Abstract
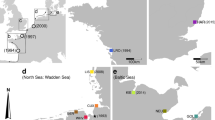
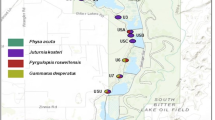
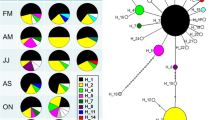
Colonization events like range expansion or biological invasions can be associated with population bottlenecks. Small population size may lead to loss of genetic diversity due to random genetic drift, to loss of heterozygosity due to increased inbreeding and should leave a signature on the genetic polymorphism and genetic structure of populations. The mating system might additionally influence the outcome of such a process. Here, we compare invasive and native populations of the hermaphroditic freshwater snail Lymnaea stagnalis. In the native range we included populations that were ice-free during the last glaciation period and populations that were glaciated and are located at the edge of the species’ native distribution range. The microsatellite data show substantial loss of genetic variation in the introduced range and no signs of high propagule pressure or admixture. The expressed polymorphism was so low that mating system analysis was not possible. In the native region, all populations display strong levels of differentiation (global F ST: 0.341) independent of colonization history and exhibit no significant pattern of inbreeding. However, the populations in more recently colonized habitats show diminished genetic diversity. Overall, these results illustrate how dramatic the reduction in genetic diversity can be for hermaphroditic animals and that gene flow in the native range can be surprisingly low despite short distances.


Similar content being viewed by others
References
Andersen BG, Borns HW Jr (1997) The ice age world: an introduction to Quaternary history and research with emphasis on North America and Northern Europe during the last 2.5 million years. Scandinavian University Press, Oslo
Avise J (2000) Phylogeography. Harvard University Press, Cambridge
Baker HG (1955) Self-compatibility and establishment after “long-distance” dispersal. Evolution 9:347–348
Belkhir K, Borsa P, Chikhi L et al. (1996–2004) GENETIX 4.05, logiciel sous Windows TM pour la génétique des populations
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B (Stat Method) 57:289–300
Bernatchez L, Wilson CC (1998) Comparative phylogeography of Nearctic and Palearctic fishes. Mol Ecol 7:431–452
Burns JH, Ashman TL, Steets JA et al. (2011) A phylogenetically controlled analysis of the roles of reproductive traits in plant invasions. Oecologia 1–9
Cain GL (1956) Studies on cross-fertilization and self-fertilization in Lymnaea stagnalis appresa. Say Biol Bull Mar Biol Lab Woods Hole 111:45–52
Cavalli-Sforza LL, Edwards AWF (1967) Phylogenetic analysis. Models and estimation procedures. Am J Hum Genet 19:233–257
Chapuis M-P, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24:621–631
Coutellec-Vreto MA, Guiller A, Daguzan J (1994) Allozyme variation in some populations of the freshwater snails Lymnaea peregra, L. auricularia and L. stagnalis (Gastropoda: Pulmonata). J Molluscan Stud 60:393–403
Crow J, Kimura M (1970) An introduction to population genetic theory. Harper & Row, New York
David P, Pujol B, Viard FR et al (2007) Reliable selfing rate estimates from imperfect population genetic data. Mol Ecol 16:2474–2487
Dillon RT (2000) The ecology of freshwater mollusca. Cambridge University Press, Cambridge
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes - application to human mitochondrial-DNA restriction data. Genetics 131:479–491
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Facon B, Pointier JP, Jarne P et al (2008) High genetic variance in life-history strategies within invasive populations by way of multiple introductions. Curr Biol 18:363–367
Felsenstein J (2005) PHYLIP (Phylogeny Inference Package). 3.6 edn. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle
Gautschi B, Tenzer I, Muller JP et al (2000) Isolation and characterization of microsatellite loci in the bearded vulture (Gypaetus barbatus) and cross-amplification in three Old World vulture species. Mol Ecol 9:2193–2195
Gillis NK, Walters LJ, Fernandes FC et al (2009) Higher genetic diversity in introduced than in native populations of the mussel Mytella charruana: evidence of population admixture at introduction sites. Divers Distrib 15:784–795
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). Available from http://www.unil.ch/izea/softwares/fstat.html
Guo SW, Thompson EA (1992) Performing the exact test of Hardy-Weinberg proportion for multiple alleles. Biometrics 48:361–372
