Abstract
Citrus Greening disease (CG) in South Africa (SA) is associated with the fastidious bacterium ‘Candidatus Liberibacter africanus’ (Laf). It has been observed that Laf isolates obtained from different geographic localities in SA differed in the rate of transmission during grafting experiments leading to the hypothesis that genetic variation of Laf may exist in this country. To determine this, 167 Laf isolates obtained from Limpopo, North West, Mpumalanga and the Western Cape were subjected to microsatellite analyses, using four polymorphic markers. From UPGMA and STRUCTURE analysis, it was shown that most sources belong to one of two major genetic groups of Laf and these comprise 25 distinct haplotypes. Four samples included within this study did not group with these two major groups, suggesting a potential third and fourth genetic group of Laf being present, which can be validated by further sampling. Results further indicate that Laf populations in SA are formed by geographic locality. The high genetic diversity observed for Laf within this study is consistent with the hypothesis that Laf originated on the African continent, warranting further genetic analysis of Laf populations from Africa. This is the first study to unveil the genetic diversity of Laf.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Citrus greening disease (CG), which is characterised by leaf mottling, the production of lopsided bitter fruit and early fruit drop, has afflicted citrus species of commercial value in South Africa since 1928 (Oberholzer et al. 1963). It was initially speculated that this disease was due to mineral deficiencies as the leaf symptoms observed were similar to that of a zinc deficiency. It was however not until the 1960’s that it was demonstrated that this disease was biological in nature and could be transmitted both mechanically and by the triozid vector, Trioza erytreae Del Guercio (Hemiptera: Triozidae) (McClean and Oberholzer 1965a, b). It has since been determined that CG in Africa is historically associated with the fastidious bacterium, ‘Candidatus Liberibacter africanus’ (Laf) (Garnier and Bové 1983; Jagoueix et al. 1994). A more invasive and serious emerging disease worldwide, known as Huanglongbing (HLB), but commonly also referred to as citrus greening, causes foliar and fruit symptoms on citrus identical to CG, but is associated with the close relatives of Laf, ‘Ca. L. asiaticus’ (Las) (Jagoueix et al. 1994) and ‘Ca. L. americanus’ (Lam) (Teixeira et al. 2005a), both of which are vectored by the liviid, Diaphorina citri Kuwayama (Hemiptera: Liviidae) (Capoor et al. 1967; Teixeira et al. 2005b). To avoid confusion we refer to the Laf associated disease as African citrus greening (ACG).
Various levels of tolerance exist amongst citrus cultivars against Laf, with sweet orange, tangelo and mandarin being severely affected by ACG, grapefruit, lemon and sour orange having moderate tolerance and lime, pomelo and trifoliate orange being tolerant against this disease (Manicom and van Vuuren 1990). These different levels of susceptibility of citrus to Laf were based on the severity of symptoms observed on different cultivars in South Africa (Manicom and van Vuuren 1990). The possibility, however, exists that the level of symptom severity being expressed may be attributed to the presence of different genetic populations of Laf (da Graça 1991). The only study on the genetic diversity of Laf, associated with ACG in South Africa was conducted in the late 1960’s when it was widely accepted that ACG in this country was due to a viral infection (McClean and Oberholzer 1965a). Schwarz (1972) proposed that different strains of the ‘greening virus’ were present in the country. These assumptions were based on graft transmission studies in which ACG affected material collected from different geographical localities were graft inoculated onto citrus seedlings and monitored over time. This study found that the rate of transmission, as well as the disease progression, differed depending on the geographical origin of the grafting material, suggesting the presence of different geographical strains of the organism associated with ACG.
In contrast to Laf, the genetic diversity of Las populations infecting citrus globally has been extensively studied. Las populations have been determined using sequence-specific approaches including PCR-restriction fragment length polymorphisms (PCR-RFLP) profiles of omp (outer membrane protein gene) sequences (Bastianel et al. 2005; Hu et al. 2011), identification of single nucleotide polymorphisms (SNP) between 16S rDNA sequences (Adkar-Purushothama et al. 2009; Tomimura et al. 2009; Gupta et al. 2011; Ghosh et al. 2013; Moreno-Enriquez et al. 2014) and ribosomal protein gene clusters (Adkar-Purushothama et al. 2009; Furuya et al. 2010) as well as the comparison of prophage sequences (Liu et al. 2011; Zheng et al. 2017).
