Abstract
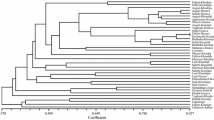
This study presents a grapevine breeding project that examines technologies for the development of new cultivars with a view to combining nematode resistance with desirable agronomic traits. The objectives of this study were to undertake a genetic-molecular characterization of a segregating population with resistance to the nematode P. brachyurus, resulting from hybridizations between different selections of Vitis, and to indicate superior segregants for future crosses. Seventy-four interspecific hybrids, seven parents, and variety Niagra were used. The genomic DNA of each individual was extracted by the CTAB method, and 10 microsatellite markers were employed. The number of fragments per primer ranged from 2 to 4. Mean observed heterozygosity (0.52) was higher than expected heterozygosity (0.42), and the fixation index averaged − 0.230. These values indicate broad genetic variability and are within the expected limits for grapevine, considering that the best performances are obtained with heterozygous individuals maintained through vegetative propagation. The genetic distance calculated by the weighted index and represented by dendrogram based UPGMA method resulted in the formation of five groups. Groups IV and V stood out with the highest number of individuals. Group IV had approximately 6% of its individuals with greater genetic similarity to the parents of V. vinifera origin. The largest group (Group V) had approximately 82% of its individuals with greater similarity to the parents of V. vinifera origin. Interspecific hybrids resistant to P. brachyurus and genetically closer to one of the parents of Vitis vinifera origin will be selected for future crosses, enabling the continuity of the grapevine breeding program.



Similar content being viewed by others
References
Bao LV, Scatoni IB, Gaggero C, Gutiérrez L, Monza J, Walker MA (2015) Genetic diversity of grape phylloxera leaf-galling populations on Vitis species in Uruguay. Am J Enol Viticult 66(1):43–53. https://doi.org/10.5344/ajev.2014.14026
Bowers JE, Bandman EB, Meredith CP (1993) DNA fingerprint characterization of some wine grape cultivars. Am J Enol Viticult 44:266–274
Bowers JE, Dangl GS, Vignani R, Meredith CP (1996) Isolation and characterization of new polymorphic simple sequence repeat loci in grape (Vitis vinifera L.). Genome 39:628–633. https://doi.org/10.1139/g96-080
Bowers JE, Dangl GS, Meredith CP (1999) Development and characterization of additional microsatellite DNA markers for grape. Am J Enol Viticult 50:243–246
Cruz DC (2016) Genes Software—extended and integrated with the R, Matlab and Selegen. Acta Sci Agron 3(4):547–552
Cunff LL, Fournier-Level A, Laucou V, Vezzulli S, Lacombe T, Adam-Blondon AF, Boursiquot JM, This P (2008) Construction of nested genetic core collections to optimize the exploitation of natural diversity in Vitis vinifera L. subsp. sativa. BMC Plant Biol 8:31. https://doi.org/10.1186/1471-2229-8-31
Dangl GS, Mendum ML, Yang J, Walker MA, Preece JE (2015) Hybridization of cultivated Vitis vinifera with wild V. californica and V. girdiana in California. Ecol Evol 5(23):5671–5684. https://doi.org/10.1002/ece3.1797
Dettweiler E, Jung A, Zyprian Zyprian Eva, Topfer R (2000) Grapevine cultivar Mueller-Thurgau and its true to type descent. Vitis 39:63–65
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Goto-Yamamoto N, Akifumi A, Nobuhito M, Shozo K (2013) SSR genotyping of wild grape species and grape cultivars of Vitis vinifera and V. vinífera × V. labrusca. J Jpn Soc Hortic Sci 82(2):125–130. https://doi.org/10.1080/14620316.2006.11512187
IBGE (2016) Instituto Brasileiro de Geografia e Estatística. Banco de dados agregados: produção agrícola municipal. Rio de Janeiro. Disponível em: http://www.sidra.ibge.gov.br/bda/tabela/listabl.asp?c=106&z=p&o=28. Accessed 18 Jan 2018
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Leão PCS, Cruz CD, Motoike SY (2013) Diversity and genetic relatedness among genotypes of Vitis spp. using microsatellite molecular markers. Rev Bras Frutic 35:799–808. https://doi.org/10.1590/S0100-29452013000300017
Moreira FM, Madini A, Marino R, Zulini L, Stefanini M, Velasco R, Kozma P, Grando MS (2011) Genetic linkage maps of two interspecific grape crosses (Vitis spp.) used to localize quantitative trait loci for downy mildew resistance. Tree Genet Genomes 7:153–167. https://doi.org/10.1007/s11295-010-0322-x
