Abstract
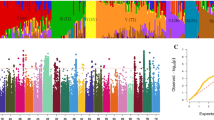
Three wheat and two barley populations were studied in order to find loci responsible for dormancy and pre-harvest sprouting. A classical quantitative trait loci analysis was combined with an association mapping approach. Many quantitative trait loci and marker trait associations could be detected on all seven chromosome groups of wheat and on the chromosomes 2H, 3H, 5H, 6H, and 7H of barley. Especially, the known regions on chromosomes 3A and 4A for wheat and 5H for barley were confirmed. Putative functions could be found via a candidate homologues search and via expressed sequence tag annotation. On chromosome 3A, the viviparous1 gene is located which is associated to preharvest sprouting and dormancy. On chromosome 4A, a protein is detected which belongs to the aquaporin family. In barley, an association with the aleurain gene on chromosome 5H was found. The expression of aleurain is regulated by abscisic acid and gibberelic acid. An influence of both hormones on dormancy and pre-harvest sprouting is known. It can be concluded that dormancy and pre-harvest sprouting are very complex traits regulated by multigenes and/or quantitative trait loci.
Similar content being viewed by others
Abbreviations
- ABA:
-
abscisic acid
- AM:
-
association mapping
- CS:
-
Chinese Spring
- D10:
-
dormancy at 10 °C
- D20:
-
dormancy at 20 °C
- DArT:
-
diversity array technology
- DI:
-
dormancy index
- EST:
-
expressed sequence tag
- GA:
-
gibberellins
- GLM:
-
general linear model
- IPK:
-
Leibniz Institute of Plant Genetics and Crop Plant Research
- ITMI:
-
International Triticeae Mapping Initiative
- LOD:
-
logarithm of odds
- MLM:
-
mixed linear model
- SOD:
-
superoxide dismutase
- MTA:
-
marker trait association
- NCBI:
-
National Centre for Biotechnology Information
- OWB:
-
Oregon Wolfe barley
- PHS:
-
pre-harvest sprouting
- QTL:
-
quantitative trait loci
References
Agrama, H.A., Eizenga, G.C., Yan, W.: Association mapping of yield and its components in rice cultivars. — Mol. Breed. 19: 341–356, 2007.
Aranzana, M.J., Kim, S., Zhao, K., Bakker, E., Horton, M., Jakob, K., Lister, C., Molitor, J., Shindo, C., Tang, C., Toomajian, C., Traw, B., Zheng, H., Bergelson, J., Dean, C., Marjoram, P., Nordborg, M.: Genome-wide association mapping in Arabidopsis identifies previously known flowering time and pathogen resistance genes. — PLoS Genet 1: e60, 2005.
Baek, K.-H., Skinner, D.Z., Ling, P., Chen, X.: Molecular structure and organization of the wheat genomic manganese superoxide dismutase gene. — Genome 49: 209–218, 2006.
Benech-Arnold, R.L., Giallorenzi, M.C., Frank, J., Rodriguez, V.: Termination of hull-imposed dormancy in developing barley grains is correlated with changes in embryonic ABA levels and sensitivity. — Seed Sci. Res. 9: 39–47, 1999.
Bethke, P.C., Fath, A., Spiegel, Y.N., Hwang, Y., Jones, R.L.: Abscisic acid, gibberellin and cell viability in cereal aleurone. — Euphytica 126: 3–11, 2002.
Börner, A., Schumann, E., Fürste, A., Cöster, H., Leithold, B., Röder, M.S., Weber, W.E.: Mapping of quantitative trait loci determining agronomic important characters in hexaploid wheat (Triticum aestivum L.). — Theor. appl. Genet. 105: 921–936, 2002.
Breseghello, F., Sorrells, M.E.: Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. — Genetics 172: 1165–1177, 2006.
Chang, C., Zhang, H.-P., Zhao, Q.-X., Feng, J.-M., Si, H.-Q., Lu, J., Ma, C.-X.: Rich allelic variations of Viviparous-1A and their associations with seed dormancy/pre-harvest sprouting of common wheat. — Euphytica 179: 343–353, 2011.
Costa, J.M., Corey, A., Hayes, P.M., Jobet, C., Kleinhofs, A., Kopisch-Obusch, A., Kramer, S.F., Kudrna, D., Li, M., Riera-Lizarazu, O., Sato, K., Szucs, P., Toojinda, T., Vales, M.I., Wolfe, R.I.: Molecular mapping of the Oregon Wolfe barleys: a phenotypically polymorphic doubled-haploid population. — Theor. appl. Genet. 103: 415–424, 2001.
Crossa, J., Burgueno, J., Dreisigacker, S., Vargas, M., Herrera-Foessel S.A., Lillemo, M., Singh, R.P., Trethowan, R., Warburton, M., Franco, J., Reynolds, M., Crouch, J.H., Ortiz, R.: Association analysis of historical bread wheat germplasm using additive genetic covariance of relatives and population structure. — Genetics 177: 1889–1913, 2007.