The use of microsatellite markers (synonymous with Variable Number Tandem Repeats –VNTR and simple sequence repeats – SSR) to determine the genetic diversity of Las populations have proven popular due to their discriminatory powers to resolve genetic populations across limited isolates (Chen et al. 2010; Katoh et al. 2011; de Paula et al. 2019). Using microsatellite markers, it was determined that Las originated in India from where it spread to the rest of Asia (Islam et al. 2012). Using this technique, it was also demonstrated that the occurrence of Las in the Americas were due to a single introduction of Las from Southeast Asia to Brazil, and two separate introduction events of Las into Florida from China and either Southeast Asia or Brazil (Chen et al. 2010; Islam et al. 2012; Matos et al. 2013).
The 16S rDNA sequence of Laf from South Africa is homologous across different geographical isolates, as well as sharing 99% sequence identity with Laf-subspecies described from this country (Roberts et al. 2015; Roberts and Pietersen 2017). This is also true for omp and other ribosomal protein sequences of Laf (Roberts et al. 2015). Upon obtaining the complete genome of Laf, it was revealed that this Liberibacter contains two prophage sequences (Lin et al. 2015). It, however, remains uncertain whether these prophage sequences are shared amongst different Laf populations in South Africa, potentially making these unsuitable markers for population studies of Laf in this country. With the complete genome of Laf now being available, the current study aimed to determine the genetic composition of Laf populations from South Africa using microsatellite markers.
Method and materials
Samples assessed
A total of 144 DNA extracts obtained from a country-wide survey conducted in 2006 (Pietersen et al. 2010) were used to perform the population study. These samples were from citrus trees showing ACG symptoms and were collected from different geographical regions within South Africa as well as from different citrus cultivars. An additional 23 DNA extracts were received from Citrus Research International’s (CRI) DNA collection and were included in fragment analysis. All 167 samples had previously tested positive for Laf. To ensure that the samples contained Laf populations and were free of ‘Ca. L. africanus subsp. clausenae’ biovar citrus (LafCl bv. citrus), the only other Liberibacter previously found to infect citrus in eastern Africa, primers described by Roberts et al. (2017) based on the omp gene, were utilised.
Microsatellite primer design
A genome-wide search for microsatellite sequences from the complete Laf genome (Genbank accession CP004021) was performed and primers flanking such regions were obtained using msatcommander software (Faircloth 2008). Initial screening of primer sets for polymorphisms was carried out using a subset of Laf-positive samples originating from different geographical localities as well as from different citrus cultivars. Reactions were set up using GoTaq® G2 Flexi DNA polymerase (Promega, USA) as follows: 5 μl of 5X Green GoTaq® Flexi buffer, 2 μl 25 mM MgCl2, 0.13 μl GoTaq® G2 Flexi DNA polymerase (5 U/μl), 200 nM per primer, 200 nM dNTP mix, 0.5 μl target DNA and made up to a final volume of 25 μl with nuclease-free water. PCR cycling was performed using the following parameters: initial denaturation of 5 min at 94 °C followed by 35 cycles of 94 °C for 30 s, 58 °C for 30 s and 72 °C for 40 s. Final extension was performed for 10 min at 72 °C. Amplification products were viewed under UV following gel electrophoresis on 3% agarose gel.
The amplification products for primer sets targeting polymorphic loci were subsequently purified enzymatically with exonuclease I (Thermo Fisher Scientific, USA) and FastAP (Werle et al. 1994). The purified products per polymorphic primer set were sequenced unidirectionally with their respective forward primer using Big Dye Terminator v3.1 cycling sequencing kit (ABI, USA) according to the manufacturer’s instructions. Sanger sequencing was conducted on an ABI 3500xL automated sequencer at the University of Pretoria, South Africa. The sequences were then inspected with Chromas v2.6 to ensure the presence of the repeat motifs targeted.