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28(19):2537–2539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Riaz S, Dangl GS, Edwards KJ, Meredith CP (2004) A microsatellite marker based framework linkage map of Vitis vinifera L. Theor Appl Genet 108:864–872. https://doi.org/10.1007/s00122-003-1488-5
Riaz S, Krivanek AF, Xu K, Walker MA (2006) Refined mapping of the Pierce’s disease resistance locus, PdR1, and Sex on an extended genetic map of Vitis rupestris × V. arizonica. Theor Appl Genet 113:1317–1329. https://doi.org/10.1007/s00122-006-0385-0
Riaz S, Karl TL, Granett J, Walker AM (2017) Population diversity of grape phylloxera in California and evidence for sexual reproduction. Am J Enol Vitic 68:218–227. https://doi.org/10.5344/ajev.2016.15114
Sant’Ana JC, Hidalgo E, De Lucas AI, Recio P, Ortiz JM, Martin JP (2007) Identification and relationships of accessions grown in the grapevine (Vitis vinifera L.) Germplasm Bank of Castilla y Léon (Spain) and the varieties authorized in the VQPRD areas of the region by SSR marker analysis. Genet Resour Crop Evol 55:573–583. https://doi.org/10.1007/s10722-007-9261-2
Sant’Ana JC, Ferreira JL, Rocha HS, Borém A, Pasqual M, Cançado GMA (2012) Comparison of a retrotransposon-based marker with microsatellite markers for discriminating accessions of Vitis vinífera. Genet Mol Res 11(2):1507–1525. https://doi.org/10.4238/2012
Santos PR, Viana AP, Gomes VM, Preisigke SC, Santos EA, Cavalcante NR, Almeida OF, Walker MA (2018) Clonal selection in interspecific Vitis spp. hybrids resistant to the root-lesion nematode Pratylenchus brachyurus by REML/BLUP. Fruits 73(3):191–197. https://doi.org/10.17660/th2018/73.3.6
Schuck MR, Biasi LA, Mariano AM, Lipski B, Riaz S, Walker MA (2011) Obtaining interspecific hybrids, and molecular analysis by microsatellite markers in grapevine. Pesq Agrop Bras 46:1480–1488. https://doi.org/10.1590/S0100-204X2011001100009
Sefc KM, Regner F, Turetschek E, Glцssl J, Steinkellner H (1999) Identification of microsatellite sequences in Vitis riparia and their applicability for genotyping of different Vitis species. Genome 42:367–373
Sefc KM, Lopes MS, Lefort F, Botta R, Roubelakisangelaskis KA, Ibanez J, Pejic I, Wagner HW, Glossi J, Steinkellener H (2000) Microsatellite variability in grapevine cultivars from different European regions and evaluation of assignment testing to assess the geographic origin of cultivars. Theor Appl Genet 100:498–505. https://doi.org/10.1007/s001220050065
Taylor AL (1967) Introduction to research on plant nematology: an FAO guide to study and control of the plant parasitic nematodes. Food And Agricultural Organization of the United Nations, Rome, p 133
This P, Dettweiler E (2003) EU-project GENRES ct96 no81 European Vitis database and results regarding the use of a common set of microsatellite markers. Acta Hortic 603:59–66. https://doi.org/10.17660/ActaHortic.2003.603.3
This P, Jung A, Boccacci P, Borrego J, Botta R, Costantini L, Crespan M, Dangl GS, Eisenheld C, Ferreira-Monteiro F, Grando S, Ibanez J, Lacombe T, Laucou V, Magalhaes R, Meredith CP, Milani N, Peterlunger E, Regner F, Zulini L, Maul E (2004) Development of a standard set of microsatellite reference alleles for identification of grape cultivars. Theor Appl Genet 109:1448–1458
Viana AP, Riaz S, Walker MA (2011) Evaluating genetic diversity and optimizing parental selections in a segregating table-grape population. Am J Enol Viticult 62:285–290. https://doi.org/10.5344/ajev.2011.10073
Viana AP, Riaz S, Walker MA (2013) Genetic dissection of agronomic traits within a segregating population of breeding table grapes. Genet Mol Res 12:951–964. https://doi.org/10.4238/2013.April.2.11
Viana AP, Resende MDV, Riaz S, Walker MA (2016) Genome selection in fruit breeding: application to table grapes. Sci Agric 73:142–149. https://doi.org/10.1590/0103-9016-2014-0323
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
dos Santos, P.R., Viana, A.P., Santos, E.A. et al. Molecular genetic diversity in segregates of Vitis: implications for the breeding of grapevine aiming at resistance to Pratylenchus brachyurus. Euphytica 215, 78 (2019). https://doi.org/10.1007/s10681-019-2403-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2403-8