Abstract
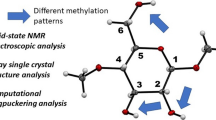
We investigate theoretically the NMR response of twisted configurations of \({\rm I}\beta\) cellulose in the tg conformation. These finite helical angle structures were constructed by a mathematical deformation of zero-angle configurations obtained via the periodic density functional energy minimizations with dispersion corrections (DFT-D2). Subsequent calculations of the \({^{13}\hbox {C}}\) nuclear magnetic resonance chemical shifts \(({\delta}^{13} \hbox {C})\) were compared with experimental findings by Erata et al. (Cellul Commun 4:128–131, 1997) and Kono et al. (Macromolecules 36:5131–5138, 2003). We determine the sensitivity of the NMR chemical shifts to helical deformation of the microfibril and find that a substantial range of helical angle, ±2 degrees/nm, is consistent with current experimental observations, with a most probable angle of ∼0.2 degree/nm. Through exhaustive combinatorial provisional assignments, we also demonstrate that there are different choices of the chemical shift \(({\delta}^{13} \hbox {C})\) assignments which are consistent with the experiments, including ones with lower deviations than existing identifications.









Similar content being viewed by others
References
Aabloo A, French AD, Mikelsaar RH, Pertsin AJ (1994) Studies of crystalline native celluloses using potential energy calculations. Cellulose 1:161–168
Adamo C, Barone V (1998) Introduction I exchange functionals with improved long-range behavior and adiabatic connection methods without adjustable parameters: the mPW and mPW1PW models. J Chem Phys 108:664–675
Atalla RH, VanderHart DL (1984) Native cellulose: a composite of two distinct crystalline forms. Science 223:283–285
Bachrach SM (2007) Quantum mechanics for organic chemistry. In: Computational organic chemistry, New York, p 811
Buhl M, Kaupp M, Malkina OL, Malkin VG (1999) The DFT route to NMR chemical shifts. J Comput Chem 20:91–105
Cheeseman JR, Trucks GW, Keith TA, Frisch MJ (1996a) A comparison of models for calculating nuclear magnetic resonance shielding tensors. J Chem Phys 104:5497–5509
Cheeseman JR, Trucks GW, Keith TA, Frisch MJ (1996) A comparison of models for calculating nuclear magnetic resonance shielding tensors. J Chem Phys 104:5497–5509
Clark T, Chandrasekhar J, Spitznagel GW, Schleyer PVR (1983) Efficient diffuse function-augmented basis sets for anion calculations. III. The 3-21+G basis set for first-row elements, Li-F. J Comput Chem 4:294–301. doi:10.1002/jcc.540040303
Erata T, Shikano T, Yunoki S, Takai M (1997) The complete assignment of the 13C CP/MAS NMR spectrum of native cellulose by using 13C labeled glucose. Cellul Commun 4:128–131
Esrafili MD, Ahmadin H (2012) DFT study of \(^{17}\text{O}\), \(^1\text{H}\) and \(^{13}\text{C}\) NMR chemical shifts in two forms of native cellulose, \(\text{I}\alpha\) and \(\text{I}\beta\). Carbohydr Res 347:99–106
Fernandes AN, Thomas LH, Altaner CM, Callow P, Forsyth VT, Apperley DC, Kennedy CJ, Jarvis MC (2011) Nanostructure of cellulose microfibrils in spruce wood. PNAS 108(47):E1195–E1203
French AD, Johnson GP (2009) Cellulose and the twofold screw axis: modeling and experimental arguments. Cellulose 16:959–973
French AD, Johnson GP (2004a) What crystals of small analogs are trying to tell us about cellulose structure. Cellulose 11:5–22
French AD, Johnson GP (2004b) Advanced conformational energy surfaces for cellobiose. Cellulose 11:449–462
French AD, Johnson GP (2006) Quantum mechanics studies of cellobiose conformations. Can J Chem 84:603–612
Frisch MJ, Trucks GW, Schlegel HB, et al (2009) Gaussian 09 revision B.01. Wallingford, CT
Gray DG (1996) Chirality in cellulose and cellulose-based materials. Polym Prepr (Div Polym Sci Am Chem Soc) 37:485–486
Gottlieb HE, Kotlyar V, Nudelman A (1997) NMR chemical shifts of common laboratory solvents as trace impurities. J Org Chem 62:7512–7515
Hadden JA, French AD, Woods RJ (2013) Unraveling cellulose microfibrils: a twisted tale. Biopolymers (Special Issue: 50th Anniversary Special Issue on Glycosciences) 99(10):746–756
Haigler CH, White AR, Brown RM, Cooper KM (1982) Alteration of invivo cellulose ribbon assembly by carboxymethylcellulose and other cellulose derivatives. J Cell Biol 94:64–69
Hanley SJ, Revol J-F, Godbout L, Gray DG (1997) Atomic force microscopy and transmission electron microscopy of cellulose from Micrasterias denticulata; evidence for a chiral helical microfibril twist. Cellulose 4:209–220
Hirai A, Tuji M, Horii F (1998) Helical sense of ribbon assemblies and splayed microfibrils of bacterial cellulose. SenI Gakkaishi 54:506–510
Hohenberg P, Kohn W (1964) Inhomogeneous electron gas. Phys Rev 136:B864–B871
Karadakov PB (2006) Ab initio calculation of NMR shielding constants. In: Webb GA (ed) Modern magnetic resonance. Springer, The Netherlands, pp 63–70
Kohn W, Sham LJ (1965) Self-consistent equations including exchange and correlation effects. Phys Rev 140:A1133–A1138
Kono H, Erata T, Takai M (2003) Determination of the through-bond carbon-carbon and carbon-proton connectivities of the native celluloses in the solid state. Macromolecules 36:5131–5138
