Abstract
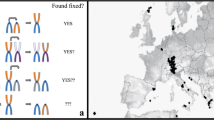
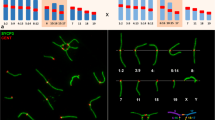
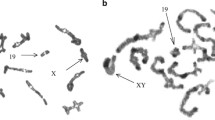
Within species, populations differing by chromosomal rearrangements (“chromosomal races”) may become reproductively isolated in association with reduced hybrid fertility due to meiotic aberrations. Speciation is also possible if hybridizing chromosomal races accumulate genetic differences because of reduced meiotic recombination in the heterozygous configuration in hybrids. Here, we examine recombination in pure races and hybrids within a model system for chromosomal speciation: the hybridization of the Poschiavo (CHPO) and Upper Valtellina (IUVA) chromosomal races of house mouse in Upper Valtellina, Italy. These races differ by Robertsonian fusions/whole-arm reciprocal translocations, such that hybrids produce a pentavalent meiotic configuration. We determined the number and position of the recombination points (using an antibody against the MutL homolog 1 [MLH1] protein) on synaptonemal complexes at pachytene in laboratory-reared CHPO, IUVA, and hybrid males, analyzing at least 112 spermatocytes per karyotypic group, up to a total of 534 spermatocytes. The mean ± standard deviation numbers of MLH1 foci per spermatocyte were 22.2 ± 3.2, 20.1 ± 2.9, 20.7 ± 2.3, and 21.9 ± 2.9 for CHPO, IUVA, CHPO × IUVA, and IUVA × CHPO, respectively. Altogether, 10,146 chromosome arms were examined, allowing multiple comparisons. Overall, recombination events were more frequently distal than proximal or interstitial. The average number of proximal MLH1 foci per chromosome arm decreased going from telocentric to metacentric bivalents to pentavalents (when present), which (together with other factors) influenced the average number of MLH1 foci per cell between CHPO, IUVA, and hybrid mice. The low frequency of proximal recombination in pentavalents of CHPO–IUVA hybrids may promote reproductive isolation between the CHPO and IUVA races, when coupled with reduced hybrid unfitness.


Similar content being viewed by others
Abbreviations
- CHPO:
-
Poschiavo [chromosomal race]
- CIII:
-
Chain-of-three
- CV:
-
Chain-of-five
- CREST:
-
Calcinosis, Raynaud's syndrome, esophageal dysmotility, sclerodactyly, telangiectasia
- DAPI:
-
4′,6-diamidino-2-phenylindole
- F0 :
-
Parental generation
- F1 :
-
First generation
- IUVA:
-
Upper Valtellina [chromosomal race]
- MLH1:
-
MutL homolog 1
- PBS:
-
Phosphate-buffered saline
- PBT:
-
Phosphate-buffered saline plus Tween-20
- Rb:
-
Robertsonian
- SC:
-
Synaptonemal complex
- SD:
-
Standard deviation
- SYCP3:
-
Synaptonemal complex protein 3
- WARTs:
-
Whole-arm reciprocal translocations
References
Anderson LK, Reeves A, Webb LM, Ashley T (1999) Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151:1569–1579
Baker RJ, Bickham JW (1986) Speciation by monobrachial centric fusions. Proc Natl Acad Sci U S A 83:8245–8248
Baker SM, Plug AW, Prolla TA, Bronner CE, Harris AC, Yao X, Christie DM, Monell C, Arnheim M, Braley A, Ashley T, Liskay RM (1996) Involvement of mouse Mlh1 in DNA mismatch repair and meiotic crossing over. Nat Genet 13:336–342
Barlow AL, Hultén MA (1998) Crossing over analysis at pachytene in man. Eur J Hum Genet 6:350–358
Bidau CJ (1993) Causes of chiasma repatterning due to centric fusions. Rev Brasil Genet 16:283–296
Bidau CJ, Giménez MD, Palmer CL, Searle JB (2001) The effects of Robertsonian fusions on chiasma frequency and distribution in the house mouse (Mus musculus domesticus) from a hybrid zone in northern Scotland. Heredity 87:305–313
Bonhomme F, Searle JB (2012) House mouse phylogeography. In: Macholán M, Baird SJE, Munclinger P, Piálek J (eds) Evolution in the house mouse. Cambridge University Press, Cambridge, pp 278–296
Borodin PM, Karamysheva TV, Belonogova NM, Torgasheva AA, Rubtsov NB, Searle JB (2008) Recombination map of the common shrew, Sorex araneus (Eulipotyphla, Mammalia). Genetics 178:621–632
Britton-Davidian J, Nadeau JH, Croset H, Thaler L (1989) Genic differentiation and origin of Robertsonian populations of the house mouse (Mus musculus domesticus Rutty). Genet Res 53:29–44
