Abstract
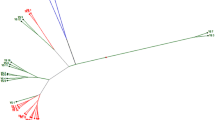
Mustard (Brassica juncea) is an important crop in both ancient and modern world. It has a broad resource of genetic diversity that is used primarily as oilseed but as vegetables, condiment and medicines also. Its superior tolerance to adverse environments, e.g., drought, high temperature and low fertility suggests its better adaptability in future possible harsh environments. Chinese vegetable mustard displays a wide spectrum of morphotypes. A collection of 95 accessions of B. juncea representing oil and vegetable mustards from China, France, India, Pakistan, and Japan were assessed to determine diversity at the molecular level using sequence-related amplified polymorphism (SRAP). Eight SRAP primer combinations identified a total of 326 scorable fragments of which 161 were polymorphic (49.39%). The percentage of polymorphism for each primer combination varied from 21.88 to 66.67%. Both Shannon-Weaver and Simpson genetic diversity index indicated that the level of genetic diversity within vegetable mustard is much higher than within oil mustard, and also winter oil mustards are genetically more diverse than spring oil mustards. Based on the Cluster and Principal Coordinates analysis, which were conducted on the similarity matrix of SRAP marker data, vegetable, spring oil and winter oil mustard were clearly divided into three distinct groups and among these three groups, spring and winter oil mustard are geneticlly closer than vegetable mustard. This suggests that bilateral gene exchange between oil and vegetable gene pools in the breeding program will effectively elevate the genetic potential in developing higher yields, more disease resistance, better quality and better adapted lines.


Similar content being viewed by others
References
Becker HC, Engqvist GM, Karlsson B (1995) Comparison of rapeseed cultivars and resynthesized lines based on allozyme and RFLP markers. Theor Appl Genet 91:62–67. doi:10.1007/BF00220859
Burkill IH (1930) The Chinese mustards in the Malay Peninsula. Gardners Bull 5:99–117
Burton WA, Salisbury PA, Potts D (2003) The potential of canola quality Brassica juncea as an oilseed crop for Australia. In: Proceedings of the 11th International Rapeseed Congress, Copenhagen, 6–10 July 2003, pp 5–7
Burton WA, Ripley VL, Potts DA, Salisbury PA (2004) Assessment of genetic diversity in selected breeding lines and cultivars of canola quality Brassica juncea and their implications for canola breeding. Euphytica 136:181–192. doi:10.1023/B:EUPH.0000030672.56206.f0
Erickson LR, Straus NA, Beversdorf WD (1983) Restriction patterns reveal origins of chloroplast genomes in Brassica amphidiploids. Theor Appl Genet 65:201–206. doi:10.1007/BF00308066
Ferriol M, Pico B, Nuez F (2003) Genetic diversity of a germplasm collection of Cucurbita pepo using SRAP and AFLP markers. Theor Appl Genet 107:271–282. doi:10.1007/s00122-003-1242-z
Fu J, Zhang MF, Qi XH (2006) Genetic diversity of traditional Chinese mustard crops Brassica juncea as revealed by phenotypic difference and RAPD markers. Genet Resour Crop Evol 53:1513–1519. doi:10.1007/s10722-005-7763-3
Gladis T, Hammer K (1992) The Brassica collection in Gatersleben: Brassica juncea, Brassica napus, Brassica nigra, and Brassica rapa (in German with English summary). Feddes Repert 103:469–507
Gómez-Campo C, Prakash S (1999) Origin and domestication. In: Gómez-Campo C (ed) Biology of Brassica coenospecies. Elsevier, Amsterdam, pp 59–106
Hinata K, Prakash S (1984) Ethnobotany and evolutionary origin of Indian oleiferous Brassicae. Indian J Genet 44:102–112
Institute of Archaeology of Chinese Academy of Sciences (1963) Xian Banpo country. Special issue of Archaeology, Archaeology Press, Beijing, pp 223
Jain A, Bhatia S, Banga SS, Prakash S, Lakshmikumaran M (1994) Potential use of random amplified polymorphic DNA (RAPD) technique to study the genetic diversity in Indian mustard (Brassica juncea) and its relationship to heterosis. Theor Appl Genet 88:116–122. doi:10.1007/BF00222403
