Abstract
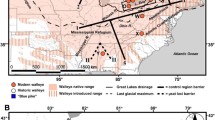
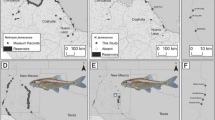
Redside dace Clinostomus elongatus (Teleostei: Cyprinidae) is a species of conservation concern that is declining throughout its range as a result of habitat fragmentation, degradation and loss. We characterized the genetic structure and diversity of redside dace populations across the species range using mitochondrial and microsatellite data to inform conservation efforts and assess how historical and recent events have shaped genetic structure and diversity within and among populations. Phylogeographic structure among 28 redside dace populations throughout southern Ontario (Canada) and the United States was assessed by sequence analysis of the mitochondrial cytochrome b and ATPase 6 and 8 genes. Populations were also genotyped using ten microsatellite loci to examine genetic diversity within and among populations as well as contemporary spatial structuring. Mitochondrial DNA sequence data revealed three geographically distinct lineages, which were highly concordant with groupings identified by microsatellite analysis. The combined genetic data refute published glacial refugia hypotheses of a single Mississippian refugium or of two lineages associated with Mississippian and Atlantic refugia. Secondary contact between the two eastern groups was documented in the Allegheny River drainage and tributaries to Lake Ontario. With the exception of several allopatric populations within the Allegheny watershed, high genetic structuring among populations suggests their isolation, indicating that recovery efforts should be population-based.






Similar content being viewed by others
References
April J, Turgeon J (2006) Phylogeography of the banded killifish (Fundulus diaphanus): glacial races and secondary contact. J Fish Biol 69:212–228
April J, Hanner RH, Dion-Côté AM, Bernatchez L (2013) Glacial cycles as an allopatric speciation pump in north-eastern American freshwater fishes. Mol Ecol 22:409–422
Avise JC (2000) Phylogeography. Harvard University Press, Massachusetts
Avise JC (2001) Cytonuclear genetic signatures of hybridization phenomena: rationale, utility, and empirical examples from fishes and other aquatic animals. Rev Fish Biol Fish 3:253–263
Bailey RM, Smith GR (1981) Origin and geography of the fish fauna of the Laurentian Great Lakes basin. Can J Fish Aquat Sci 38:1539–1561
Bandelt H, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Beausoleil JMJ, Doucet SM, Heath DD, Pitcher TE (2012) Spawning coloration, female choice and sperm competition in the redside dace, Clinostomus elongatus. Anim Behav 83:969–977
Beneteau CL, Mandrak NE, Heath DD (2009) The effects of river barriers and range expansion of the population genetic structure and stability in Greenside Darter (Etheostoma blennioides) populations. Conserv Genet 10:477–487
Beneteau CL, Walter RP, Mandrak NE, Heath DD (2012) Range expansion by invasion: genetic characterization of invasion of the greenside darter (Etheostoma blennioides) at the northern edge of its distribution. Biol Invasion 14:191–201
Berendzen PB, Simons AM, Wood RM (2003) Phylogeography of the northern hogsucker, Hypentelium nigricans (Teleostei: Cypriniformes): genetic evidence for the existence of the ancient Teays River. J Biogeogr 30:1139–1152
Berendzen PB, Dugan JF, Feltz JJ (2008) Establishing conservation units and population genetic parameters of fishes of greatest conservation need distributed in Southeast Minnesota. Final report for the State Wildlife Grants Program, Division of Ecological Resources, Minnesota Department of Natural Resources. Available via http://files.dnr.state.mn.us/eco/nongame/projects/consgrant_reports/2008/2008_berendzen_etal.pdf. Accessed 15 Sept 2017
Bernatchez L, Wilson CC (1998) Comparative phylogeography of Nearctic and Palearctic fishes. Mol Ecol 7:431–452
Bessert ML, Orti G (2003) Microsatellite loci for paternity analysis in the fathead minnow, Pimephales promelas (Teleostei: Cyprinidae). Mol Ecol Notes 3:532–534
Borden WC, Krebs RA (2009) Phylogeography and postglacial dispersal of smallmouth bass (Micropterus dolomieu) into the Great Lakes. Can J Fish Aquat Sci 66:2142–2156
