Abstract
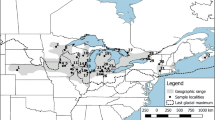
Blanding’s turtle (Emys blandingii) has declined substantially in North America due to anthropogenic activities, leaving populations smaller and increasingly fragmented spatially. We sampled 212 turtles to evaluate variation at eight microsatellite loci within and among 18 populations of E. blandingii across its primary range in the midwestern United States (Illinois, Iowa, Minnesota, and Nebraska). All loci and populations were highly polymorphic. Our analyses also detected considerable genetic structure within and among the sampled localities, and revealed ancestral gene flow of E. blandingii in this region north and east from an ancient refugium in the central Great Plains, concordant with post-glacial recolonization timescales. The data further implied unexpected ‘links’ between geographically disparate populations in Nebraska and Illinois. Our study encourages conservation decisions to be mindful of the genetic uniqueness of populations of E. blandingii across its primary range.



Similar content being viewed by others
References
Alacs EA, Janzen FJ, Scribner KT (2007) Genetic issues in freshwater turtle and tortoise conservation. Chelonian Res Monogr 4:107–123
Amato ML, Brooks RJ, Fu J (2008) A phylogeographic analysis of populations of the wood turtle (Glyptemys insculpta) throughout its range. Mol Ecol 17:570–581
Austin JD, Lougheed SC, Neidrauer L, Chek AA, Boag PT (2002) Cryptic lineages in a small frog: the post-glacial history of the spring peeper, Pseudacris crucifer (Anura: Hylidae). Mol Phylogenet Evol 25:316–329
Avise JC (2010) Perspective: conservation genetics enters the genomics era. Conserv Genet 11:665–669
Avise JC, Arnold J, Ball RM, Bermingham E, Lamb T, Neigel JE, Reeb CA, Saunders NC (1987) Intraspecific phylogeography—the mitochondrial DNA bridge between population genetics and systematics. Annu Rev Ecol Syst 18:489–522
Beaudry F, deMaynadier PG, Hunter ML Jr (2008) Identifying road mortality threat at multiple spatial scales for semi-aquatic turtles. Biol Conserv 141:2550–2563
Burnham KP, Anderson DR (1998) Model selection and inference: a practical information-theoretic approach. Springer-Verlag, New York
Byrne M, Yeates DK, Joseph L et al (2008) Birth of a biome: insights into the assembly and maintenance of the Australian arid zone biota. Mol Ecol 17:4398–4417
Christiansen JL (1998) Perspectives on Iowa’s declining amphibians and reptiles. J Iowa Acad Sci 105:109–114
Congdon JD, Keinath DA (2006) Blanding’s Turtle (Emydoidea blandingii): a technical conservation assessment. USDA Forest Service, Rocky Mountain Region. http://www.fs.fed.us/r2/projects/scp/assessments/blandingsturtle.pdf. Accessed Feb 2013
Congdon JD, Van Loben Sels RC (1993) Relationships of reproductive traits and body-size with attainment of sexual maturity and age in Blanding’s Turtles (Emydoidea blandingii). J Evol Biol 6:547–557
Congdon JD, Dunham AE, Van Loben Sels RC (1993) Delayed sexual maturity and demographics of Blanding’s turtles (Emydoidea blandingii)—implications for conservation and management of long-lived organisms. Conserv Biol 7:826–833
Congdon JD, Nagle RD, Osentoski MR, Kinney OM, Van Loben Sels RC (2003) Life history and demographic aspects of aging in the long-lived turtle (Emydoidea blandingii). In: Finch CE, Robine J-M, Christen Y (eds) Brain and longevity. Springer, Germany, pp 15–31
Earl DA, vonHoldt BM (2012) Structure harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Ehlers J, Gibbard P (2008) Extent and chronology of Quaternary glaciation. Episodes 31:211–218
Ernst CH, Lovich JE (2009) Turtles of the United States and Canada, 2nd edn. Johns Hopkins, Baltimore
Ersts PJ (2010) Geographic Distance Matrix Generator (version 1.2.3). American Museum of Natural History, Center for Biodiversity and Conservation. http://biodiversityinformatics.amnh.org/open_source/gdmg/documentation.php. Accessed Feb 2011
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes—application to human mitochondrial-DNA restriction data. Genetics 131:479–491
Fontanella FM, Feldman CR, Siddall ME, Burbrink FT (2008) Phylogeography of Diadophis punctatus: extensive lineage diversity and repeated patterns of historical demography in a trans-continental snake. Mol Phylogenet Evol 46:1049–1070