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new markers system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461. doi:10.1007/s001220100570
Lu GY, Yang GS, Fu TD (2004) Molecular mapping of a dominant genic male sterility gene Ms in rapeseed (Brassica napus). Plant Breed 123:262–265. doi:10.1111/j.1439-0523.2004.00957.x
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273. doi:10.1073/pnas.76.10.5269
Palmer JD, Shields CR, Cohen DB, Orton TJ (1983) Chloroplast DNA evolution and the origin of amphidiploid Brassica species. Theor Appl Genet 65:181–189. doi:10.1007/BF00308062
Potts DJ, Rakow GW, Males DR (1999) Canola-quality Brassica juncea, a new oilseed crop for the Canadian praries. In: Proceedings of the 10th International Rapeseed Congress, Canberra, CD-ROM
Pradhan AK, Sodhi YS, Mukhopadhyay A, Pental D (1993) Heterosis breeding in Indian mustard (Brassica juncea L. Czern & Cross): analysis of component characters contributing to heterosis for yield. Euphytica 69:219–229. doi:10.1007/BF00022368
Prain D (1898) The mustards cultivated in Bengal. Agr Ledger 5:1–80
Prakash S, Bhat SR, Quiros CF, Kirti PB, Chopra VL (2009) Brassica and its close allies: cytogenetics and evolution. Plant Breed Rev 31:21–187
Qi XH, Zhang MF, Yang JH (2007) Molecular phylogeny of Chinese vegetable mustard (Brassica juncea) based on the internal transcribed spacers (ITS) of nuclear ribosomal DNA. Genet Resour Crop Evol 54:1709–1716. doi:10.1007/s10722-006-9179-0
Qi XH, Yang JH, Zhang MF (2008) AFLP-based genetic diversity assessment among Chinese vegetable mustards (Brassica juncea (L.) Czern). Genet Resour Crop Evol 55:705–711
Seyis F, Friedt W, Lühs W (2006) Yield of Brassica napus L. hybrids developed using resynthesized rapeseed material sown at different locations. Field Crops Res 96:176–180. doi:10.1016/j.fcr.2005.06.005
Sinskaia EN (1928) The oleiferous plants and root crops of the family Cruciferae. Bull Appl Bot Genet Plant Breed 19:1–648
Song KM, Osborn TC, Williams PH (1988) Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 1. Genome evolution of diploid and amphidiploid species. Theor Appl Genet 75:784–794. doi:10.1007/BF00265606
Srivastava A, Gupta V, Pental D, Pradhan AK (2001) AFLP-based genetic diversity assessment amongst agronomically important natural and some newly synthesized lines of Brassica juncea. Theor Appl Genet 102:193–199. doi:10.1007/s001220051635
Sun VG (1970) Breeding plants of Brassica. J Agric Assoc China 71:41–52
U N (1935) Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J Bot 7:389–452
Uchimiya H, Wildman SG (1978) Evolution of fraction I protein in relation to origin of amphidiploid Brassica species and other member of Cruciferae. J Hered 69:299–303
Vavilov NI (1949) The origin, variation, immunity and breeding of cultivated plants. Chron Bot 13:1–364
Woods DL, Capcara JJ, Downey RK (1991) The potential of mustard (Brassica juncea (L.) Coss.) as an edible oil crop on the Canadian Prairies. Can J Plant Sci 71:195–198
Yang YG, Zhou GF, Chen XQ, Chen CL, Zhou GF, Liao KL (1989) A study on classification of mustard. Acta Hortic Sin 16:114–121
Zhao JJ, Wang XW, Deng B, Lou P, Wu J, Sun RF (2005) Genetic relationships within Brassica rapa as inferred from AFLP fingerprints. Theor Appl Genet 110:1301–1314. doi:10.1007/s00122-005-1967-y
Acknowledgements
The authors are very grateful to Prof. Shyam Prakash for critical reading the manuscript and providing valuable comments. We thank Prof. Li Xixiang, Institute for Vegetable and Flowers (IVF) and The Germplasm Resources Information Network (GRIN) for providing seeds of vegetable mustards and their passport data. This project is sponsored by 863-Hi-techresearch and development program of China (2006AA10Z1E4) and National Natural Science Foundation of China (39830270).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, Xm., Chen, By., Lu, G. et al. Genetic diversity in oil and vegetable mustard (Brassica juncea) landraces revealed by SRAP markers. Genet Resour Crop Evol 56, 1011–1022 (2009). https://doi.org/10.1007/s10722-009-9420-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-009-9420-8