Bossu CM, Beaulieu JM, Ceas PA, Near TJ (2013) Explicit tests of palaeodrainage connections of southeastern North America and the historical biogeography of Orangethroat Darters (Percidae: Etheostoma: Ceasia). Mol Ecol 22:5397–5417
COSEWIC (2007) COSEWIC assessment and updated status report on the Redside Dace Clinostomus elongatus in Canada. Committee on the Status of Endangered Wildlife in Canada. Ottawa, ON
COSEWIC (2014) Guidelines for recognizing designatable units. Committee on the status of endangered wildlife in Canada. Available at http://www.cosewic.gc.ca/default.asp?lang=en&n=DD31EAEE-1. Accessed 15 Sept 2017
Daniels RA, Wisniewski SJ (1994) Feeding ecology of redside dace, Clinostomus elongatus. Ecol Freshw Fish 3:176–183
Dimsoski P, Toth GP, Bagley MJ (2000) Microsatellite characterization in central stoneroller Campostoma anomalum (Pisces: Cyprinidae). Mol Ecol 9:2155–2234
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR (2014) NeEstimator v2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214
Dowling TE, Naylor GJP (1997) Evolutionary relationships of minnows in the genus Luxilus (Teleostei : Cyprinidae) as determined from cytochrome b sequences. Copeia 1997: 758–765
Dyke AS (2004) An outline of North American deglaciation with emphasis on central and northern Canada. In: Ehlers J, Gibbard PL (Eds.) Quaternary glaciations: extent and chronology: part II: North America. Elsevier, Amsterdam, pp 373–424
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Eliades N-G, Eliades DG (2009) HAPLOTYPE ANALYSIS: software for analysis of haplotypes data. http://www.uni-goettingen.de/en/134935.html. Accessed 15 Sept 2017
England PR, Luikart G, Waples RS (2010) Early detection of population fragmentation using linkage disequilibrium estimation of effective population size. Conserv Genet 11:2425–2430
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Frankham R, Briscoe DA, Ballou JD (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Galbraith HS, Zanatta DT, Wilson CC (2015) Comparative analysis of riverscape genetic structure in rare, threatened and common freshwater mussels. Conserv Genet 16:845–857
Galtier N, Nabholz B, Glémin S, Hurst GDD (2009) Mitochondrial DNA as a marker of molecular diversity: a reappraisal. Mol Ecol 18:4541–4550
George AL, Kuhajda BR, Williams JD, Cantrell MA, Rakes PL, Shute JR (2009) Guidelines for propagation and translocation for freshwater fish conservation. Fisheries 34:529–545
Ginson R, Walter RP, Mandrak NE, Beneteau CL, Heath D (2015) Hierarchical analysis of genetic structure in the habitat-specialist eastern sand darter (Ammocrypta pellucida). Ecol Evol 5:695–708
Harpending HC (1994) Signature of ancient population growth in a low-resolution mitochondrial DNA mismatch distribution. Hum Biol 66:591–600
Helfman GS (2007) Fish conservation: a guide to understanding and restoring global aquatic biodiversity and fishery resources. Island Press, Washington, DC
Herdendorf CE (2013) Research overview: holocene development of Lake Erie. Ohio J Sci 112:24–36
Hocutt CH, Wiley EO (1986) The zoogeography of North American freshwater fishes. Wiley, New York
Hocutt CH, Jenkins RE, Stauffer JR Jr (1986) Zoogeography of the fishes of the central Appalachians and central Atlantic coastal plain. In: Hocutt CH, Wiley EO (Eds.) The zoogeography of North American freshwater fishes. Wiley, New York, pp 161–211
Houston DD, Shiozawa DK, Riddle BR (2010) Phylogenetic relationships of the western North American cyprinid genus Richardsonius, with an overview of phylogeographic structure. Mol Phylogenet Evol 55:259–273
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Kalinowski ST (2005) HP-Rare 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol Ecol Notes 5:187–189
Larson G, Schaetzl R (2001) Origin and evolution of the Great Lakes. J Great Lakes Res 27:518–546
Lewis CM, Moore TC, Rea DK, Dettman DL, Smith AM, Mayer LA (1995) Lakes of the Huron basin: their record of runoff from the Laurentide Ice Sheet. Quat Sci Rev 13:891–922
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Litvak MK, Mandrak NE (1993) Ecology of freshwater baitfish use in Canada and the United States. Fisheries 18:6–13