Hale ML, Bevan R, Wolff K (2001) New polymorphic microsatellite markers for the red squirrel (Sciurus vulgaris) and their applicability to the grey squirrel (S. carolinensis). Mol Ecol Notes 1:47–49
Hauser L, Carvalho GR, Hughes RN et al (1992) Clonal structure of the introduced freshwater snail Potamopyrgus antipodarum (Prosobranchia: Hydrobiidae), as revealed by DNA fingerprinting. Proc Biol Sci 249:19–25
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58:247–276
Hewitt GM (1999) Post-glacial re-colonization of European biota. Biol J Linn Soc 68:87–112
Hörandl E (2011) Evolution and biogeography of alpine apomictic plants. Taxon 60:390–402
Hutton FW (1881) Art. XIX.—notes on some pulmonate Mollusca. Trans Proc R Soc NZ 14:150–158
Hutton FW (1884) Art. VI.—the freshwater shells of New Zealand belonging to the family Limnæidæ. Trans Proc R Soc NZ 17:54–58
Jensen JL, Bohonak AJ, Kelley ST (2005) Isolation by distance, web service. BMC Genet 6
Knott KE, Puurtinen M, Kaitala V (2003) Primers for nine microsatellite loci in the hermaphroditic snail Lymnaea stagnalis. Mol Ecol Notes 3:333–335
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability in populations. Evolution 29:1–10
Puurtinen M, Knott KE, Suonpaa S et al (2004) Genetic variability and drift load in populations of an aquatic snail. Evolution 58:749–756
Puurtinen M, Knott KE, Suonpaa S et al (2007) Predominance of outcrossing in Lymnaea stagnalis despite low apparent fitness costs of self-fertilization. J Evol Biol 20:901–912
Raymond M, Rousset F (1995) Genepop (version-1.2)—population-genetics software for exact tests and ecumenicism. J Hered 86:248–249
Roman J, Darling JA (2007) Paradox lost: genetic diversity and the success of aquatic invasions. Trends Ecol Evol 22:454–464
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rozen S, Skaletsky HJ (1998) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Städler T, Frye M, Neiman M et al (2005) Mitochondrial haplotypes and the New Zealand origin of clonal European Potamopyrgus, an invasive aquatic snail. Mol Ecol 14:2465–2473
Stebbins GL (1957) Self fertilization and population variability in the higher plants. Am Nat 91:337–354
Svendsen JI, Alexanderson H, Astakhov VI et al (2004) Late Quaternary ice sheet history of northern Eurasia. Quat Sci Rev 23:1229–1271
Tenzer I, degli Ivanissevich S, Morgante M et al (1999) Identification of microsatellite markers and their application to population genetics of Venturia inaequalis. Phytopathology 89:748–753
Van Kleunen M, Manning John C, Pasqualetto V et al (2008) Phylogenetically independent associations between autonomous self-fertilization and plant invasiveness. Am Nat 171:195–201
Vandel A (1928) La parthénogénèse géographique contribution à l’étude biologiue et cytologique de la parthénogénèse naturelle. Bulletin Biologique de la France et de la Belgique 62:164–281
Verhoeven KJF, Simonsen KL, McIntyre LM (2005) Implementing false discovery rate control: increasing your power. Oikos 108:643–647
Walsh PS, Metzger DA, Higuchi R (1991) Chelex-100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10:506–513
Weir BS (1996) Genetic data analysis II. Sinauer Associates, Sunderland
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Woodruff DS, Mulvey M, Yipp MW (1985) Population-genetics of Biomphalaria straminea in Hong-Kong—a neotropical schistosome-transmitting snail recently introduced into China. J Hered 76:355–360
Acknowledgments
We thank Christine Reber Funk and Paul Schmid-Hempel for their support in developing microsatellite primers. We are grateful to Stefano Rezzonico, Karoliina Räsänen, Trent Garner and Martin Surbeck for their support in the lab and field. The paper benefited greatly from the comments of the editor and three anonymous referees. Research was funded by a Marie Curie Fellowship of the European Community at Newcastle University (contract number HPMT-CT-2001-00272, KK) and the Academy of Finland Centre of Excellence for Evolutionary Research and Swiss National Science Foundation (JJ).
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
Rights and permissions
About this article
Cite this article
Kopp, K.C., Wolff, K. & Jokela, J. Natural range expansion and human-assisted introduction leave different genetic signatures in a hermaphroditic freshwater snail. Evol Ecol 26, 483–498 (2012). https://doi.org/10.1007/s10682-011-9504-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10682-011-9504-8