Capillary electrophoresis
The forward primer of each of the polymorphic primer sets identified were labelled with the fluorescent dyes FAM, NED and VIC, respectively (Table 1). PCR amplification using these fluorescently labelled primers were performed in simplex on all 167 Laf-positive citrus samples using GoTaq® G2 Flexi DNA polymerase (Promega, USA) system, as previously described. Following visualization of amplification products with electrophoresis on a 2% agarose gel, the amplification products from each primer set were pooled in a 1:1:1:1 ratio per sample. The pooled samples were then further diluted in a 1:100 ratio with nuclease-free water. Prior to capillary electrophoresis, 1 μl of each pooled-diluted sample was added to 10.7 μl of Hi-Di Formamide and 0.3 μl GeneScan Liz500® (Thermo Fisher Scientific, USA). The reaction cocktail was then heated to 95 °C for 3 min and cooled on ice before being loaded onto a 3500 Genetic Analyzer (Applied Biosystems, USA) for fragment analysis. Analysis of the data obtained was conducted using Geneious® 11.1.5 software.
Genetic diversity analyses
From the multilocus allelic data obtained following fragment analysis, different genotypes were identified and compiled into a clonally corrected dataset. This dataset was used to determine the number of alleles per locus (Na), number of effective alleles (Ne) and haploid genetic diversity (H) using GenAIEx 6.503 (Peakall and Smouse 2006) for each loci.
Genetic structure analyses
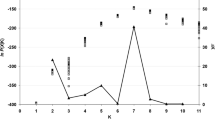
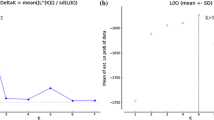
Using Nei’s genetic distance, a UPGMA dendrogram, derived from a genetic tri-square matrix obtained in GenAIEx, was constructed using MEGA X (Kumar et al. 2018). This was performed to obtain a visual representation of the genetic relationship amongst Laf isolates included in this study. A principal coordinate analysis (PCoA) analysis was performed in GenAIEx to further determine the genetic diversity and genetic distance of Laf populations from South Africa. To validate the results obtained from UPGMA and PCoA analysis, a Bayesian modelling analysis using STRUCTURE 2.3.1 (Pritchard et al. 2000) was performed in which the number of clusters (K) was determined by running 100 independent runs of K = 1–10. A burn-in period of 25,000 and a run length of 50,000 was set for each run. The results from STRUCTURE were assessed in STRUCTURE harvester (http://taylor0.biology.ucla.edu/structureHarvester/).
Results
All 167 samples tested positive for Laf senso stricto with none appearing to contain LafCl bv. citrus, which was previously found to infect citrus in Eastern Africa (Roberts et al. 2017), following end-point PCR using the omp primers designed by Roberts et al. (2017). In total, 50 primer sets for the amplification of microsatellite loci were designed and tested for polymorphisms. Of these, only five detected polymorphisms, of which four were able to amplify their intended target for all Laf-positive samples included in initial screening. The sequences of these polymorphic primer sets are listed in Table 1.
For the four loci assessed by fragment analysis, the number of alleles ranged from 2 to 8 and the haploid genetic diversity from 0.053 to 0.802 (Table 1). When the genetic diversity per province was determined, the highest genetic diversity was observed in the Western Cape (0.442), whereas the lowest diversity was observed in Limpopo (0.399) (Table 2).
UPGMA analyses revealed the presence of 29 distinct Laf haplotypes being present in South Africa (Fig. 1). Sixteen of these observed haplotypes contain multiple Laf isolates found in different provinces and citrus types and 13 haplotypes consist of only a single Laf isolate. This analysis further demonstrated the presence of two major genetic Laf populations (denoted as Group I and Group II), with a collective of 163 Laf isolates included in this study being represented within these two groups. The four remaining isolates (4/167) grouped outside of these two clusters in a potential third and fourth genetic population. However, further sampling will be required to justify the existence of these clusters.
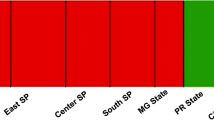
STRUCTURE analysis indicated that the number of genetic groups was two (K = 2) (Fig. S1), further demonstrating the need for future sampling to include samples from areas not represented within this study such as KwaZulu-Natal, the Southern Cape (i.e Knysna) and Gauteng.