MacKenzie DI, Nichols JD, Royle JA, Pollock KH (2006) Occupancy estimation and modeling: inferring patterns and dynamics of species occurrence. Elsevier, Burlington, MA
Maggs CA, Castilh R, Foltz D, Henzler C, Jolly MT, Kelly J, Olsen J, Perez KE, Stam W, Väinölä R, Viard F, Wares J (2008) Evaluating signatures of glacial refugia for North Atlantic benthic marine taxa. Ecology 89:S108–S122
Mandrak NE, Crossman E (1992) Postglacial dispersal of freshwater fishes into Ontario. Can J Zool 70:2247–2259
McCusker MR, Mandrak NE, Egeh B, Lovejoy NR (2014) Population structure and conservation genetic assessment of the endangered Pugnose Shiner, Notropis anogenus. Conserv Genet 15:343–353
McDermid JL, Wozney JK, Kjartanson SL, Wilson CC (2011) Quantifying historical, contemporary, and anthropogenic influences on the genetic structure and diversity of lake sturgeon (Acipenser fulvescens) populations in northern Ontario. J Appl Ichthyol 27:12–23
Moritz C (1994) Defining ‘evolutionarily significant units’ for conservation. Trends Ecol Evol 9:373–375
NatureServe (2015) NatureServe Explorer: an online encyclopedia of life [web application]. Version 7.1, Arlington, VA. Available via http://explorer.natureserve.org Accessed 24 July 2016
Near TL, Page LM, Mayden RL (2001) Intraspecific phylogeography of Percina evides (Percidae: Etheostomatinae): an additional test of the Central Highlands pre-Pleistocene vicariance hypothesis. Mol Ecol 10:2235–2240
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Nei M, Tajima F, Tateno Y (1983) Accuracy of estimated phylogenetic trees from molecular data. J Mol Evol 19:153–170
Novinger DC, Coon TG (2000) Behavior and physiology of the redside dace, Clinostomus elongatus, a threatened species in Michigan. Environ Biol Fish 57:315–326
Osborne MN, Diver TA, Turner TF (2013) Introduced populations as genetic reservoirs for imperiled species: a case study of the Arkansas River Shiner. Conserv Genet 14:637–647
Overpeck JT, Webb RS, Webb T (1992) Mapping eastern North American vegetation change of the past 18 ka: No-analogs and the future. Geology 20:1071–1074
Page LM, Burr BM (2011) Peterson field guide to freshwater fishes of North America North of Mexico. 2nd edn., Houghton Mifflin Harcourt, Boston
Parker BJ, McKee P, Campbell RR (1988) Status of the redside dace, Clinostomus elongatus, in Canada. Can Field Nat 102:163–169
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pitcher TW, Beneteau CL, Walter RP, Wilson CC, Mandrak NE, Heath DD (2009) Isolation and characterization of microsatellite loci in the redside dace, Clinostomus elongatus. Conserv Genet Resour 1:381–383
Poos MS, Jackson DA (2012) Impact of species-specific dispersal and regional stochasticity on estimates of population viability in stream metapopulations. Landsc Ecol 27:405–416
Poos MS, Lawrie D, Tu C, Jackson DA, Mandrak N (2012) Estimating local and regional population sizes for an endangered minnow, redside dace (Clinostomus elongatus), in Canada. Aquat Conserv Mar Freshw Ecosyst 22:47–57
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ray JM, Wood RM, Simons AM (2006) Phylogeography and post-glacial colonization patterns of the rainbow darter, Etheostoma caeruleum (Teleostei: Percidae). J Biogeogr 33:1550–1558
Raymond M, Rousset F (1995) GENEPOP (Version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Redside Dace Recovery Team (2010) Recovery strategy for redside dace (Clinostomus elongatus) in Ontario. Ontario Ministry of Natural Resources, Peterborough, ON
Reid SM, Yunker G, Jones N (2009) Evaluation of single-pass electrofishing for stream fish community monitoring. Fish Manag Ecol 16:1–9
Reid SM, Kidd A, Wilson CC (2012) Validation of buccal swabs for noninvasive DNA sampling of small-bodied imperiled fishes. J Appl Ichthyol 28:290–292
Rice WR (1989) Analyzing tables of statistical tests. Evol Int J Org Evol 43:223–225
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schmidt RE (1986) Zoogeography of the Northern Appalachians. In: Hocutt CH, Wiley EO (Eds.) The Zoogeography of North American freshwater fishes. Wiley, New York, pp 137–159
Scott WB, Crossman EJ (1973) Freshwater fishes of Canada. Bull Fish Res Board Can 184:1–966