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Fritz U, Schmidt C, Ernst CH (2011) Competing generic concepts for Blanding’s, Pacific and European pond turtles (Emydoidea, Actinemys and Emys)-which is best? Zootaxa 2791:41–53
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Hey J, Nielsen R (2004) Multilocus methods for estimating population sizes, migration rates and divergence time, with applications to the divergence of Drosophila pseudoobscura and D. persimilis. Genetics 167:747–760
Hoffmann M, Hilton-Taylor C, Angulo A et al (2010) The impact of conservation on the status of the world’s vertebrates. Science 330:1503–1509
Holman JA (1995) Pleistocene amphibians and reptiles in North America. Oxford Monographs on Geology and Geophysics No. 32, Oxford
Howes BJ, Brown JW, Gibbs HL, Herman TB, Mockford SW, Prior KA, Weatherhead PJ (2009) Directional gene flow patterns in disjunct populations of the black ratsnake (Pantherophis obsoletus) and the Blanding’s turtle (Emydoidea blandingii). Conserv Genet 10:407–417
Howeth JG, McGaugh SE, Hendrickson DA (2008) Contrasting demographic and genetic estimates of dispersal in the endangered Coahuilan box turtle: a contemporary approach to conservation. Mol Ecol 17:4209–4221
Jackson CJ, Kaye JM (1974) The occurrence of Blanding’s turtle, Emydoidea blandingii, in the Late Pleistocene of Mississippi (Testudines: Testudinae). Herpetologica 30:417–419
Janzen FJ, Krenz JG, Haselkorn TS, Brodie ED Jr, Brodie ED III (2002) Molecular phylogeography of common garter snakes (Thamnophis sirtalis) in western North America: implications for regional historical forces. Mol Ecol 11:1739–1751
Kimura M, Ohta T (1978) Stepwise mutation model and distribution of allelic frequencies in a finite population. Proc Natl Acad Sci USA 75:2868–2872
King TL, Julian SE (2004) Conservation of microsatellite DNA flanking sequence across 13 emydid genera assayed with novel bog turtle (Glyptemys muhlenbergii) loci. Conserv Genet 5:719–725
Lee-Yaw JA, Irwin JT, Green DM (2008) Postglacial range expansion from northern refugia by the wood frog, Rana sylvatica. Mol Ecol 17:867–884
Lewis P, Zaykin D (2008) GDA v1.1 http://www.eeb.uconn.edu/people/plewis/software.php. Accessed Feb 2010
McGuire JM, Scribner KT, Congdon JD (2013) Spatial aspects of movements, mating patterns, and nest distributions influence gene flow among population subunits of Blanding’s turtles (Emydoidea blandingii). Conserv Genet. doi:10.1007/s10592-013-0493-8
Mickelson DM, Colgan PM (2003) The southern Laurentide Ice Sheet. Develop Quat Sci 1:1–16
Mockford SW, Snyder M, Herman TB (1999) A preliminary examination of genetic variation in a peripheral population of Blanding’s turtle, Emydoidea blandingii. Mol Ecol 8:323–327
Mockford SW, McEachern L, Herman TB, Synder M, Wright JM (2005) Population genetic structure of a disjunct population of Blanding’s turtle (Emydoidea blandingii) in Nova Scotia, Canada. Biol Conserv 123:373–380
Mockford SW, Herman TB, Snyder M, Wright JM (2007) Conservation genetics of Blanding’s turtle and its application in the identification of evolutionarily significant units. Conserv Genet 8:209–219
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Placyk JS, Burghardt GM Jr, Small RL, King RB, Casper GS, Robinson JW (2007) Post-glacial recolonization of Michigan by the common gartersnake (Thamnophis sirtalis) inferred from mtDNA sequences. Mol Phylogenet Evol 43:452–467
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rambaut A, Drummond AJ (2007) Tracer v1.4. http://beast.bio.ed.ac.uk/Tracer. Accessed Feb 2010
Raymond M, Rousset F (1995) GENEPOP (Version-1.2)—population-genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rhodin AGJ, van Dijk PP (2011) Emydoidea blandingii. In: IUCN 2012. IUCN Red List of Threatened Species. Version 2012.1 www.iucnredlist.org. Accessed 15 Aug 2012
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Rubin CS, Warner RE, Bouzat JL, Paige KN (2001) Population genetic structure of Blanding’s turtles (Emydoidea blandingii) in an urban landscape. Biol Conserv 99:323–330