Within the two major Laf genetic populations (i.e. Group I and Group II) Group I is primarily composed of Laf isolates found in the northern provinces, i.e. North West, Limpopo and Mpumalanga. Group II, however, contains most of the Laf isolates from the Western Cape (22/27) included in this study, isolates from Mpumalanga and North West and only a single isolate from Limpopo (Fig. 2). When assessed by citrus type, Laf isolated from Lemon mainly fell within Group II (20 of 23 lemon isolates included), and Laf isolated from sweet orange mainly clustered in Group I (100 of 120 isolates). Of the soft citrus Laf isolates, 10/16 isolates clustered with Group II haplotypes whereas all three Laf isolates from grapefruit clustered in Group I. These two citrus types are however underrepresented within the current study, and therefore further studies are needed to conclusively determine whether host selection plays a role in shaping Laf populations.
PCoA analysis supported the presence of the 29 distinct Laf haplotypes in South Africa. Isolates from Limpopo, North West and Mpumalanga (orange squares, blue diamonds and yellow crosses on Fig. 3), were shown to be made up of a number of single haplotypes, mainly obtained from sweet orange, which are not shared by other provinces. Interestingly, a number of isolates from Mpumalanga and the Western Cape (Yellow crosses and grey triangles on Fig. 3) grouped together, suggesting that these haplotypes are either shared amongst these two provinces or closely related.
Principal coordinate analysis (PCoA) performed on clonal corrected dataset, representing the genetic distance and diversity of Laf populations in South Africa across different provinces and citrus types. Blue  : North West isolates, Orange
: North West isolates, Orange  : Limpopo isolates, Yellow
: Limpopo isolates, Yellow  : Mpumalanga isolates, Grey
: Mpumalanga isolates, Grey  : Western Cape isolates
: Western Cape isolates
Discussion
The haploid genetic diversity for Laf populations in South Africa was shown to be 0.480, which is unexpectedly high, considering that Las populations from China and India have haploid genetic diversities of 0.342 and 0.360, respectively (Islam et al. 2012). Additionally, two distinct genetic groups containing a collective 25 haplotypes could be distinguished, with an additional four haplotypes being assigned to a potential third and fourth genetic group.
Microsatellite repeats are known to be influenced by environmental conditions (Zhou et al. 2014) and therefore it is not surprising that the two major genetic groups of Laf as described in this study were dominantly formed based on the geographical locality of the isolates studied. Group I is dominated by Laf isolates from the Northern provinces (i.e Limpopo, North West and Mpumalanga) whereas group II mainly contained Laf isolates from the Western Cape. Of these four provinces represented, the Western Cape is separated from the three aforementioned Northern provinces by both physical barriers and climatic conditions. The Western Cape comprises of a mountainous landscape, isolated from other citrus producing regions represented in this study, to the North by the arid landscape of the Northern Cape. Laf has not yet been identified to the east of the Western Cape, in the Eastern Cape, which potentially forms a bridge for the movement of Laf populations by its triozid vector to the eastern coastal production areas, therefore further isolating Laf in the Western Cape. Additionally, this province has a Mediterranean climate with a winter rainfall season, whereas Limpopo, Mpumalanga and the North West are all in the temperate climatic zones and receive rain during the summer months. The formation of Laf populations based on geographical locality, further supports the findings by Schwarz (1972), suggesting that different Laf populations have varying transmission rates when graft inoculated onto citrus. This would however require further studies to determine whether these populations are transmitted by different rates under controlled conditions.
When considering the history of ACG in SA, this disease was first observed in Rustenburg, North West during the late 1920’s from where it spread to Mpumalanga, Limpopo and KwaZulu-Natal (Pretorius and van Vuuren 2006). It was not until 1994 that the first ACG-like symptoms were observed on commercial citrus in the Western Cape with another 4 years to confirm the presence of Laf from affected samples (Garnier et al. 2000). This timeline would suggest that Laf was introduced into the Western Cape from the Northern provinces. Furthermore, the PCoA analyses revealed that Laf populations from Mpumalanga and the Western Cape are closely related, indicating the likelihood that Laf was introduced into the Western Cape from Mpumalanga. Since the introduction of Laf into the Western Cape, numerous haplotypes of Laf, as indicated by both the high haploid genetic diversity of 0.442 and UPGMA analysis, may have been formed, possibly aided by the environmental conditions as previously explained. However, the high genetic diversity of Laf observed in the Western Cape would suggest that Laf was present in this province prior to its detection from commercial citrus in 2000, going undetected for years in non-commercial settings. This alternative theory, is however not supported by the timeline of reports of ACG in citrus in the Western Cape, and unlikely seeing as symptoms of ACG would have alerted farmers within this province to an incursion of Laf. It is thus more likely that the climate of the Western Cape is conducive to the formation of genetic diverse populations.
