Abstract
Fusarium oxysporum f. sp. cubense is the causal agent of Panama disease of banana. A rapid and reliable diagnosis is the foundation of integrated disease management practices in commodity crops. For this diagnostic purpose, we have developed a reliable molecular method to detect Foc race 4 isolates in Taiwan. By PCR amplification, the primer set Foc-1/Foc-2 derived from the sequence of a random primer OP-A02 amplified fragment produced a 242 bp size DNA fragment which was specific to Foc race 4. With the optimized PCR parameters, the molecular method was sensitive and could detect small quantities of Foc DNA as low as 10 pg in 50 to 2,000 ng host genomic DNA with high efficiency. We also demonstrated that by using our PCR assay with Foc-1/Foc-2 primer set, Foc race 4 could be easily distinguished from other Foc races 1 and 2, and separated other formae speciales of F. oxysporum.







Similar content being viewed by others
References
Beckman, C. H. (1990). Host responses to the pathogen. In R. C. Ploetz (Ed.), Fusarium wilt of banana (pp. 107–114). St. Paul: APS.
Beckman, C. H., & Roberts, E. M. (1995). On the nature and genetic basis for resistance and tolerance to fungal wilt diseases of plants. Advances in Botanical Research, 21, 35–77. doi:10.1016/S0065-2296(08)60008-7.
Bentley, S., Pegg, K. G., Moore, N. Y., Davis, R. D., & Buddenhagen, I. W. (1998). Genetic variation among vegetative compatibility groups of Fusarium oxysporum f. sp. Cubense analysed by DNA fingerprinting. Phytopathology, 88, 1283–1288. doi:10.1094/PHYTO.1998.88.12.1283.
Brake, V. M., Pegg, K. G., Irwin, J. A. G., & Langdon, P. W. (1990). Vegetative compatibility groups within Australian populations of Fusarium oxysporum f. sp. cubense, the cause of Fusarium wilt of bananas. Australian Journal of Agricultural Research, 41, 863–870. doi:10.1071/AR9900863.
Chang, J. Y. (2005). Molecular identification of Fusarium oxysporum f. sp. cubense and its detection in infected banana seedlings. Taichung, Taiwan, ROC: National Chung Hsing University, Master’s thesis.
Chang, P. F. L., Chang, C. Y., Lin, E. T., Chen, I. R., & Huang, J. W. (2003). Detection of Fusarium oxysporum f. sp. cubense based on RAPD and PCR analysis. Plant Pathology Bulletin, 12, 277 Abstract in Chinese.
Daniells, J., Davis, D., Peterson, R., & Pegg, K. (1995). Goldfinger: Not as resistant to sigatoka/yellow sigatoka as first thought. Infomusa, 4, 6.
Dellaporta, S. L., Wood, J., & Hicks, J. B. (1983). A plant DNA mini-preparation: version II. Plant Molecular Biology Reporter, 1, 19–21. doi:10.1007/BF02712670.
Diener, A. C., & Ausubel, F. M. (2005). Resistance to Fusarium oxysporum 1, a dominant Arabidopsis disease-resistance gene, is not race specific. Genetics, 171, 305–321. doi:10.1534/genetics.105.042218.
Forsyth, L. M., Smith, L. J., & Aitken, E. A. B. (2006). Identification and characterization of non-pathogenic Fusarium oxysporum capable of increasing and decreasing Fusarium wilt severity. Mycological Research, 110, 929–935. doi:10.1016/j.mycres.2006.03.008.
Fungaro, M. H. P., Vissotto, P. C., Sartori, D., Vilas-Boas, L. A., Furlaneto, M. C., & Taniwaki, M. H. (2004). A molecular method for detection of Aspergillus carbonarius in coffee beans. Current Microbiology, 49, 123–127. doi:10.1007/s00284-004-4273-z.
Gerlach, K. S., Bentley, S., Moore, N. Y., Pegg, K. G., & Aitken, A. B. (2000). Characterisation of Australian isolates of Fusarium oxysporum f. sp. cubense by DNA fingerprinting analysis. Australian Journal of Agricultural Research, 51, 945–953. doi:10.1071/AR99172.
Getha, K., & Vikineswary, S. (2002). Antagonistic effects of Streptomyces violaceusniger strain G10 on Fusarium oxysporum f. sp. cubense race 4: indirect evidence for the role of antibiosis in the antagonistic process. Journal of Industrial Microbiology & Biotechnology, 28, 303–310. doi:10.1038/sj.jim.7000247.
Groenewald, S., Van Den Berg, N., Marasas, W. F., & Viljoen, A. (2006). The application of high-throughput AFLPs in assessing genetic diversity in Fusarium oxysporum f. sp. cubense. Mycological Research, 110, 297–305. doi:10.1016/j.mycres.2005.10.004.
