Abstract
Purpose
The identification of biomarkers of hormonal therapy (HT) failure would allow tailored monitoring in metastatic breast cancer (mBC) patients. PIK3CA gene mutation is one of the most frequent events in mBC and is associated with HT resistance. We evaluated the early prognostic value of cell-free DNA (cfDNA) PIK3CA detection in first-line HT-treated mBC patients.
Methods
Between June 2012 and January 2014, 39 patients were prospectively included in a dedicated clinical trial (NCT01612871). Blood sampling was performed before (M0) and 4 weeks (M1), 3 months (M3) and 6 months (M6) after HT initiation, and at tumor progression. Patients were followed until progression or until the end of the study (2 years). Mutation detection was performed using droplet-based digital PCR (ddPCR). Progression-free survival (PFS) was used as primary endpoint.
Results
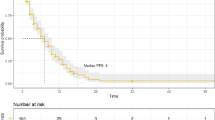
Median age at inclusion was 63 years (range 40–86). Most patients (34/39) received an aromatase inhibitor and presented a non-measurable disease (71.8%). PIK3CA mutations were reported in 10 (27.8%) and 5 (14.3%) cases at M0 and M1, respectively. The persistence of a detectable circulating mutation at M1 was highly correlated with a worse progression-free survival (PFS), rate at 1 year: 40% versus 76.7%; p = 0.0053).
Conclusions
Four-week persistence of cfDNA PIK3CA mutation appears highly correlated with PFS.
Trial registration
NCT01612871, registered on June 6th, 2012; https://clinicaltrials.gov/ct2/show/NCT01612871.


Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the Department of Biostatistics, Institut Claudius Regaud, IUCT-O Toulouse, France, upon reasonable request.
Abbreviations
- BC:
-
Breast cancer
- cfDNA:
-
Cell-free DNA
- CDK:
-
Cyclin-dependent kinase
- ER+:
-
Estrogen-receptor-positive
- HT:
-
Hormonal therapy
- mBC:
-
Metastatic breast cancer
- mTOR:
-
Mammalian target of rapamycin
- PI3K:
-
Phosphatidylinositol 3-kinase
- PI3KCA:
-
PI3K catalytic subunit alpha
- PFS:
-
Progression-free survival
- RB:
-
Retinoblastoma
- 95% CI:
-
95% confidence interval
References
Provenzano A, Kurian S, Abraham J (2013) Overcoming endocrine resistance in breast cancer: role of the PI3 K and the mTOR pathways. Expert Rev Anticancer Ther 13(2):143–147. https://doi.org/10.1586/era.12.173
Fu X, Osborne CK, Schiff R (2013) Biology and therapeutic potential of PI3 K signaling in ER+/HER2-negative breast cancer. Breast 22(Suppl 2):S12–18. https://doi.org/10.1016/j.breast.2013.08.001
Lopez-Knowles E, Segal CV, Gao Q, Garcia-Murillas I, Turner NC, Smith I, Martin LA, Dowsett M (2014) Relationship of PIK3CA mutation and pathway activity with antiproliferative response to aromatase inhibition. Breast Cancer Res 16(3):R68. https://doi.org/10.1186/bcr3683
Ramirez-Ardila DE, Helmijr JC, Look MP, Lurkin I, Ruigrok-Ritstier K, van Laere S, Dirix L, Sweep FC, Span PN, Linn SC, Foekens JA, Sleijfer S, Berns EM, Jansen MP (2013) Hotspot mutations in PIK3CA associate with first-line treatment outcome for aromatase inhibitors but not for tamoxifen. Breast Cancer Res Treat 139(1):39–49. https://doi.org/10.1007/s10549-013-2529-7
Lai K, Killingsworth MC, Lee CS (2015) Gene of the month: PIK3CA. J Clin Pathol 68(4):253–257. https://doi.org/10.1136/jclinpath-2015-202885
Araki K, Miyoshi Y (2018) Mechanism of resistance to endocrine therapy in breast cancer: the important role of PI3 K/Akt/mTOR in estrogen receptor-positive, HER2-negative breast cancer. Breast Cancer 25(4):392–401. https://doi.org/10.1007/s12282-017-0812-x
Higgins MJ, Jelovac D, Barnathan E, Blair B, Slater S, Powers P, Zorzi J, Jeter SC, Oliver GR, Fetting J, Emens L, Riley C, Stearns V, Diehl F, Angenendt P, Huang P, Cope L, Argani P, Murphy KM, Bachman KE, Greshock J, Wolff AC, Park BH (2012) Detection of tumor PIK3CA status in metastatic breast cancer using peripheral blood. Clin Cancer Res 18(12):3462–3469. https://doi.org/10.1158/1078-0432.CCR-11-2696
