Abstract
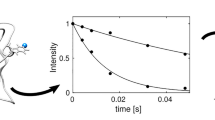
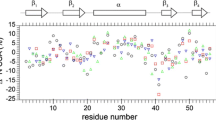
Recently a suite of six CPMG relaxation dispersion experiments has been described for quantifying millisecond time-scale exchange processes in proteins. The methodology has been applied to study the folding reaction of a G48M Fyn SH3 domain mutant that exchanges between the native state, and low populated unfolded and intermediate states. A complex non-linear global optimization protocol allows extraction of the kinetics and thermodynamics of the 3-site exchange process from the experimental data, as well as reconstruction of the amide group chemical shifts of the excited states. We show here, through a series of Monte-Carlo simulations on various synthetic data sets, that the 3-site exchange parameters extracted for this system on the basis of 15N single-quantum (SQ) dispersion profiles exclusively, recorded at a single temperature, are significantly in error. While a temperature dependent 15N study improves the robustness of extracted parameters, as does a combined analysis of 15N and 1H SQ data sets measured at a single temperature, the best agreement is observed in cases where the full complement of six dispersion profiles per residue is analyzed.
Similar content being viewed by others
References
M.J. Grey C.Y. Wang A.G. Palmer (2003) J. Am. Chem. Soc. 125 14324–14335 Occurrence Handle10.1021/ja0367389
R. Ishima D. Torchia (2003) J. Biomol. NMR 25 243–248 Occurrence Handle10.1023/A:1022851228405
D.M. Korzhnev K. Kloiber L.E. Kay (2004a) J. Am. Chem. Soc. 126 7320–7329
D.M. Korzhnev P. Neudecker A. Mittermaier V.Y. Orekhov L.E. Kay (2005) J. Am. Chem. Soc. 127 15602–15611
D.M. Korzhnev X. Salvatella M. Vendruscolo A.A. Di Nardo A.R. Davidson C.M. Dobson L.E. Kay (2004b) Nature 430 586–590 Occurrence Handle2004Natur.430..586K Occurrence Handle10.1038/nature02655
J.P. Loria M. Rance A.G. Palmer (1999) J. Am. Chem. Soc. 121 2331–2332 Occurrence Handle10.1021/ja983961a
A. Mittermaier D.M. Korzhnev L.E. Kay (2005) Biochemistry 44 15430–15436 Occurrence Handle10.1021/bi051771o
V.Y. Orekhov D.M. Korzhnev L.E. Kay (2004) J. Am. Chem. Soc. 126 1886–1891
A.G. Palmer Suffix3rd M.J. Grey C. Wang (2005) Methods Enzymol 394 430–465
A.G. Palmer C.D. Kroenke J.P. Loria (2001) Methods Enzymol. 339 204–238
W.H. Press B.P. Flannery S.A. Teukolsky W.T. Vetterling (1988) Numerical Recipes in C Cambridge University Press Cambridge
D. Tolkatchev P. Xu F. Ni (2003) J. Am. Chem. Soc. 125 12432–12442 Occurrence Handle10.1021/ja021238l
M. Tollinger N.R. Skrynnikov F.A.A. Mulder J.D. Forman-Kay L.E. Kay (2001) J. Am. Chem. Soc. 123 11341–11352 Occurrence Handle10.1021/ja011300z
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Neudecker, P., Korzhnev, D.M. & Kay, L.E. Assessment of the Effects of Increased Relaxation Dispersion Data on the Extraction of 3-site Exchange Parameters Characterizing the Unfolding of an SH3 Domain. J Biomol NMR 34, 129–135 (2006). https://doi.org/10.1007/s10858-006-0001-2
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s10858-006-0001-2