The almost 100 years of ACG being known to infect commercial citrus in South Africa has resulted in Laf adaptation and genetic structures across different provinces. This geographical separation of genetic Liberibacter populations has been observed for populations of Las (Chen et al. 2010; Islam et al. 2012; Katoh et al. 2011, 2012) as well as ‘Ca. L. solanacearum’ (Lso) (Lin et al. 2012), suggesting that the citrus host has little influence on the genetic make-up of Liberibacter populations. Katoh et al. (2015), suggested that the number of alleles within polymorphic loci are influenced by psyllid transmission. As previously stated, Laf has had ample time for multiple passages of through psyllid vectors and hosts allowing for genetically unique populations to form. Additionally, based on the absence of Laf on other continents, it is believed that Laf is indigenous to the African continent, but not necessarily on citrus, which was only introduced during the 1600’s, but rather from an indigenous Rutaceous species. It would, however, be valuable to conduct a comparative population study on Laf isolates across Africa, using the multilocus microsatellite markers described herein. Such a study will give valuable insights into the origin and evolutionary processes that help shape Laf senso stricto from citrus on the African continent.
With this said, the current study was biased towards Laf isolates from sweet orange, with 124 of the 167 isolates included in this study being obtained from sweet oranges. This is not surprising as the production of sweet orange in the four provinces represented (30,764 ha), far exceeds that of soft citrus (8915 ha), grapefruit (6054 ha) and lemons (5693 ha) (CGA Annual report 2018). The majority of Laf isolates from sweet orange fell within genetic Group I, and was mainly represented by isolates from the Northern provinces (i.e. Limpopo, North West and Mpumalanga). As for Laf isolated from lemon, these were mainly confined to Genetic group II, with 20 of the 23 lemon isolates included in the current study falling in this group. However, 13 of the 23 Laf isolates from lemons included in this study originated from the Western Cape, further indicating that Laf isolates are predominantly shaped by geographic locality rather than citrus type.
In addition to the two major genetic groups of Laf as revealed in this study, four Laf isolates fell outside of these groupings, suggesting the existence of a potential third and fourth genetic group of Laf being present in the country. Of the eight commercial citrus production provinces in the country, only four were represented. This may contribute to the weak support from this study for the existence of other genetic Laf populations. Additionally, only four polymorphic markers were assessed in this study, which is comparatively few considering that population studies of Las typically comprises of seven to eight polymorphic primers (Islam et al. 2012; de Paula et al. 2019). By including Laf isolates from other geographical localities, additional polymorphic loci could potentially be obtained from the primers described herein. Using these additional markers and including Laf isolates from other provinces not represented in the current study, greater support for the existence of a third and fourth genetic group could potentially be obtained.
The microsatellite markers described herein can be utilised for future studies in the population structure of Laf isolates from across Africa. This is the first study on the genetic variability of Laf in South Africa.
References
Adkar-Purushothama, C. R., Quaglino, F., Casati, P., Gottravalli Ramanayaka, J., & Bianco, P. A. (2009). Genetic diversity among ‘Candidatus Liberibacter asiaticus’ isolated based on single nucleotide polymorphisms in 16S rRNA and ribosomal protein genes. Annals of Microbiology, 59(4), 681–688.
Bastianel, C., Garnier-Semancik, G., Renaudin, J., Bové, J. M., & Eveillard, S. (2005). Diversity of ‘Candidatus Liberibacter asiaticus’ based on the omp gene sequence. Applied and Environmental Microbiology, 71(11), 6473–6478.