Capanna E (1982) Robertsonian numerical variation in animal speciation: Mus musculus, an emblematic model. In: Barigozzi C (ed) Mechanisms of speciation. Liss, New York, pp 155–177
Capanna E, Corti M (1982) Reproductive isolation between two chromosomal races of Mus musculus in the Rhaetian Alps (Northern Italy). Mammalia 46:107–109
Capanna E, Redi CA (1995) Whole-arm reciprocal translocation (WART) between Robertsonian chromosomes: finding of a Robertsonian heterozygous mouse with karyotype derived through WARTs. Chromosom Res 3:135–137
Castiglia R, Capanna E (2002) Chiasma repatterning across a chromosomal hybrid zone between chromosomal races of Mus musculus domesticus. Genetica 114:35–40
Castiglia R, Annesi F, Capanna E (2002) Contact zones between chromosomal races of Mus musculus domesticus: III. Molecular and chromosomal evidence of restricted gene flow between the CD race (2n = 22) and the ACR race (2n = 24). Heredity 89:219–224
Dumas D, Britton-Davidian J (2002) Chromosomal rearrangements and evolution of recombination: comparison of chiasma distribution patterns in standard and Robertsonian populations of the house mouse. Genetics 162:1355–1366
Dumont BL, Payseur BA (2011) Evolution of the genomic recombination rate in murid rodents. Genetics 187:643–657
Feder JL, Egan SP, Nosil P (2012) The genomics of speciation-with-gene-flow. Trends Genet 28:342–350
Franchini P, Castiglia R, Capanna E (2008) Reproductive isolation between chromosomal races of the house mouse Mus musculus domesticus in a parapatric contact area revealed by an analysis of multiple unlinked loci. J Evol Biol 21:502–513
Garagna S, Broccoli D, Redi CA, Searle JB, Cooke HJ, Capanna E (1995) Robertsonian metacentrics of the house mouse lose telomeric sequences but retain some minor satellite DNA in the pericentromeric area. Chromosoma 103:685–692
Garagna S, Marziliano N, Zuccotti M, Searle JB, Capanna E, Redi CA (2001) Pericentromeric organization at the fusion point of mouse Robertsonian translocation chromosomes. Proc Natl Acad Sci U S A 98:171–175
Giménez MD, White TA, Hauffe HC, Panithanarak T, Searle JB (2013) Understanding the basis of diminished gene flow between hybridizing chromosome races of the house mouse. Evolution 67:1446–1462
Hauffe HC, Searle JB (1992) A disappearing speciation event? Nature 357:26
Hauffe HC, Searle JB (1993) Extreme karyotypic variation in a Mus musculus domesticus hybrid zone: the tobacco mouse story revisited. Evolution 47:1374–1395
Hauffe HC, Searle JB (1998) Chromosomal heterozygosity and fertility in house mice (Mus musculus domesticus) from Northern Italy. Genetics 150:1143–1154
Hauffe HC, Giménez MD, Searle JB (2012) Chromosomal hybrid zones in the house mouse. In: Macholán M, Baird SJE, Munclinger P, Piálek J (eds) Evolution in the house mouse. Cambridge University Press, Cambridge, pp 407–430
Hultén MA (2011) On the origin of crossover interference: a chromosome oscillatory movement (COM) model. Mol Cytogenet 4:10
Jones GH, Franklin FC (2006) Meiotic crossing-over: obligation and interference. Cell 126:246–248
King M (1993) Species evolution: the role of chromosome change. Cambridge University Press, Cambridge
Lercher MJ, Hurst LD (2003) Imprinted chromosomal regions of the human genome have unusually high recombination rates. Genetics 165:1629–1632
Li W, Fann CS, Ott J (1998) Low-order polynomial trends of female-to-male map distance ratios along human chromosomes. Hum Hered 48:266–270
Manterola M, Page J, Vasco C, Berríos S, Parra MT, Viera A, Rufas JS, Zuccotti M, Garagna S, Fernández-Donoso R (2009) A high incidence of meiotic silencing of unsynapsed chromatin is not associated with substantial pachytene loss in heterozygous male mice carrying multiple simple Robertsonian translocations. PLoS Genet 5:e1000625
Matveevsky SN, Pavlova SV, Acaeva MM, Kolomiets OL (2012) Synaptonemal complex analysis of interracial hybrids between the Moscow and Neroosa chromosomal races of the common shrew Sorex araneus showing regular formation of a complex meiotic configuration (ring-of-four). Comp Cytogenet 6:301–314
