Abstract
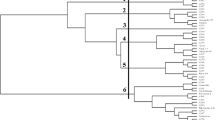
The cultivated diploid, Gossypium arboreum L., (A genome) is an invaluable genetic resource for improving modern tetraploid cotton (G. hirsutum L. and G. barbadense L.) cultivars. The objective of this research is to select a set of informative and robust microsatellites for studying genetic relationships among accessions of geographically diverse G. arboreum cultivars. From more than 1,500 previously developed simple sequence repeat (SSR) markers, 115 genomic (BNL) and EST-derived (MUCS and MUSS) markers were used to evaluate the allelic diversity of a core panel of G. arboreum accessions. These SSR data enabled advanced genome analyses. A set of 25 SSRs were selected based both upon their high level of informativeness (PIC ≥ 0.50) and the production of clear PCR bands on agarose gels. Subsequently, 96 accessions representing a wide spectrum of diversity of G. arboreum cultivars were analyzed with these markers. The 25 SSR loci revealed 75 allelic variants (polymorphisms) ranging from 2 to 4 alleles per locus. The Neighborjoining (NJ) method, based on genetic dissimilarities, revealed that cultivars from geographically adjacent countries tend to cluster together. Outcomes of this research should be useful in decreasing redundancy of effort and in constructing a core collection of G. arboreum, important for efficient use of this genetic resource in cotton breeding.


Similar content being viewed by others
References
Abdalla AM, Reddy OUK, El-Zik KM, Pepper AE (2001) Genetic diversity and relationships of diploid and tetraploid cottons revealed using AFLP. Theor Appl Genet 102:222–229. doi:10.1007/s001220051639
Alvarez I, Wendel JF (2006) Cryptic interspecific introgression and genetic differentiation within Gossypium aridum (Malvaceae) and its relatives. Evol Int J Org Evol 60:505–517
Anderson JA, Churchill GA, Autrique JE, Tanksley SD, Sorrells ME (1993) Optimizing parental selection for genetic linkage maps. Genome 36:181–186. doi:10.1139/g93-024
Bowman DT (2000) Attributes of public and private cotton breeding programs. J Cotton Sci 4:130–136
Fryxell PA (1992) A revised taxonomic interpretation of Gossypium L. (Malvaceae). Rheedea 2:108–165
Fryxell PA, Craven LA, Stewart JMD (1992) A revision of Gossypium sect. Grandicalyx (Malvaceae), including the description of six new species. Syst Bot 17:91–114. doi:10.2307/2419068
Guo WZ, Wang K, Zhang TZ (2003) A and D genome evolution in Gossypium revealed using SSR molecular markers. Acta Genet Sin 30:183–188
Gupta PK, Varshney RK (2000) The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica 113:163–185. doi:10.1023/A:1003910819967
Iqbal MJ, Aziz N, Saeed NA, Zafar Y, Malik KA (1997) Genetic diversity evaluation of some elite cotton varieties by RAPD-analysis. Theor Appl Genet 94:139–144. doi:10.1007/s001220050392
Iqbal MJ, Reddy OUK, El-Zik KM, Pepper AE (2001) A genetic bottleneck in the evolution under domestication of upland cotton Gossypium hirsutum L. examined using DNA fingerprinting. Theor Appl Genet 103:547–554. doi:10.1007/PL00002908
Jain S, Jain RK, McCouch S (2004) Genetic analysis of Indian aromatic and quality rice (Oryza sativa L.) germplasm using panels of fluorescently-labeled microsatellite markers. Theor Appl Genet 109:965–977. doi:10.1007/s00122-004-1700-2
Khan SA, Hussain D, Askari E, Stewart JMD, Malik KA, Zafar Y (2000) Molecular phylogeny of Gossypium species by DNA fingerprinting. Theor Appl Genet 101:931–938. doi:10.1007/s001220051564
Lacape M, Dessauw D, Rajab M, Noyer JL, Hau B (2007) Microsatellite diversity in tetraploid Gossypium germplasm : assembling a highly informative genotyping set of cotton SSRs. Mol Breed 19:45–58. doi:10.1007/s11032-006-9042-1
Liu S, Saha S, Stelly D, Burr B, Cantrell RG (2000a) Chromosomal assignment of microsatellite loci in cotton. J Hered 91(4):326–332. doi:10.1093/jhered/91.4.326
Liu S, Cantrell RG, McCarty JCJ, Stewart JMD (2000b) Simple sequence repeat-based assessment of genetic diversity in cotton race stock accessions. Crop Sci 40:1459–1469
Liu D, Guo X, Lin Z, Nie Y, Zhang X (2006) Genetic diversity of Asian cotton (Gossypium arboreum L.) in China evaluated by microsatellite analysis. Genet Resour Crop Evol 53:1145–1152. doi:10.1007/s10722-005-1304-y
Lu HJ, Myers GO (2002) Genetic relationships and discrimination of ten influential Upland cotton varieties using RAPD markers. Theor Appl Genet 105:325–331. doi:10.1007/s00122-002-0947-8
