Abstract
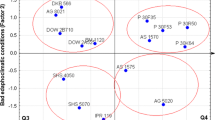
The aim of this study was to use a multi-environment trial approach from a mixed model point of view for factor analysis (FA) of the stability and adaptability of hybrids. Twenty-eight hybrids were analyzed in 35 environments across four seasons/years (summer season 2010, winter season 2011, summer season 2011 and winter season 2012). Several of these hybrids were analyzed during the first seasons and were not evaluated in later seasons or vice versa. Therefore, the dataset used in this study simulated the dynamics of a genetic breeding program with removal and inclusion of genotypes over the years. A biplot of the factor scores and loadings showed that the environments were more similar within seasons than between seasons, thereby suggesting that a given site may behave differently year after year. The season was more effective in discovery mega-environments. The FA models may be directly interpreted as GGE biplot analysis since the first factor score had a perfect fit (\(r^{2} = 0.99\)) with the empirical best linear unbiased predictors of the genotypes. Given the assumption of normality for the factor scores, confidence ellipses could be created to directly compare the genotypes in the biplot. Stability and adaptability could be analyzed in unbalanced experiments with the removal and inclusion of genotypes over time. This approach allowed certain trends in a breeding program to be measured by directly comparing hybrids developed for the first or winter season. The biplot interpretation was direct and intuitive, and it has the same properties as the GGE biplot obtained by singular value decomposition.







Similar content being viewed by others
References
Annicchiarico P (2002) Genotype × environment interactions: challenges and opportunities for plant breeding and cultivar recommendations. Food and Agricultural Organization, Rome, Italy. (FAO Plant Production and Protection Paper, 174)
Burgueño J, Crossa J, Cornelius PL, Yang RC (2008) Using factor analytic models for joining environments and genotypes without crossover genotype × environment interaction. Crop Sci 48:1291–1305
Burgueño J, Crossa J, Cotes JM, San Vicente F, Das B (2011) Prediction assessment of linear mixed models for multienvironment trials. Crop Sci 51:944–954
Burgueño J, de los Campos G, Weigel K, Crossa J (2012) Genomic prediction of breeding values when modeling genotype × environment interaction using pedigree and dense molecular markers. CropSci. doi:10.2135/cropsci2011.06.0299.x
Crossa J (1990) Statistical analysis of multilocations trials. Adv Agron 44:55–85
Crossa J, Burgueño J, Cornelius PL, Mclaren G, Trethowan R, Krishnamachari A (2006) Modeling genotype × environment interaction using additive genetic covariances of relatives for predicting breeding values of wheat genotypes. Crop Sci 46:1722–1733
Crossa J, Vargas M, Joshi AK (2010) Linear, bilinear, and linear–bilinear fixed and mixed models for analyzing genotype × environment interaction in plant breeding and agronomy. Can J Plant Sci 90:561–574
Crossa J, Vargas M, Cossani CM, Alvarado G, Burgueño J, Mathews KL, Reynolds MP (2013) Evaluation and interpretation of interactions. Agron J 105:1–12
Dempster AP, Laird NM, Rubin DF (1977) Maximum likelihood from incomplete data with EM algorithm. J R Stat Soc 39:1
Gauch HG (1992) Statistical analysis of regional yield trials: AMMI analysis of factorial designs. Elsevier, Amsterdam
Gauch HG (2007) MATMODEL version 3.0: open source software for AMMI and related analyses. Crop and Soil Sci, Cornell University, Ithaca, New York
Gauch HG, Zobel RW (1990) Imputing missing yield trial data. Theor Appl Genet 79:753–761
Gauch HG, Zobel RW (1997) Identifying mega-environments and targeting genotypes. Crop Sci 37:311–326
Gauch HG, Piepho HP, Annicchiarico P (2008) Statistical analysis of yield trials by AMMI and GGE: further considerations. Crop Sci 48:866–889
Gilmour AR, Cullis BR, Gogel BJ, Welham SJ, Thompson R (2005) ASReml user guide, release 2.0. VSN Int., Hemel Hempstead, UK
Henderson CR (1984) Applications of linear models in animal breeding. University of Guelph, Guelph, Ontario
Kelly AM, Smith AB, Eccleston JA, Cullis BR (2007) The accuracy of varietal selection using factor analytic models for multi-environment plant breeding trials. Crop Sci 47:1063–1070
Kelly AM, Cullis BR, Gilmour AR, Eccleston JA, Thompson R (2009) Estimation in a multiplicative mixed model involving a genetic relationship matrix. Genet Sel Evol 41:33
Meyer K (2009) Factor-analytic models for genotype × environment type problems and structured covariance matrices. Genet Sel Evol 41:1–11
Paderewski J, Rodrigues PC (2014) The usefulness of EM-AMMI to study the influence of missing data pattern and application to Polish post-registration winter wheat data. Aust J Crop Sci 8:640–645
Piepho HP (1998) Empirical best linear unbiased prediction in cultivar trials using factor-analytic variance-covariance structures. Theor Appl Genet 97:195–201
Resende MDV, Thompson R (2004) Factor analytic multiplicative mixed models in the analysis of multiple experiments. Rev de Mat e Estat 22:1–22
Smith AB, Cullis BR, Gilmour A (2001a) The analysis of crop variety evaluation data in Australia. Aust N. Z. J Stat 43:129–145
Smith AB, Cullis BR, Thompson R (2001b) Analyzing variety by environment data using multiplicative mixed models and adjustments for spatial field trend. Biometrics 57:1138–1147
Smith AB, Cullis BR, Thompson R (2005) The analysis of crop cultivar breeding and evaluation trials: an overview of current mixed model approaches. J Agric Sci 143:449–462
Thompson R, Cullis B, Smith A, Gilmour A (2003) A sparse implementation of the average information algorithm for factor analytic and reduced rank variance models. Biometrics. Aust N. Z. J Stat 45:445–459
Yan W (2010). Comment on “biplot analysis of genotype × environment interaction: proceed with caution,” by R.-C. Yang, J. Crossa, P.L. Cornelius, and J. Burgueño in Crop Science. Crop Sci 50:1121–1123
Yan W, Kang M (2003) GGE biplot analysis: a graphical tool for breeders, geneticists, and agronomists. CRC Press, Boca Raton
Yan W, Hunt LA, Sheng Q, Szlavnics Z (2000) Cultivar evaluation and mega-environment investigation based on the GGE biplot. Crop Sci 40:597–605
Yan W, Kang MS, Ma B, Woods S, Cornelius PL (2007) GGE biplot vs. AMMI analysis of genotype-by-environment data. Crop Sci 47:643–655
Yang R, Crossa J, Cornelius Pl, Burgueño J (2009) Biplot analysis of genotype × environment Interaction: proceed with caution. Crop Sci 49:1564–1576
Zobel RW, Wright MJ, Gauch HG Jr (1988) Statistical analysis of a yield trial. Agron J 80:388–393
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
de Figueiredo, A.G., Von Pinho, R.G., Silva, H.D. et al. Application of mixed models for evaluating stability and adaptability of maize using unbalanced data. Euphytica 202, 393–409 (2015). https://doi.org/10.1007/s10681-014-1301-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-014-1301-3