1995) Plant J 7(5):809–822
Simpson PJ, Tantitadapitak C, Reed AM, Mather OC, Bunce CM, White SA, Ride JP (2009) J Mol Biol 392(2):465–480
Srivastava S, Chandra A, Bhatnagar A, Srivastava SK, Ansari NH (1995) Biochem Biophys Res Commun 217(3):741–746
Srivastava S, Dixit BL, Cai J, Sharma S, Hurst HE, Bhatnagar A, Srivastava SK (2000) Free Radic Biol Med 29(7):642–651
Turoczy Z, Kis P, Torok K, Cserhati M, Lendvai A, Dudits D, Horvath GV (2011) Plant Mol Biol 75(4–5):399–412. doi:10.1007/s11103-011-9735-7
Vander Jagt DL, Robinson B, Taylor KK, Hunsaker LA (1992) J Biol Chem 267(7):4364–4369
Vander Jagt DL, Hassebrook RK, Hunsaker LA, Brown WM, Royer RE (2001) Chem Biol Interact 130–132(1–3):549–562
Weber J, Kayser A, Rinas U (2005) Microbiology 151(Pt 3):707–716
Welle R, Schroder G, Schiltz E, Grisebach H, Schroder J (1991) Eur J Biochem 196(2):423–430
Yadav SK, Singla-Pareek SL, Ray M, Reddy MK, Sopory SK (2005) Biochem Biophys Res Commun 337(1):61–67
Acknowledgments
We thank Dr. Jon P. Ride (University of Birmingham) and Prof. Pradit Pongtongkam (Kasetsart University) for fruitful discussion. This work was funded by the Kasetsart University Research and Development Institute (KURDI), and the Faculty of Science, Kasetsart University.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Narawongsanont, R., Kabinpong, S., Auiyawong, B. et al. Cloning and Characterization of AKR4C14, a Rice Aldo–Keto Reductase, from Thai Jasmine Rice. Protein J 31, 35–42 (2012). https://doi.org/10.1007/s10930-011-9371-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-011-9371-8