Macaulay M, Ramsay L, Powell W, Waugh R (2001) A representative, highly informative “genotyping set” of barley SSRs. Theor Appl Genet 106:801–809. doi:10.1007/s001220000487
Masi P, Spagnoletti Zeuli PL, Donini P (2003) Development and analysis of multiplex microsatellite markers sets in common bean (Phaseolus vulgaris L.). Mol Breed 11:303–313. doi:10.1023/A:1023443109985
Mehetre SS, Aher AR, Gawande VL, Patil VR, Mokate AS (2003) Induced polyploidy in Gossypium: a tool to overcome interspecific incompatibility of cultivated tetraploid and diploid cottons. Curr Sci 84(12):1510–1512
Meredith WR Jr (1991) Contributions of introductions to cotton improvement. In: Shands HL, Weisner LE (eds) Use of plant introductions in cultivar improvement. Part 1 CSSA. Madison, WI, pp 127–146
Multani DS, Lyon BR (1995) Genetic fingerprinting of Australian cotton cultivars with RAPD markers. Genome 38:1005–1008. doi:10.1139/g95-132
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273. doi:10.1073/pnas.76.10.5269
Park Y-H, Alabady MS, Ulloa M, Sickler B, Wilkins TA, Yu J, Stelly DM, Kohel RJ, El-Shihy OM, Cantrell RG (2005) Genetic mapping of new cotton fiber loci using EST-derived microsatellites in an interspecific recombinant inbred line cotton population. Mol Genet Genomics 274:428–441. doi:10.1007/s00438-005-0037-0
Reddy OUK, Pepper AE, Abdurakhmonov I, Saha S, Jenkins JN, Brooks TD, Bolek Y, El-Zik KM (2001) New dinucleotide and trinucleotide microsatellite markers for cotton genomic research. J Cotton Sci 5:103–113
Rohlf FJ (2000) NTSYS-pc: Numerical Taxonomy and Multivariate Analysis System, Version 2.1. User Guide. Exeter Software, New York
Saha S, Wu J, Jenkins JN, McCarty JC, Gutierrez OA Jr, Stelly DM, Percy RG, Raska DA (2004) Effect of chromosome substitutions from Gossypium barbadense L. 3–79 into G.hirsutum L. TM-1 on agronomic and fiber traits. J Cotton Sci 8:162–169
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Evol Biol 4:406–425
Smulders MJM, Bredemeijer G, Rus-Kortekaas W, Arens P, Vosman B (1997) Use of short microsatellites from database sequences to generate polymorphisms among Lycopersicon esculentum cultivars and accessions of other Lycopersicon species. Theor Appl Genet 97:264–272. doi:10.1007/s001220050409
Song QJ, Quigley CV, Nelson RL, Carter TE, Boerma HR, Strachan JL, Cregan PB (1999) A selected set of trinucleotide simple sequence repeat markers for soybean cultivar identification. Plant Var Seeds 12:207–220
Stewart JMD (1995) Potential for crop improvement through germplasm enhancement and genetic engineering. In: Constable G, Forester N (eds) Challenging the future. Proc World Cotton Research Conference-1. CSIRO, Melbourne, pp 313–327
Struss D, Plieske J (1998) The use of microsatellite markers for detection of genetic diversity in barley populations. Theor Appl Genet 97:308–315. doi:10.1007/s001220050900
Tatineni V, Canterell RG, Davis DD (1996) Genetic diversity in elite cotton germplasm determined by morphological characteristics and RAPDs. Crop Sci 36:186–192
Ulloa M, Meredith WR Jr, Percy R, Moser H (1999) Genetic variability within improved germplasm of Gossypium hirsutum and G. barbadense cottons. Crop Sci abstr, Annu Mtg ASA, CSSA, SSSA, Madison, WI, USA, p 73
Ulloa M, Brubaker C, Chee P (2007) Cotton. In: Kole C (ed) Genome mapping & molecular breeding. vol 6: technical crops. Springer, New York, pp 1–49
Van Esbroeck GA, Bowman DT, May OL (1997) Pedigrees and distinguishing features of upland and pima cotton germplasm lines released from 1970 to 1996. N C Agric Res Serv Bull 312
Vergara GV, Bughrara SS (2003) AFLP analyses of genetic diversity in Bentgrass. Crop Sci 43:2162–2171
Vigouroux Y, Jaqueth JS, Matsuoka Y, Smith OS, Beavis WD, Smith SC, Doebley J (2002) Rate and pattern of mutation at microsatellite loci in maize. Mol Biol Evol 19:1251–1260
Xu QH, Zhang XL, Nie YC (2001) Genetic diversity evaluation of cultivars (G. hirsutum L.) from the Changjiang River valley and Yellow River valley by RAPD markers. Acta Genet Sin 28:683–690
Zhu LF, Zhang XL, Nie YC (2003) Analysis of genetic diversity in upland cotton (Gossypium hirsutum L.) cultivars from China and foreign countries by RAPDs and SSRs. J Agric Biotechnol 11:450–455
Acknowledgement
We thank the Delta & Pine Land Company for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kantartzi, S.K., Ulloa, M., Sacks, E. et al. Assessing genetic diversity in Gossypium arboreum L. cultivars using genomic and EST-derived microsatellites. Genetica 136, 141–147 (2009). https://doi.org/10.1007/s10709-008-9327-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-008-9327-x