Bertucci F, Ng CKY, Patsouris A, Droin N, Piscuoglio S, Carbuccia N, Soria JC, Dien AT, Adnani Y, Kamal M, Garnier S, Meurice G, Jimenez M, Dogan S, Verret B, Chaffanet M, Bachelot T, Campone M, Lefeuvre C, Bonnefoi H, Dalenc F, Jacquet A, De Filippo MR, Babbar N, Birnbaum D, Filleron T, Le Tourneau C, Andre F (2019) Genomic characterization of metastatic breast cancers. Nature 569(7757):560–564. https://doi.org/10.1038/s41586-019-1056-z
Bussolati G, Leonardo E (2008) Technical pitfalls potentially affecting diagnoses in immunohistochemistry. J Clin Pathol 61(11):1184–1192. https://doi.org/10.1136/jcp.2007.047720
Singh VM, Salunga RC, Huang VJ, Tran Y, Erlander M, Plumlee P, Peterson MR (2013) Analysis of the effect of various decalcification agents on the quantity and quality of nucleic acid (DNA and RNA) recovered from bone biopsies. Ann Diagn Pathol 17(4):322–326. https://doi.org/10.1016/j.anndiagpath.2013.02.001
Goto K, Ichinose Y, Ohe Y, Yamamoto N, Negoro S, Nishio K, Itoh Y, Jiang H, Duffield E, McCormack R, Saijo N, Mok T, Fukuoka M (2012) Epidermal growth factor receptor mutation status in circulating free DNA in serum: from IPASS, a phase III study of gefitinib or carboplatin/paclitaxel in non-small cell lung cancer. J Thorac Oncol 7(1):115–121. https://doi.org/10.1097/JTO.0b013e3182307f98
Kim TW, Peeters M, Thomas AL, Gibbs P, Hool K, Zhang J, Ang A, Bach BA, Price T (2018) Impact of emergent circulating tumor DNA RAS mutation in panitumumab-treated chemoresistant metastatic colorectal cancer. Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-17-3377
Board RE, Wardley AM, Dixon JM, Armstrong AC, Howell S, Renshaw L, Donald E, Greystoke A, Ranson M, Hughes A, Dive C (2010) Detection of PIK3CA mutations in circulating free DNA in patients with breast cancer. Breast Cancer Res Treat 120(2):461–467. https://doi.org/10.1007/s10549-010-0747-9
Baselga J, Im SA, Iwata H, Cortes J, De Laurentiis M, Jiang Z, Arteaga CL, Jonat W, Clemons M, Ito Y, Awada A, Chia S, Jagiello-Gruszfeld A, Pistilli B, Tseng LM, Hurvitz S, Masuda N, Takahashi M, Vuylsteke P, Hachemi S, Dharan B, Di Tomaso E, Urban P, Massacesi C, Campone M (2017) Buparlisib plus fulvestrant versus placebo plus fulvestrant in postmenopausal, hormone receptor-positive, HER2-negative, advanced breast cancer (BELLE-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol 18(7):904–916. https://doi.org/10.1016/S1470-2045(17)30376-5
Alix-Panabieres C, Pantel K (2016) Clinical applications of circulating tumor cells and circulating tumor DNA as liquid biopsy. Cancer Discov 6(5):479–491. https://doi.org/10.1158/2159-8290.CD-15-1483
Hayes DF, Ethier S, Lippman ME (2006) New guidelines for reporting of tumor marker studies in breast cancer research and treatment: REMARK. Breast Cancer Res Treat 100(2):237–238. https://doi.org/10.1007/s10549-006-9253-5
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM, Statistics Subcommittee of the NCIEWGoCD (2005) REporting recommendations for tumour MARKer prognostic studies (REMARK). Eur J Cancer 41(12):1690–1696. https://doi.org/10.1016/j.ejca.2005.03.032
Lamy PJ, Castan F, Lozano N, Montelion C, Audran P, Bibeau F, Roques S, Montels F, Laberenne AC (2015) Next-generation genotyping by digital PCR to detect and quantify the BRAF V600E Mutation in melanoma biopsies. J Mol Diagn 17(4):366–373. https://doi.org/10.1016/j.jmoldx.2015.02.004
Hu ZY, Xie N, Tian C, Yang X, Liu L, Li J, Xiao H, Wu H, Lu J, Gao J, Hu X, Cao M, Shui Z, Xiao M, Tang Y, He Q, Chang L, Xia X, Yi X, Liao Q, Ouyang Q (2018) Identifying circulating tumor DNA mutation profiles in metastatic breast cancer patients with multiline resistance. EBioMedicine 32:111–118. https://doi.org/10.1016/j.ebiom.2018.05.015
Rothe F, Laes JF, Lambrechts D, Smeets D, Vincent D, Maetens M, Fumagalli D, Michiels S, Drisis S, Moerman C, Detiffe JP, Larsimont D, Awada A, Piccart M, Sotiriou C, Ignatiadis M (2014) Plasma circulating tumor DNA as an alternative to metastatic biopsies for mutational analysis in breast cancer. Ann Oncol 25(10):1959–1965. https://doi.org/10.1093/annonc/mdu288
Liu YR, Jiang YZ, Zuo WJ, Yu KD, Shao ZM (2014) PIK3CA mutations define favorable prognostic biomarkers in operable breast cancer: a systematic review and meta-analysis. OncoTargets Ther 7:543–552. https://doi.org/10.2147/OTT.S60115