Merico V, Pigozzi MI, Esposito A, Merani MS, Garagna S (2003) Meiotic recombination and spermatogenic impairment in Mus musculus domesticus carrying multiple simple Robertsonian translocations. Cytogenet Genome Res 103:321–329
Merico V, de Barboza GD, Vasco C, Ponce R, Rodriguez V, Garagna S, Tolosa de Talamoni N (2008) A mitochondrial mechanism is involved in apoptosis of Robertsonian mouse male germ cells. Reproduction 135:797–804
Moses MJ (1980) New cytogenetic studies on mammalian meiosis. In: Serio M, Martini L (eds) Animal models in human reproduction. Raven, New York, pp 169–190
Nachman MW, Boyer SN, Searle JB, Aquadro CF (1994) Mitochondrial DNA variation and the evolution of Robertsonian chromosomal races of house mice, Mus domesticus. Genetics 136:1105–1120
Ng SH, Madeira R, Parvanov ED, Petros LM, Petkov PM, Paigen K (2009) Parental origin of chromosomes influences crossover activity within the Kcnq1 transcriptionally imprinted domain of Mus musculus. BMC Mol Biol 13:10–43
Panithanarak T, Hauffe HC, Dallas JF, Glover A, Ward RG, Searle JB (2004) Linkage-dependent gene flow in a house mouse chromosomal hybrid zone. Evolution 58:184–192
Peters AH, Plug AW, van Vugt MJ, de Boer P (1997) A drying-down technique for the spreading of mammalian meiocytes from the male and female germline. Chromosom Res 5:66–68
Piálek J, Hauffe HC, Rodríguez-Clark KM, Searle JB (2001) Raciation and speciation in house mice from the Alps: the role of chromosomes. Mol Ecol 10:613–625
Piálek J, Hauffe HC, Searle JB (2005) Chromosomal variation in the house mouse. Biol J Linn Soc 84:535–563
Pinho C, Hey J (2010) Divergence with gene flow: models and data. Ann Rev Ecol Evol Syst 41:215–230
Robinson WP, Lalande M (1995) Sex-specific meiotic recombination in the Prader–Willi/Angelman syndrome imprinted region. Hum Mol Genet 4:801–806
Rieseberg LH (2001) Chromosomal rearrangements and speciation. Trends Ecol Evol 16:351–358
Schramm S, Fraune J, Naumann R, Hernandez-Hernandez A, Höög C, Cooke HJ, Alsheimer M, Benavente R (2011) A novel mouse synaptonemal complex protein is essential for loading of central element proteins, recombination, and fertility. PLoS Genet 7:e1002088
Searle JB (1993) Chromosomal hybrids in eutherian mammals. In: Harrison RG (ed) Hybrid zones and the evolutionary process. Oxford University Press, New York, pp 309–353
Smadja CM, Butlin RK (2011) A framework for comparing processes of speciation in the presence of gene flow. Mol Ecol 20:5123–5140
Tease C (1998) Chiasma distributions and chromosome segregation in male and female translocation heterozygous mice analysed using FISH. Chromosoma 107:549–558
Tease C, Hultén MA (2004) Inter-sex variation in synaptonemal complex lengths largely determine the different recombination rates in male and female germ cells. Cytogenet Genome Res 107:208–215
Vasco C, Manterola M, Page J, Zuccotti M, de la Fuente R, Redi CA, Fernandez-Donoso R, Garagna S (2012) The frequency of heterologous synapsis increases with aging in Robertsonian heterozygous male mice. Chromosom Res 20:269–278
Wallace BMN, Searle JB, Everett CA (2002) The effect of multiple simple Robertsonian heterozygosity on chromosome pairing and fertility of wild-stock house mice (Mus musculus domesticus). Cytogenet Genome Res 96:276–286
White MJD (1978a) Modes of speciation. Freeman, San Francisco
White MJD (1978b) Chain processes in chromosomal speciation. Syst Zool 27:285–298
Acknowledgments
We acknowledge the funding from the Natural Environment Research Council. We thank the two anonymous reviewers for their very helpful comments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Fengtang Yang.
Valeria Merico, Mabel D. Giménez, Chiara Vasco, Jeremy B. Searle, Heidi C. Hauffe, and Silvia Garagna contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 111 kb)
Rights and permissions
About this article
Cite this article
Merico, V., Giménez, M.D., Vasco, C. et al. Chromosomal speciation in mice: a cytogenetic analysis of recombination. Chromosome Res 21, 523–533 (2013). https://doi.org/10.1007/s10577-013-9377-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-013-9377-5