Abstract
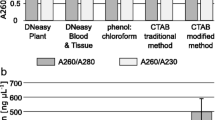
To study the molecular mechanism of the Antarctic sea–ice alga in adaptation to polar sea–ice environments, the RNA was prepared for cDNA library construction of Chlamydomonas sp. ICE-L. Three different methods were tested to prepare total RNA from this psychrophilic, unicellular green alga rich in protein and polysaccharide. Lauryl sodium sulfate- based method allowed a most effective extraction of high-quality total RNA compared to the other methods. Total RNA extracted with this protocol was used for cDNA library construction. The recombination rate of constructed cDNA library was 98.60%, the primary titer was 7.15 × 106 pfu, and an average sequence length was 1.2 kb. These results show that with a high-quality RNA preparation, a cDNA library can be constructed successfully for Chlamydomonas sp. ICE-L.


Similar content being viewed by others
References
Bekesiova I, Nap JP, Mlynarova L (1999) Isolation of high quality DNA and RNA from leaves of the carnivorous plant Drosera rotundifolia. Plant Mol Biol Reptr 17:269–277
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate–phenol–chloroform extraction. Anal Biochem 162:156–159
Dong JZ, Dunstan DI (1996) A reliable method for extraction of RNA from various conifer tissues. Plant Cell Rep 15:516–521
Gasic K, Hernandez A, Korban SS (2004) RNA extraction from different apple tissues rich in polyphenols and polysaccharides for cDNA library construction. Plant Mol Biol Reptr 22:437a–437g
Geuna F, Hartings H, Scienza A (1998) A new method for rapid extraction of high quality RNA from recalcitrant tissues of grapevine. Plant Mol Biol Reptr 16:61–67
Gyeongseo J, Lee CG, Kang SH, Jin E (2007) Annotation and expression profile analysis of cDNAs from the Antarctic diatom Chaetoceros neogracile. J Microbiol Biotechnol 17:1330–1337
Hwang YS, Jung G, Jin E (2008) Transcriptome analysis of acclimatory responses to thermal stress in Antarctic algae. Biochem Biophys Res Commun 367:635–641
Iandolino AB, Silva FGD, Lim H, Choi H, Williams LE, Cook DR (2004) High-quality RNA, cDNA, and derived EST libraries from grapevine (Vitis vinifera L.). Plant Mol Biol Report 22:269–278
Janech MG, Krell A, Mock T, Kang JS, Raymond JA (2006) Ice-binding proteins from sea ice diatoms (Bacillariophyceae). J Phycol 42:410–416
Jung G, Lee CG, Kang SH, Jin E (2007) Annotation and expression profile analysis of cDNAs from the Antarctic diatom Chaetoceros neogracile. J Microbiol Biotechnol 17:1330–1337
Junttila S, Lim KJ, Rudd S (2009) RNA isolation for cDNA library construction from the reindeer lichen Cladonia rangiferina. BMC Research Notes 2:204. doi:10.1186/1756-0500-2-204
Kan GF, Zheng Z, Jiang YH, Miao JL, Zhang BT, Li GY (2006) Salt resistance of Antarctic ice microalga Chlamydomonas sp. Journal of Fishery Sciences of China 13:73–78
Kim SH, Lee YK, Hong CB, Lee IK (1997) A simple method for RNA extraction from marine macroalgae. Algae 12:53–56
Liu CL, Huang XH, Wang XL, Zhang XC, Li GY (2006) Phylogenetic studies on two strains of Antarctic ice algae based on morphological and molecular characteristics. Phycologia 45:190–198
Miao JL, Shi HQ, Jiang YH, Zhang BT, Hou XG (2002) Studies on biochemical composition of Antarctic ice–microalgae and its relation to the cold acclimation. Adv Mar Sci 20:43–50
Miao JL, Kan GF, Li GY (2003) Changes of the Antarctic alga exposed to UV2B in configuration and ultrastructure as well as main biochemical composition. Chinese J Marine Drugs 6:1–5
Mock T, Thomas DN (2005) Recent advances in sea–ice microbiology. Environ Microbiol 7:605–619
Mock T, Krell A, Glockner G, Kolukisaoglu U, Valentin K (2005) Analysis of expressed sequence tags (ESTs) from the polar diatom Fragilariopsis cylindrus. J Phycol 42:78–85
Pinto FL, Thapper A, Sontheim W, Lindblad P (2009) Analysis of current and alternative phenol based RNA extraction methodologies for cyanobacteria. BMC Mol Biol 10:79–86
Provasolil (1968) Media and prospects for the cultivation of marine algae. In: Watanabe A, Hattori A (eds) Cultures and collections of algae. Proceeding of the U.S., Japan Conference, Hakone. Japanese Society of Plant Physiology, pp 63–75
Su X, Gibor A (1988) A method for RNA isolation from marine macro-algae. Anal Biochem 174:650–657
Thanh T, Omar H, Abdullah MP, Chi VTQ, Noroozi M, Ky H, Napis S (2009) Rapid and effective method of RNA isolation from green microalga Ankistrodesmus convolutus. Mol Biotechnol 43:148–153
Vanessa DRF, Angela PT, Mariana CO, Pio C (2008) RNA isolation method for polysaccharide rich algae: agar producing Gracilaria tenuistipitata (Rhodophyta). J Appl Phycol 20:9–12
Yao JT, Fu WD, Wang XL, Duan DL (2009) Improved RNA isolation from Laminaria japonica Aresch (Laminariaceae, Phaeophyta). J Appl Phycol 21:233–238
Zeng Y, Yang T (2002) RNA isolation from high viscous samples rich in polyphenols and polysaccharides. Plant Molecular Reporter 20:417a–417e
Zhang ZD, Shrager J, Jain M, Chang CW, Vallon O, Grossman AR (2004) Insights into the survival of Chlamydomonas reinhardtii during sulfur starvation based on microarray analysis of gene expression. Eukaryotic Cell 3(5):1331–1348
Zhu M, Zhang XC, Mao YX, Yan BL, Teng YJ, Lu ZH (2003) Effects of temperature, salinity and illumination on the growth of Thalassiasira sp. Mar Sci 27(12):58–61
Acknowledgments
This work was financially supported by a grant of National Natural Science Foundation (40876106) for Chenlin Liu and grants of polar Stratagem Foundations (2006 and 2007) for Chenlin Liu and Shenghao Liu.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, G., Liu, C., Liu, S. et al. High-quality RNA preparation for cDNA library construction of the Antarctic sea–ice alga Chlamydomonas sp. ICE-L. J Appl Phycol 22, 779–783 (2010). https://doi.org/10.1007/s10811-010-9519-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-010-9519-5