Capoor, S. P., Rao, D. G., & Viswanath, S. M. (1967). Diaphorina citri Kuwayama, a vector of the greening disease of citrus in India. Indian Journal of Agricultural Sciences, 37, 572–575.
Chen, J., Den, X., Sun, X., Jones, D., Irey, M., & Civerolo, E. (2010). Guangdong and Florida populations of ‘Candidatus Liberibacter asiaticus’ distinguished by a genomic locus with short tandem repeats. Phytopathology, 100, 567–572.
Citrus Growers’ Association of South Africa (CGA) (2018). Annual Report 2018. http://c1e39d912d21c91dce811d6da9929ae8.cdn.ilink247.com/ClientFiles/cga/CitrusGowersAssociation/Company/Documents/CGA%20AR%202018es(1).pdf. Accessed 18 Feb 2019
da Graça, J. V. (1991). Citrus greening disease. Annual Review of Phytopathology, 29, 109–136.
de Paula, L. B., Lin, H., Stuchi, E. S., Francisco, C. S., Safady, N. G., & Coletta-Filho, H. D. (2019). Genetic diversity of ‘Candidatus Liberibacter asiaticus’ in Brazil analysed in different geographic regions and citrus varieties. European Journal of Plant Pathology, 154, 863–872.
Faircloth, B. C. (2008). Msatcommander: Detection of microsatellite repeat arrays and automated, locus-specific primer design. Molecular Ecology Resources, 8(1), 92–94.
Furuya, N., Matsukura, K., Tomimura, K., Okuda, M., Miyata, S., & Iwanami, T. (2010). Sequence homogeneity of the serA-trmU-tufB-secE-nusG-rplKAJL-rpoB gene cluster and the flanking regions of Candidatus Liberibacter asiaticus isolates around Okinawa Main Island in Japan. Journal of General Plant Pathology, 76(2), 122–131.
Garnier, M., & Bové, J. M. (1983). Transmission of the organism associated with citrus greening disease from sweet orange to periwinkle by dodder. Phytopathology, 73, 1358–1363.
Garnier, M., Bové, J. M., Jagoueix-Eveillard, S., Cronje, C. P. R., Sander, G. M., Korsten, L., & le Roux, H. F. (2000). Presence of “Candidatus Liberibater africanus” in the Western cape province of South Africa. In J. V. da Graça, R. F. Lee, & R. K. Yokomi (Eds.), Proceedings of the 14th conference of the International Organization of Citrus Virologists (pp. 369–372). Riverside: University of California.
Ghosh, D., Bhose, S., Mukherjee, K., & Baranwal, V. K. (2013). Sequence and evolutionary analysis of ribosomal DNA from Huanglongbing (HLB) isolates of Western India. Phytoparasitica, 41(3), 295–305.
Gupta, K., Baranwal, V., & Haq, Q. (2011). Sequence analysis and comparison of 16S rDNA, 23S rRNA and 16S/23S intergenic spacer region of greening bacterium associated with yellowing disease (Huanglongbing) of Kinnow mandarin. Indian Journal of Microbiology, 52, 13–21.
Hu, W. Z., Wang, X. F., Zhou, Y., Li, Z. A., Tang, K. Z., & Zhou, C. Y. (2011). Diversity of the omp gene in Candidatus Liberibacter asiaticus in China. Journal of Plant Pathology, 93(1), 211–214.
Islam, M.–. S., Glynn, J. M., Bai, Y., Duan, Y.–. P., Coletta-Filho, H. D., Kuruba, G., Civerolo, E. L., & Lin, H. (2012). Multilocus microsatellite analysis of ‘Candidatus Liberibacter asiaticus’ associated with citrus Huanglongbing worldwide. BMC Microbiology, 12, 39. https://doi.org/10.1186/1471-2180-12-39.
Jagoueix, S., Bové, J. M., & Garnier, M. (1994). The phloem-limited bacterium of greening disease is a member of the α subdivision if the Proteobacteria. International Journal of Systematic Bacteriology, 44, 379–386.
Katoh, H., Subandiyah, S., Tomimura, K., Okuda, M., Su, H.–. J., & Iwanami, T. (2011). Differentiation of ‘Candidatus Liberibacter asiaticus’ isolates by variable-number tandem-repeat analysis. Applied and Environmental Microbiology, 77(5), 1910–1917.
