Abstract
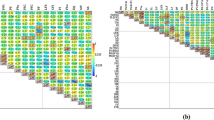
In Tanzania sweet potato ranks as the third most important crop after cassava and potato. We studied the phenotypic diversity of morphological plant and root descriptor traits in accessions of the sweet potato germplasm collection of Sokoine University of Agriculture, Morogoro and Sugarcane Research Institute, Kibaha, Tanzania, using phenotypic characters. A total number of 105 sweet potato accessions of different geographic origins were studied in field trials of The Sugarcane Research Institute at Kibaha Tanzania, and data were recorded for 27 phenotypic characters. Estimates of pair-wise phenotypic similarities using the Manhattan coefficient varied from 0.023 to 0.814, with a mean of 0.285. Cluster analysis was conducted using the unweighted pair group method with arithmetic mean (UPGMA) and Principal Coordinate Analysis (PCO). The clustering of phenotypic data resulted in a dendrogram which was discordant with geographic origin and AFLP data. The analysis of variance (ANOVA) revealed highly significant variation among the accessions for 21 out of the 27 characters studied. Phenotypic analyses revealed a wider range of variability than AFLP analyses. Comparison of molecular and phenotypic data using the Mantel test showed a very low correlation (r 2 = 0.0007). Molecular and phenotypic classifications are discordant, and both are necessary to classify the germplasm correctly and to clarify genetic relationships among sweet potato accessions.


Similar content being viewed by others
References
Abdellaoui R, Rouaissi M, Chenennaoui S, Ben Naceur M, Ben Hmida J, Leila BBK (2007) Genetic diversity of Tunisian local barley accessions analyzed with morphological and RAPD makers: relationship between the two methods. Kor J Genet 29:315–322
AGROBASE (2001) AGROBASE Generation II, Users manual. Agronomix Software, Inc., Winnipeg
Baker RH, Yu XB, DeSalle R (1998) Assessing the relative contribution of molecular and morphological characters in simultaneous analysis trees. Mol Phylogenet Evol 9:427–436
Campos ET, Espinosa MAG, Warburton ML, Varela AS, Monter AV (2005) Characterization of mandarin (Citrus spp.) using morphological and AFLP markers. Interciencia 30:687–693
CIP (1999) Annual report. International Potato Center, Lima
Crespan M (2004) Evidence on the evolution of polymorphism of microsatellite markers in varieties of Vitis vinifera L. Theor Appl Genet 108:231–237
Diaz JP, Schmiediche P, Austin DF (1996) Polygon of crossability between eleven species of Ipomoea: section Batatas (Convolvulaceae). Euphytica 88:189–200
Elameen A, Fjellheim S, Larsen A, Rognli OA, Sundheim L, Msolla E, Masumba S, Mtunda K, Klemsdal SS (2008) Analysis of genetic diversity in a sweet potato (Ipomoea batatas L.) germplasm collection from Tanzania as revealed by AFLP. Genet Resour Crop Evol 55:397–408
Fajardo S, La Bonte DR, Jarret RL (2002) Identifying and selecting genetic diversity in Papua New Guinea sweet potato Ipomoea batatas (L.) Lam. Germplasm collected as botanical seed. Genet Resour Crop Evol 49:463–470
FAO, Production Yearbook (2004) FAO Statistic Series No 134. Rome, Italy
Gepts P (1993) The use of molecular and biochemical markers in crop evolution studies. Evol Biol 27:51–94
Green RC (2005) Sweet potato transfers in Polynesian prehistory. In: Ballard C, Brown P, Bourke RM, Harwood T (eds) The sweet potato in Oceania: a reappraisal. The University of Sydney, Sydney, pp 43–62
Halliburton R (2004) Introduction to population genetics. Upper Saddle River, Pearson Prentice hall, pp 231–333
Harlan JR (1992) Crops and man, 2nd edn. American Society of Agronomy and Crop Science Society of America, Madison, p 12
He GH, Prakash CS, Jarret RL (1995) Analysis of genetic diversity in sweet potato (Ipomoea batatas) germplasm collection using DNA amplification fingerprinting. Genome 38:938–945
Huamán Z (1991) Descriptors for sweet potato. CIP, AVRDC, IBPGR, Rome, pp 43–44
Huamán Z (1992) Morphologic identification of duplicates in collections of Ipomoea batatas. CIP Research Guide 36. International Potato Centre, Lima, Peru
Huamán Z, Aguilar C, Ortiz R (1999) Selecting a Peruvian sweet potato core collection on the basis of morphological, eco-geographical, and disease and pest reaction data. Theor Appl Genet 98:840–844
Hutchings MJ, de Kroon H (1994) Foraging in plants: the role of morphological plasticity in resource acquisition. Adv Ecol Res 25:159–238
Jones A, Dukes PD, Schalk JM (1986) Sweet potato breeding. In: Basset MJ (ed) Breeding vegetable crops, Av Publishing C, USA, pp 1–35
Kuo CG (1991) Conservation and distribution of sweet potato germplasm. In: Dodds JH (ed) In vitro methods for conservation of plant genetic resources. Chapman and Hall, New York, pp 123–147
Lin KH, Lai YC, Chang KY, Chen YF, Hwang YS, Lo HF (2007) Improving breeding efficiency for quality and yield of sweet potato. Bot Stud 48:283–292
Magoon ML, Krishnan R, Bai KV (1970) Cytological evidence in the origin of sweet potato. Theor Appl Genet 40:360–366