Hwang, S. C., & Ko, W. H. (2004). Cavendish banana cultivars resistant to Fusarium wilt acquired through somaclonal variation in Taiwan. Plant Disease, 88, 580–588. doi:10.1094/PDIS.2004.88.6.580.
Jurado, M., Vázquez, C., Marín, S., Sanchis, V., & González-Jaéna, M. T. (2006). PCR-based strategy to detect contamination with mycotoxigenic Fusarium species in maize. Systematic and Applied Microbiology, 29, 681–689. doi:10.1016/j.syapm.2006.01.014.
Klemsdal, S. S., & Elen, O. (2006). Development of a highly sensitive nested-PCR method using a single closed tube for detection of Fusarium culmorum in cereal samples. Letters in Applied Microbiology, 42, 544–548. doi:10.1111/j.1472-765X.2006.01880.x.
Koike, M., Watanabe, M., Nagao, H., & Ohshima, S. (1995). Molecular analysis of Japanese isolates of Verticillium dahliae and V. alboatrum. Letters in Applied Microbiology, 21, 75–78. doi:10.1111/j.1472-765X.1995.tb01010.x.
Liu, E. T. (2003). Analysis of Fusarium oxysporum f. sp. cubense using random amplified polymorphic DNA and polymerase chain reaction techniques. Taichung, Taiwan, ROC: National Chung Hsing University, Master’s thesis.
Mes, J. J., Weststeijn, E. A., Herlaar, F., Lambalk, J. J. M., Wijbrandi, J., Haring, M. A., et al. (1999). Biological and molecular characterization of Fusarium oxysporum f. sp. lycopersici divides race 1 isolates into separate virulence groups. Phytopathology, 89, 156–160. doi:10.1094/PHYTO.1999.89.2.156.
Möller, E. M., Chełkowski, J., & Geiger, H. H. (1999). Species-specific PCR assays for the fungal pathogens Fusarium moniliforme and Fusarium subglutinans and their application to diagnose maize ear rot disease. Journal of Phytopathology, 147, 497–508. doi:10.1111/j.1439-0434.1999.tb03856.x.
Moore, N. Y., Bentley, S., Pegg, K. G., & Jones, D. R.(1995). Fusarium wilt of banana. Musa Disease Fact Sheet no. 5, International Network for the Improvement of Banana and Plantain, Montpellier, France.
Nash, S. M., & Snyder, W. C. (1962). Quantitative estimations by plate counts of propagules of the bean root rot Fusarium in field soils. Phytopathology, 52, 567–572.
Nicholson, P., Simpson, D. R., Weston, G., Rezanoor, H. N., Lees, A. K., Parry, D. W., et al. (1998). Detection and quantification of Fusarium culmorum and Fusarium graminearum in cereals using PCR assays. Physiological and Molecular Plant Pathology, 53, 17–37. doi:10.1006/pmpp.1998.0170.
O’Donnell, K., Kistler, H. C., Cigelnik, E., & Ploetz, R. C. (1998). Multiple evolutionary origins of the fungus causing Panama disease of banana: Concordant evidence from nuclear and mitochondrial gene genealogies. Proceedings of the National Academy of Sciences of the United States of America, 95, 2044–2049. doi:10.1073/pnas.95.5.2044.
Parry, D. W., & Nicholson, P. (1996). Development of a PCR assay to detect Fusarium poae in wheat. Plant Pathology, 45, 383–391. doi:10.1046/j.1365-3059.1996.d01-133.x.
Ploetz, R. C. (1990). Population biology of Fusarium oxysporum f. sp. cubense. In R. C. Ploetz (Ed.), Fusarium wilt of banana (pp. 63–76). St. Paul: APS.
Ploetz, R. C. (1994). Panama disease: Return of the first banana menace. International Journal of Pest Management, 40, 326–336.
Ploetz, R. C., & Correll, J. C. (1988). Vegetative compatibility among races of Fusarium oxysporum f. sp. cubense. Plant Disease, 72, 325–328. doi:10.1094/PD-72-0325.
Ploetz, R. C., Herbert, J., Sebasigari, K., Hernandez, J. H., Pegg, K. G., Ventura, J. A., et al. (1990). Importance of Fusarium wilt in different banana growing regions. In R. C. Ploetz (Ed.), Fusarium wilt of banana (pp. 9–26). St. Paul: APS.
Ploetz, R. C., & Pegg, K. G. (2000). Fusarium wilt. In D. R. Jones (Ed.), Diseases of banana, abacá and enset (pp. 143–159). Wallingford: CABI.
Schilling, A. G., Möller, E. M., & Geiger, H. H. (1996). Polymerase chain reaction-based assays for species-specific detection of Fusarium culmorum, F. graminearum, and F. avenaceum. Phytopathology, 86, 515–523. doi:10.1094/Phyto-86-515.