Katoh, H., Davis, R., Smith, M. W., Weinert, M., & Iwanami, T. (2012). Differentiation of Indian, east Timorese, Papuan and Floridian ‘Candidatus Liberibacter asiaticus’ isolates on the basis of simple sequence repeat and single nucleotide polymorphism profiles at 25 loci. Annals of Applied Biology, 160, 291–297.
Katoh, H., Inoue, H., & Iwanami, T. (2015). Changes in variable number of tandem repeats in ‘Candidatus Liberibacter asiaticus’ through insect transmission. PLoS One, 10(9), e0138699.
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35, 1547–1549.
Lin, H., Islam, M. S., Bai, Y., Wen, A., Lan, S., Gumstad, N. C., & Civerolo, E. L. (2012). Genetic diversity of ‘Candidatus Liberibacter solanacearum’ strains in the United States and Mexico as revealed by simple sequence repeat markers. European Journal of Plant Pathology, 132, 297–308.
Lin, H., Pietersen, G., Han, C., Read, D. A., Lou, B., Gupta, G., & Civerolo, E. L. (2015). Complete genome sequence of ‘Candidatus Liberibacter africanus,’ a bacterium associated with citrus Huanglongbing. Genome Announcements, 3(4), e00733–e00715.
Liu, R., Zhang, P., Pu, X., Xing, X., Chen, J., & Deng, X. (2011). Analysis of a prophage gene frequency revealed population variation of ‘Candidatus Liberibacter asiaticus’ from two citrus-growing provinces in China. Plant Disease, 95, 431–435.
Manicom, B. Q. & van Vuuren, S. P. (1990). Symptoms of greening disease with special emphasis on African greening. In B. Aubert, S. Tontyaporn, & D. Buabgsuwon (Eds.) proceedings of the 4th international Asia Pacific Conference on Citrus Rehabilitation. Chiang Mai, Thailand, 4-10th Feb. 1990.
Matos, L. A., Hilf, M. E., Chen, J., & Folimonova, S. F. (2013). Validation of ‘variable number of tandem repeat’ – Based approach for examination of ‘Candidatus Liberibacter asiaticus’ diversity and its application for the analysis of the pathogen populations in the areas of recent introduction. PLoS One, 8(11), e78994.
McClean, A. P. D., & Oberholzer, P. C. J. (1965a). Greening disease of the sweet orange: Evidence that it is caused by a transmissible virus. South African Journal of Agricultural Science, 8, 253–276.
McClean, A. P. D., & Oberholzer, P. C. J. (1965b). Citrus psylla, a vector of the greening disease of sweet orange. South African Journal of Agricultural Science, 8, 297–298.
Moreno-Enriquez, A., Minero-Garcia, Y., Ramirez-Prado, J. H., Loeza-Kuk, E., Uc-Vargeuz, A., & Moreno-Valenzuela, O. A. (2014). Comparative analysis of 16S ribosomal RNA of Candidatus Liberibacter asiaticus associated with Huanglongbing disease of Persian lime and Mexican lime reveals a major haplotype with worldwide distribution. African Journal of Microbiology Research, 8(30), 2861–2873.
Oberholzer, P. C. J., von Staden, D. F. A., & Basson, W. J. (1963). Greening disease of sweet orange in South Africa. In W. C. Price (Ed.), Proceedings of the 3rd conference of the International Organization of Citrus Virologists (pp. 213–219). Riverside: University of California.
Peakall, R., & Smouse, P. E. (2006). GENALEX 6: Genetic analysis in excel. Population genetic software for teaching and research. Molecular Ecology Notes, 6, 288–295.
Pietersen, G., Arrebola, E., Breytenbach, J. H. J., Korsten, L., le Roux, H. F., la Grange, H., Lopes, S. A., Meyer, J. B., Pretorius, M. C., Schwerdtfeger, M., Vuuren, S. P., & Yamamoto, P. (2010). A survey for ‘Candidatus Liberibacter’ species in South Africa confirms the presence of only ‘Ca. L. africanus’ in commercial citrus. Plant Disease, 94, 244–249.