Simon TP, Burskey JL (2014) Spatial distribution and dispersal patterns of central North American freshwater crayfish (Decapoda: Cambaridae) with emphasis on implications of glacial refugia. Int J Biodivers. doi:10.1155/2014/282079
Sivasundar A, Bermingham E, Orti G (2001) Population structure and biogeography of migratory freshwater fishes (Prochilodus: Characiformes) in major South American rivers. Mol Ecol 10:407–417
Smith CL (1985) The inland fishes of New York State. New York State Department of Environmental Conservation, Albany, NY
Soltis DE, Morris AB, McLachlan JS, Manos PS, Soltis PS (2006) Comparative phylogeography of unglaciated eastern North America. Mol Ecol 15:4261–4293
Sweeten J (2012) Redside dace (Clinostomus elongatus) in Mill Creek, Wabash County, Indiana: a strategy for research and augmentation. Indiana Department of Natural Resources, Wabash County, IN
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Takezaki N, Nei M, Tamura K (2010) POPTREE2: Software for constructing population trees from allele frequency data and computing other population statistics with Windows interface. Mol Biol Evol 27:747–752
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Todd TN, Hatcher CO (1993) Genetic variability and glacial origins of yellow perch (Perca flavescens) in North America. Can J Fish Aquat Sci 50:1828–1834
Underhill JC (1986) The fish fauna of the Laurentian Great Lakes, the St. Lawrence lowlands, Newfoundland and Labrador. In: Hocutt CH, Wiley EO (eds) The zoogeography of North American freshwater fishes. Wiley, New York, pp 105–136
Van Oosterhout C, Hutchinson WF, Willis DP, Shipley P (2004) MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Vélez-Espino LA, Koops M (2008). Recovery potential modelling of redside dace (Clinostomus elongatus) in Canada. Canadian Science Advisory Secretariat. Research Document 2008/005. Fisheries and Oceans Canada, Ottawa, ON
Ver Steeg K (1946) The Teays River. Ohio J Sci 46:297–307
Waples RS (2006) Distinct population segments. In: Goble DD, Davis FW, Michael Scott JM (Eds.) The endangered species act at thirty: conserving biodiversity in human-dominated landscapes. Vol. 2, Island Press, Washington, pp 127–149
Waples RS, Do C (2010) Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl 3:244–262
Wilson CC, Hebert PD (1998) Phylogeography and postglacial dispersal of lake trout (Salvelinus namaycush) in North America. Can J Fish Aquat Sci 55:1010–1024
Wozney KM, Haxton TJ, Kjartanson S, Wilson CC (2011) Genetic assessment of lake sturgeon (Acipenser fulvescens) population structure in the Ottawa River. Environ Biol Fishes 90:183–195
Acknowledgements
Funding for this project was provided by the Ontario Ministry of Natural Resources and Forestry (OMNRF) and Fisheries and Oceans Canada. Samples were generously provided by Jeff Anderson (Conservation Halton), Douglas Carlson (New York State Department of Environmental Conservation), Jon Clayton (Credit Valley Conservation), Konrad Schmidt and Jenny Kruckenberg (North American Native Fishes Association), Holly Jennings and John Pagel (USDA Forest Service, Michigan), Brant Fisher (Indiana Department of Natural Resources), Brian Zimmerman (Ohio State University), David Thorne and Isaac Gibson (West Virginia Division of Natural Resources), John Lyons (Wisconsin Department of Natural Resources), Nate Tessler (EnviroScience, Inc.), David Miko (Pennsylvania Fish and Boat Commission), and Matthew Thomas (Kentucky Department of Fish and Wildlife Resources). Aaron Clauser (Clauser Environmental), Wayne Starnes (North Carolina State Museum of Natural Sciences), Aaron Snell (Streamside Ecological Services, Inc), and Douglas Fischer (Pennsylvania Fish and Boat Commission) also provided valuable advice and feedback. Lab work assistance was provided by Caleigh Smith, Kristyne Smith, and Anne Kidd. Alan Dextrase (OMNRF) and Joanna Freeland (Trent University) and two anonymous reviewers provided constructive comments on earlier drafts of the manuscript
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Serrao, N.R., Reid, S.M. & Wilson, C.C. Conservation genetics of redside dace (Clinostomus elongatus): phylogeography and contemporary spatial structure. Conserv Genet 19, 409–424 (2018). https://doi.org/10.1007/s10592-017-1012-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-017-1012-0