Krishnan R, Brinkley JS, Seeger R, Pople JA (1980) Self-consistent molecular orbital methods. XX. A basis set for correlated wave functions. J Chem Phys 72:650–654
Kubicki JD, Mohamed MN-A, Watts HD (2013a) Quantum mechanical modeling of the structures, energetics and spectral properties of \(\text{I}\alpha\) and \(\text{I}\beta\) cellulose. Cellulose 20:9–23. doi: 10.1007/s10570-012-9838-6
Kubicki JD, Watts HD, Zhao Z, Zhong L (2013b) Quantum mechanical calculations on cellulosewater interactions: structures, energetics, vibrational frequencies and NMR chemical shifts for surfaces of \(\text{I}\alpha\) and \(\text{I}\beta\) cellulose. Cellulose 118. doi:10.1007/s10570-013-0029-x
Matthews JF, Skopec CE, Mason PE, Zuccato P, Torget RW, Sugiyama J, Himmel ME, Brady JW (2006) Computer simulation studies of microcrystalline cellulose I beta. Carbohydr Res 341:138–152
Nimlos MR, Matthews JF, Crowley MF, Walker RC, Chukkapalli G, Brady JW, Adney WS, Cleary JM, Zhong L, Himmel ME (2007) Molecular modeling suggests induced fit of Family I carbohydrate-binding modules with a broken-chain cellulose surface. Protein Eng Des Sel 20:179–187
Nishiyama Y, Langan P, Chanzy H (2002) Crystal structure and hydrogen-bonding system in cellulose \(\text{I}\beta\) from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 124(31):9074–9082
Nishiyama Y, Sugiyama J, Chanzy H, Langan P (2003) Crystal structure and hydrogen bonding system in cellulose Ia from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 125(47):14300–14306
Paavilainen S, Rog T, Vattulainen I (2011) Analysis of twisting of cellulose nanofibrils in atomistic molecular dynamics simulations. J Phys Chem B 115:3747–3755
Papajak E, Zheng J, Xu X et al (2011) Perspectives on basis sets beautiful: seasonal plantings of diffuse basis functions. J Chem Theory Comput 7:3027–3034
Perdew J, Chevary J, Vosko S (1992) Atoms, molecules, solids, and surfaces: applications of the generalized gradient approximation for exchange and correlation. Phys Rev B 46:6671–6687
Qian X (2008) The effect of cooperativity on hydrogen bonding interactions in native cellulose \(\text{I}\beta\) from ab initio molecular dynamics simulations. Mol Simul 34(2):183–191
Sarotti AM, Pellegrinet SC (2009) A multi-standard approach for GIAO (13)C NMR calculations. J Org Chem 74:7254–7260
Schreckenbach G, Ziegler T (1995) Calculation of NMR shielding tensors using gauge-including atomic orbitals and modern density functional theory. J Phys Chem 99:606611. doi:10.1021/j100002a024
Sternberg U, Koch F-T, Priess W, Witter R (2003) Crystal structure refinements of cellulose polymorphs using solid- state \(^{13}\text{C}\) chemical shifts. Cellulose 10:189–199
Taylor RE, French AD, Gamble GR, Himmelsbach DS, Stipanovic RD, Thibodeaux DP, Wakelyn PJ, Dybowski C (2008) \(^1\text{H}\) and \(^{13}\text{C}\) solid-state NMR of gossypium barbadense (Pima) cotton. J Mol Struct 878:177–184
VanderHart DL, Atalla RH (1984) Studies of macrostructure in native cellulose using solid-state \(^{13}\text{C}\) NMR. Macromolecules 17(8):1465–1472
Watts H, Mohamed M, Kubicki J (2011) Comparison of multistandard and TMS-standard calculated NMR shifts for coniferyl alcohol and application of the multistandard method to lignin dimers. J Phys Chem B 115:1958–1970
Watts HD, Mohamed MNA, Kubicki JD (2014) A DFT study of vibrational frequencies and 13C NMR chemical shifts of model cellulosic fragments as a function of size. Cellulose 21(1):53–70
Wolinski K, Hinton JF, Pulay P (1990) Efficient implementation of the gauge-independent atomic orbital method for NMR chemical shift calculations. J Am Chem Soc 112:8251–8260. doi:10.1021/ja00179a005
Yui T, Nishimura S, Akiba S, Hayashi S (2006) Swelling behavior of the cellulose \(\text{I}\beta\) crystal models by molecular dynamics. Carbohydr Res 341:2521–2530
Yui T, Hayashi S (2007) Molecular dynamics simulations of solvated crystal models of cellulose \(\text{I}_{\alpha}\) and \(\text{IIII}_I\). Biomacromolecules 8:817–824
Yui T, Hayashi S (2009) Structural stability of the solvated cellulose \(\text{IIII}_I\) crystal models: a molecular dynamics study. Cellulose 16:151–165
Zhao Z, Shklyaev OE, Nili A, Mohamed MN-A, Kubicki JD, Crespi VH, Zhong L (2013) Cellulose microfibril twist, mechanics, and implication for cellulose biosynthesis. J Phys Chem A 117:2580–2589
Acknowledgments
This work was supported as part of The Center for LignoCellulose Structure and Formation, an Energy Frontier Research Center funded by the U.S. Department of Energy, Office of Science, Basic Energy Sciences under Award # DE-SC0001090. Computational support was provided by the Research Computation and Cyberinfrastructure group at The Pennsylvania State University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shklyaev, O.E., Kubicki, J.D., Watts, H.D. et al. Constraints on \({\rm I}\beta\) cellulose twist from DFT calculations of \(^{13}\hbox {C}\) NMR chemical shifts. Cellulose 21, 3979–3991 (2014). https://doi.org/10.1007/s10570-014-0448-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-014-0448-3