Pretorius, M. C., & van Vuuren, S. P. (2006). Managing Huanglongbing (Citrus greening disease) in the Western cape. South African Fruit Journal, 5(4), 59–62.
Pritchard, J., Stephens, M., & Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics, 155, 945–959.
Roberts, R., & Pietersen, G. (2017). A novel subspecies of ‘Candidatus Liberibacter africanus’ found on native Teclea gerrardii (family: Rutaceae) from South Africa. Antonie Van Leeuwenhoek, 110, 437–444.
Roberts, R., Steenkamp, E. T., & Pietersen, G. (2015). Novel lineages of ‘Candidatus Liberibacter africanus’ associated with native rutaceous hosts of Trioza erytreae in South Africa. International Journal of Systematic and Evolutionary Microbiology, 65, 723–731.
Roberts, R., Cook, G., Grout, T. G., Khamis, F., Rwomushana, I., Nderitu, P. W., Seguni, Z., Materu, C. L., Steyn, C., Pietersen, G., Ekesi, S., & le Roux, H. F. (2017). Resolution of the identity of ‘Candidatus Liberibacter’ species from Huanglongbing-affected citrus in East Africa. Plant Disease, 101(8), 1481–1488.
Schwarz, R. E. (1972). Strains of the greening pathogen. In W. C. Price (Ed.), Proceedings of the 5th conference of the International Organization of Citrus Virologists (pp. 40–44). Riverside: University of California.
Teixeira, D. C., Ayres, J., Kitajima, E. W., Danet, L., Jagoueix-Eveillard, S., Saillard, C., & Bové, J. M. (2005a). First report of a huanglongbing-like disease of citrus in Sao Paulo state, Brazil and association of a new liberibacter species, ‘Candidatus Liberibacter americanus’, with the disease. Plant Disease, 89, 107.
Teixeira, D. C., Saillard, C., Eveillard, S., Danet, J. L., da Costa, P. I., Ayres, A. J., & Bové, J. (2005b). ‘Candidatus Liberibacter americanus’ associated with citrus huanglongbing (greening disease) in Sao Paulo state, Brazil. International Journal of Systematic and Evolutionary Microbiology, 55, 1857–1862.
Tomimura, K., Miyata, S. I., Furuya, N., Kubota, K., Okuda, M., Subandiyah, S., Hung, T. H., Su, H. J., & Iwanami, T. (2009). Evaluation of genetic diversity among ‘Candidatus Liberibacter asiaticus’ isolates collected in Southeast Asia. Phytopathology, 99, 1062–1069.
Werle, E., Schneider, C., Renner, M., Völker, M., & Fiehn, W. (1994). Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Research, 22, 4354–4355.
Zheng, Z., Wu, F., Kumagai, L. B., Polek, M., Deng, X., & Chen, J. (2017). Two “Candidatus Liberibacter asiaticus” strains recently found in California harbour different prophages. Phytopathology, 107(6), 662–668.
Zhou, K., Aertsen, A., & Michiels, C. W. (2014). The role of variable DNA tandem repeats in bacterial adaption. FEMS Microbiological Reviews, 38, 119–141.
Acknowledgements
We would like to acknowledge Citrus Research International for funding as well as Dr. Glynnis Cook for providing additional Laf isolates from the CRI DNA collection. This work was partially supported by the European Union’s Horizon 2020 research and innovation programme under grant agreement No. 727459 (TROPICSAFE).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The work presented in the manuscript has not been published before or submitted to another journal for consideration. The work presented herein is the original work of the authors listed and all authors are aware of the manuscript submission to EJPP.
Conflict of interest
The authors declare that there are no conflict of interest.
Research involving human and animal rights
No humans or animals were used to conduct the research presented in this manuscripts.
Informed consent
The authors have given informed consent regarding the submission of the manuscript.
Supplementary Information
ESM 1
(DOCX 22.0 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Roberts, R., Lin, H. & Pietersen, G. Genetic diversity of ‘Candidatus Liberibacter africanus’ in South Africa based on microsatellite markers. Eur J Plant Pathol 159, 259–268 (2021). https://doi.org/10.1007/s10658-020-02180-w
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-020-02180-w