Shaffer HB (2009) Turtles (Testudines). In: Hedges SB, Kumar S (eds) The timetree of life. Oxford University Press, Oxford, pp 398–401
Smith PW (1957) An analysis of post-Wisconsin biogeography of the prairie peninsula region based on distributional phenomena among terrestrial vertebrate populations. Ecology 38:205–218
Sommer RS, Lindqvist C, Persson A, Bringsoe H, Rhodin AGJ, Schneeweiss N, Siroky P, Bachmann L, Fritz U (2009) Unexpected early extinction of the European pond turtle (Emys orbicularis) in Sweden and climatic impact on its Holocene range. Mol Ecol 18:1252–1262
Spinks PQ, Shaffer HB (2009) Range-wide molecular analysis of the western pond turtle (Emys marmorata): cryptic variation, isolation by distance, and their conservation implications. Mol Ecol 14:2047–2064
Starkey DE, Shaffer HB, Burke RL, Forstner MRJ, Iverson JB, Janzen FJ, Rhodin AGJ, Ultsch GR (2003) Molecular systematics, phylogeography, and the effects of Pleistocene glaciation in the painted turtle (Chrysemys picta) complex. Evolution 57:119–128
Stiff BJ, Hansel AK (2004) Quaternary glaciations in Illinois. In: Ehlers J, Gibbard PL (eds) Quaternary glaciations: extent and chronology 2: Part II North America. Elsevier, Amsterdam, pp 71–82
Stuart SN, Chanson JS, Cox NA, Young BE, Rodrigues ASL, Fischman DL, Waller RW (2004) Status and trends of amphibian declines and extinctions worldwide. Science 306:1783–1786
Sutherland WJ, Aveling R, Bennun L et al (2012) A horizon scan of global conservation issues for 2012. Trends Ecol Evol 27:12–18
Ursenbacher S, Carlsson M, Helfer V, Tegelstrom H, Fumagalli L (2006) Phylogeography and Pleistocene refugia of the adder (Vipera berus) as inferred from mitochondrial DNA sequence data. Mol Ecol 15:3425–3437
Van Devender TR, King JE (1975) Fossil Blanding’s turtles, Emydoidea blandingii (Holbrook), and the late Pleistocene vegetation of western Missouri. Herpetologica 31:208–212
van Oosterhout C, Weetman D, Hutchinson WF (2006) Estimation and adjustment of microsatellite null alleles in nonequilibrium populations. Mol Ecol Notes 6:255–256
Walsh PS, Metzger DA, Higuchi R (1991) Chelex-100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10:506–513
Weir BS (1996) Genetic data analysis II. Sinauer, Sunderland
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population-structure. Evolution 38:1358–1370
Weisrock DW, Janzen FJ (2000) Comparative molecular phylogeography of North American softshell turtles (Apalone): implications for regional and wide-scale historical evolutionary forces. Mol Phylogenet Evol 14:152–164
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191
Wooding S, Ward RH (1997) Phylogeography and Pleistocene evolution in the North American black bear. Mol Biol Evol 14:1096–1105
You FM, Luo MC, Gu YQ, Lazo GR, Deal K, Dvorak J, Anderson OD (2007) GenoProfiler: batch processing of high-throughput capillary fingerprinting data. Bioinformatics 23:240–242
Acknowledgments
Thanks to Steve Dinkelacker, Robb Goldsberry, Chris Phillips, and Sara Ruane for assistance with collecting and to two anonymous reviewers for constructive criticisms. Turtles and tissues for this project were obtained under Iowa DNR Scientific Collecting permits to JLC and TJV, Illinois Endangered Species permits to FJJ and SH, Minnesota DNR permits to JMR, and a Nebraska Game & Parks Commission permit to Steve Dinkelacker. Funding for this research was provided in part by a contract from the McHenry County Conservation Foundation and National Science Foundation grants IBN-0080194, DEB-0089680, and DEB-0640932 to FJJ, and by a National Aeronautics and Space Administration Iowa Space Grant Consortium grant to FJJ and JLC. Additional funding for sequencing and bioinformatics support was provided by National Science Foundation DEB-0508665 to FJJ and EMM, as well as a grant from the EEOB Department at ISU and a James Cornette Fellowship to AS.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sethuraman, A., McGaugh, S.E., Becker, M.L. et al. Population genetics of Blanding’s turtle (Emys blandingii) in the midwestern United States. Conserv Genet 15, 61–73 (2014). https://doi.org/10.1007/s10592-013-0521-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-013-0521-8