Hortobagyi GN, Stemmer SM, Burris HA, Yap YS, Sonke GS, Paluch-Shimon S, Campone M, Petrakova K, Blackwell KL, Winer EP, Janni W, Verma S, Conte P, Arteaga CL, Cameron DA, Mondal S, Su F, Miller M, Elmeliegy M, Germa C, O’Shaughnessy J (2018) Updated results from MONALEESA-2, a phase III trial of first-line ribociclib plus letrozole versus placebo plus letrozole in hormone receptor-positive, HER2-negative advanced breast cancer. Ann Oncol 29(7):1541–1547. https://doi.org/10.1093/annonc/mdy155
Andre F, Ciruelos E, Rubovszky G, Campone M, Loibl S, Rugo HS, Iwata H, Conte P, Mayer IA, Kaufman B, Yamashita T, Lu YS, Inoue K, Takahashi M, Papai Z, Longin AS, Mills D, Wilke C, Hirawat S, Juric D, Group S-S, the S-SG (2019) Alpelisib for PIK3CA-mutated, hormone receptor-positive advanced breast cancer. N Engl J Med 380(20):1929–1940. https://doi.org/10.1056/NEJMoa1813904
O’Leary B, Hrebien S, Morden JP, Beaney M, Fribbens C, Huang X, Liu Y, Bartlett CH, Koehler M, Cristofanilli M, Garcia-Murillas I, Bliss JM, Turner NC (2018) Early circulating tumor DNA dynamics and clonal selection with palbociclib and fulvestrant for breast cancer. Nat Commun 9(1):896. https://doi.org/10.1038/s41467-018-03215-x
Vora SR, Juric D, Kim N, Mino-Kenudson M, Huynh T, Costa C, Lockerman EL, Pollack SF, Liu M, Li X, Lehar J, Wiesmann M, Wartmann M, Chen Y, Cao ZA, Pinzon-Ortiz M, Kim S, Schlegel R, Huang A, Engelman JA (2014) CDK 4/6 inhibitors sensitize PIK3CA mutant breast cancer to PI3 K inhibitors. Cancer Cell 26(1):136–149. https://doi.org/10.1016/j.ccr.2014.05.020
Herrera-Abreu MT, Palafox M, Asghar U, Rivas MA, Cutts RJ, Garcia-Murillas I, Pearson A, Guzman M, Rodriguez O, Grueso J, Bellet M, Cortes J, Elliott R, Pancholi S, Baselga J, Dowsett M, Martin LA, Turner NC, Serra V (2016) Early adaptation and acquired resistance to CDK4/6 inhibition in estrogen receptor-positive breast cancer. Cancer Res 76(8):2301–2313. https://doi.org/10.1158/0008-5472.CAN-15-0728
Ma F, Zhu W, Guan Y, Yang L, Xia X, Chen S, Li Q, Guan X, Yi Z, Qian H, Yi X, Xu B (2016) ctDNA dynamics: a novel indicator to track resistance in metastatic breast cancer treated with anti-HER2 therapy. Oncotarget 7(40):66020–66031. https://doi.org/10.18632/oncotarget.11791
Peduzzi P, Concato J, Feinstein AR, Holford TR (1995) Importance of events per independent variable in proportional hazards regression analysis. II. Accuracy and precision of regression estimates. J Clin Epidemiol 48(12):1503–1510
Dirican E, Akkiprik M, Ozer A (2016) Mutation distributions and clinical correlations of PIK3CA gene mutations in breast cancer. Tumour Biol 37(6):7033–7045. https://doi.org/10.1007/s13277-016-4924-2
Acknowledgements
The authors want to thank Dr. Hélène de Forges for her substantive writing and editing assistance.
Funding
This ancillary study was supported by the “Fond pour la Recherche Val d’Aurelle” grant. This research was supported by the SIRIC Montpellier Cancer Grant INCa_Inserm_DGOS_12553.
Author information
Authors and Affiliations
Contributions
Study concept/design/funding: WJ, P-JL, EL-C, FD, and AD. Reagents/materials/analysis tools contribution: NG, P-JL, NL, and EL-C. Patients’ inclusion and follow-up: WJ, FD, AD, J-LL, SP, LG, GR, and HR. Acquisition of data: NG, WJ, P-JL, and EL-C. Statistical analysis: LC and TF. Analysis and interpretation of data: WJ, P-JL, and EL-C. Drafting of the manuscript: NG, WJ, P-JL, EL-C, FD, LC, and TF. All the authors participated to the critical revision and validation of the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This study was approved by the CPP (Ethics Committee) Sud-Ouest et Outre-Mer III and registered under the reference No. 2012/25. This study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jacot, W., Dalenc, F., Lopez-Crapez, E. et al. PIK3CA mutations early persistence in cell-free tumor DNA as a negative prognostic factor in metastatic breast cancer patients treated with hormonal therapy. Breast Cancer Res Treat 177, 659–667 (2019). https://doi.org/10.1007/s10549-019-05349-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05349-y