Mantel NA (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Naskar SK (1996) Genetic divergence for yield contributing traits in sweet potato (Ipomoea batatas). In: Kurup GT, Palaniswami MS, Potty VP, Padmaja G, Labeerathumma S, Pillai SV (eds) Tropical tuber crops: problems, prospects and future strategies. Science Publishers, New Hampshire, pp 133–136
NRC (National Research Council) (1993) Agricultural crop issues and policies: managing global genetic resources, vol I. National Research Council (USA), Washington DC, p 123
Ozias-Akins P, Jarret RL (1994) Nuclear DNA content and ploidy levels in the genus Ipomoea. J Am Soc Hort Sci 119:110–115
Price EAC, Marshall C (1999) Clonal plants and environmental heterogeneity: an introduction to the proceedings. Plant Ecol 141:3–7
Rao VR, Debouck T, Iwanga M (1994) The role of international organisations in root and tuber crop conservation. In: The first ministry of agriculture, forestry and fisheries. International workshop on genetic resources—root and tuber crops, 15–17 March 1994, Tsukuba, MAFF, Japan, pp 7–22
Rohlf FJ (2000) NTSYS-pc, numerical taxonomy and multivariate analysis system, v.2.02. Exeter software, New York
Roldan-Ruiz I, Van euwijk FA, Gilliland TJ, Dubreuil P, Dillmann C, Lallemand J, De Loose M, Baril CP (2001) A comparative study of molecular and morphological methods of describing relationships between perennial ryegrass (Lolium perenne L.) varieties. Theor Appl Genet 103:1138–1150
Sagredo B, Hinrichsen P, López H, Cubillos A, Munoz C (1998) Genetic variation of sweet potatoes (Ipomoea batatas L.) cultivated in Chile determined by RAPDs. Euphytica 101:193–198
SAS (2002–2003) SAS for Windows Release 9.1, SAS Institute, Inc., Cary
Sihachakr D, Ducreux G (1987) Plant regeneration from protoplast culture of sweet potato (Ipomoea batatas Lam.). Plant Cell Rep 6:326–328
Sneath PA, Sokal RR (1973) Numerical taxonomy. Freeman, San Francisco, p 573
Solis JS, Ulloa DM, Rodriguez LA (2007) Molecular description and similarity relationships among native germplasm potatoes (Solanum tuberosum ssp. tuberosum L.) using morphological data and AFLP markers. Electronic J Biotech 10:436–443
Spooner DM, Mclean K, Ramsay G, Waugh R, Bryan GJ (2005) Single domestication for potato based on multilocus amplified fragment length polymorphism genotyping. PNAS 102:14694–14699
Tairo F, Mneney E, Kullaya A (2008) Morphological and agronomical characterization of sweet potato (Ipomoea batatas Lam.) germplasm collection from Tanzania. Afr J Plant Sci 8:77–85
Tsegaye E, Sastry D, Dechassa N (2007) Genetic variability for the yield and other agronomic traits in sweet potato. Indian J Hort 64:237–240
van Kleunen M, Fischer M, Schmid B (2002) Experimental life-history evolution: selection on the allocation to sexual reproduction and its plasticity in a clonal plant. Evolution 56:2168–2177
Vaughan JG, Geissler CA (1997) The new Oxford book of food plants. Oxford University Press, Oxford
Veasey EA, Silva JSD, Rosa MS, Borges A, Bressan ED, Peroni N (2007) Phenology and morphological diversity of sweet potato (Ipomoea batatas) landraces of the Vale do Ribeira. Scientia Agricola 64:416–427
Veteloainen M, Gammelga E, Valkonen JPT (2005) Diversity of Nordic landrace potatoes (Solanum tuberosum L.) revealed by AFLPs and morphological characters. Genet Resour Crop Evol 52:999–1010
Wachira FN, Waugh R, Hackett CA, Powell W (1995) Detection of genetic diversity in tea (Camellia sinensis) using RAPD markers. Genome 38:201–210
Wendel JF, Doyle JJ (1998) Phylogenetic incongruence: window into genome history and molecular evolution. In: Soltis DE, Soltis PS, Doyle JJ (eds) Molecular systematics of plants II: DNA sequencing. Kluwer Academic Publishers, Boston, pp 265–296
Zdunic G, Pejic I, Kontic JK, Vukicevic D, Vokurka A, Pezo I, Maletic E (2008) Comparison of genetic and morphological data for inferring similarity among native Dalmatian (Croatia) grapevine cultivars (Vitis vinifera L.). J Food Agric Environ 6:333–336
Zhang D, Cervantes J, Huaman EC, Ghislain M (2000) Assessing genetic diversity of sweet potato (Ipomoea batatas (L.) Lam.) cultivars from tropical America using AFLP. Genet Resour Crop Evol 47:659–665
Acknowledgments
The authors thank the staff at the Germplasm Centre of The Sugarcane Research Institute, Kibaha, and at Sokoine University of Agriculture, Morogoro, Tanzania, for their cooperation and for supplying us with the phenotypic data of sweet potato accessions used in the study. This work has been partly supported by NORAD (Norwegian Agency for Development Cooperation, project 021 TARPII-SUA).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Elameen, A., Larsen, A., Klemsdal, S.S. et al. Phenotypic diversity of plant morphological and root descriptor traits within a sweet potato, Ipomoea batatas (L.) Lam., germplasm collection from Tanzania. Genet Resour Crop Evol 58, 397–407 (2011). https://doi.org/10.1007/s10722-010-9585-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-010-9585-1