Snyder, W., & Hanson, H. (1940). The species concept in Fusarium. American Journal of Botany, 27, 64–67. doi:10.2307/2436688.
Stover, R. H., & Malo, S. E. (1972). The occurrence of fusarial wilt in normally resistance ‘dwarf Cavendish’ banana. Plant Disease Reporter, 56, 1000–1003.
Su, H. J., Chuang, T. Y., & Kong, W. S. (1977). Physiological race of Fusarial wilt fungus attacking Cavendish banana of Taiwan. Special publication no. 2 pp. 1–21. Pingtung: Taiwan Banana Research Institute.
Su, H. J., Hwang, S. C., & Ko, W. H. (1986). Fusarial wilt of Cavendish bananas in Taiwan. Plant Disease, 70, 814–818. doi:10.1094/PD-70-814.
Turner, A. S., Lees, A. K., Rezanoor, H. N., & Nicholson, P. (1998). Refinement of PCR-detection of Fusarium avenaceum and evidence from DNA marker studies for phenetic relatedness to Fusarium tricinctum. Plant Pathology, 47, 278–288. doi:10.1046/j.1365-3059.1998.00250.x.
Waite, B. H., & Stover, R. H. (1960). Studies on Fusarium wilt of bananas, VI. Variability and cultivar concept in Fusarium oxysporum f. sp. cubense. Canadian Journal of Botany, 38, 985–994. doi:10.1139/b60-087.
Wilson, A., Simpson, D., Chandler, E., Jennings, P., & Nicholson, P. (2004). Development of PCR assays for the detection and differentiation of Fusarium sporotrichioides and Fusarium langsethiae. FEMS Microbiology Letters, 233, 69–76. doi:10.1016/j.femsle.2004.01.040.
Yergeau, E., Filion, M., Vujanovic, V., & St-Arnaud, M. (2005). A PCR-denaturing gradient gel electrophoresis approach to assess Fusarium diversity in asparagus. Journal of Microbiological Methods, 60, 143–154. doi:10.1016/j.mimet.2004.09.006.
Yoder, W. T., & Christianson, L. M. (1998). Species-specific primers resolve members of Fusarium section Fusarium taxonomic status of the edible “Quorn” fungus reevaluated. Fungal Genetics and Biology, 23, 68–80. doi:10.1006/fgbi.1997.1027.
Acknowledgements
We thank Dr. Peter P. Ueng of USDA-ARS for providing DNA samples of reference isolates of Foc race 1, 2, and 4, and his critical review of this manuscript. We are grateful to Drs. Y.-S. Lin (NCHU), K.-S. Chen (FTHEB, ARI), S.-P.-Y. Hsieh (NCHU), S.-C. Hwang (TBRI), Miss H.-L. Lee (TDARES), WVC/AVRDC, and ARI for providing the tested microorganisms, and to Dr. W.-H. Ko for critical reading and useful suggestions for this manuscript. We also thank Miss L.J. Smith for VCG identification, Dr. R.C. Ploetz for information about Foc race 4, Miss Y.-L. Wan and Mr. C.-C. Su for technical assistance. This research was supported in part by Bureau of Animal and Plant Health Inspection and Quarantine, and Department of International Affairs, Council of Agriculture, Executive Yuan, Taiwan, R.O.C. under grant numbers 89ST-6.2-BQ-65(06), 91AS-7.3.1-BQ-B2(3), 93AS-1.9.2-BQ-B1, 96AS-4.1.2-IC-I1(2), and 97AS-4.1.2-IC-I1(2); by the Ministry of Education, Taiwan, R.O.C. under the ATU plan; and also by National Chung Hsing University, Taiwan, R.O.C.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
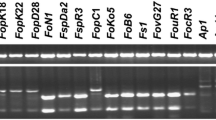
Amplification of PCR products using the primer set Foc-1/Foc-2 specific to Fusarium oxysporum f. sp. cubense (Foc) race 4 isolates. The fungal isolates used for extracting genomic DNA (gDNA) as PCR templates (50 ng) were as listed in Table 1. The DNA gels were subjected to Southern hybridisation (shown as the panel below each ethidium bromide-stained DNA gel pattern with light background) using the Biotin-labelled OPA02404 as a probe. The location of a 242-bp DNA band specific to F. oxysporum f. sp. cubense race 4 isolates (labeled as Foc) is indicated on the left. Fon-H0103 is the Fusarium wilt pathogen of watermelon to serve as a negative control here. N = negative control using sterile dH2O as the PCR template. M = molecular markers of Gen-100 DNA ladder (DOC 3.34 MB).
Rights and permissions
About this article
Cite this article
Lin, YH., Chang, JY., Liu, ET. et al. Development of a molecular marker for specific detection of Fusarium oxysporum f. sp. cubense race 4. Eur J Plant Pathol 123, 353–365 (2009). https://doi.org/10.1007/s10658-008-9372-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-008-9372-4