Flintham, J.E.: Different genetic components control coatimposed and embryo-imposed dormancy in wheat. — Seed Sci. Res. 10: 43–50, 2000.
Flintham, J., Adlam, R., Bassoi, M., Holdsworth, M., Gale, M.: Mapping genes for resistance to sprouting damage in wheat. — Euphytica 126: 39–45, 2002.
Foley, M.E., Fennimore, S.A.: Genetic basis for seed dormancy. — Seed Sci. Res. 8: 173–182, 1998.
Forrest, K.L., Bhave, M.: Physical mapping of wheat aquaporin genes. — Theor. appl. Genet. 120: 863–873, 2010.
Francki, M.G., Walker, E., Crawford, A.C., Broughton, S., Ohm, H.W., Barclay, I., Wilson, R.E., McLean, R.: Comparison of genetic and cytogenetic maps of hexaploid wheat (Triticum aestivum L.) using SSR and DArT markers. — Mol. Genet. Genomics 281: 181–191, 2009.
Ganal, M.W., Röder, M.S.: Microsatellite and SNP markers in wheat breeding. — In: Varshney, R.K., Tuberosa, R. (ed.): Genomics Assisted Crop Improvement. Vol. 2: Genomics Applications in Crops. Pp. 1–24. Springer, New York 2007.
Gao, W., Clancy, J.A., Han, F., Prada, D., Kleinhofs, A., Ullrich, S.E.: Molecular dissection of a dormancy QTL region near the chromosome 7 (5H) L telomere in barley. — Theor. appl. Genet. 107: 552–559, 2003.
Gerjets, T., Scholefield, D., Foulkes, M.J., Lenton, J.R., Holdsworth, M.J.: An analysis of dormancy, ABA responsiveness, after-ripening and pre-harvest sprouting in hexaploid wheat (Triticum aestivum L.) caryopses. — J. exp. Bot. 61: 597–607, 2010.
Groos, C., Gay, G., Perretant, M.-R., Gervais, L., Bernard, M., Dedryver, F., Charmet, G.: Study of the relationship between pre-harvest sprouting and grain color by quantitative trait loci analysis in a whitexred grain breadwheat cross. — Theor. appl. Genet. 104: 39–47, 2002.
Han, F., Ullrich, S.E., Clancy, J.A., Romagosa, I.: Inheritance and fine mapping of a major barley seed dormancy QTL. — Plant Sci. 143: 113–118, 1999.
Higa, A., Hidaka, T., Minai, Y., Matsuoka, Y., Haga, M.: Active oxygen radicals induce peroxidase activity in rice blade tissues. — Biosci. Biotechnol. Biochem. 65: 1852–1855, 2001.
Hildebrandt, T.M., Grieshaber, M.K.: Three enzymatic activities catalyze the oxidation of sulfide to thiosulfate in mammalian and invertebrate mitochondria. — FEBS J. 275: 3352–3361, 2008.
Huang, J., Zhao, X., Huihui, Y., Ouyang, Y., Wang, L., Zhang, Q.: The ankyrin repeat gene family in rice: genome-wide identification, classification and expression profiling. — Plant mol. Biol. 71: 207–226, 2009.
Jansen, C., Korell, M., Eckey, C., Biednkopf, D., Kogel, K.-H.: Identification and transcriptional analysis of powdery mildew-induced barley genes. — Plant Sci. 168: 373–380, 2005.
Kato, K., Nakamura, W., Tabiki, T., Miura, H., Sawada, S.: Detection of loci controlling seed dormancy on group 4 chromosomes of wheat and comparative mapping with rice and barley genomes. — Theor. appl. Genet. 102: 980–985, 2001.
Kervinen, T., Peltonen, S., Teeri, T.H., Karjalainen, R.: Differential expression of phenylalanine ammonia-lyase genes in barley induced by fungal infection or elicitors. — New Phytol. 139: 293–300, 1998.
Kikuchi, S., Taketa, S., Ichii, M., Kawasaki, S.: Efficient fine mapping of the naked caryopsis gene (nud) by HEGS (High Efficiency Genome Scanning)/AFLP in barley. — Theor. appl. Genet. 108: 73–78, 2003.
Kleinhofs, A., Kilian, A., Saghai Maroof, M., Biyashev, R., Hayes, P., Chen, F., Lapitan, N., Fenwick, A., Blake, T., Kanazin, V., Ananiev, E., Dahleen, L., Kudrna, D., Bollinger, J., Knapp, S., Liu, B., Sorrells, M., Heun, M., Franckowiak, J., Hovman, D., Skadsen, R., Stevenson, B.: A molecular, isozyme and morphological map of the barley (Hordeum vulgare) genome. — Theor. appl. Genet. 86: 705–713, 1993.
Kraakman, A.T.W., Niks, R.E., Van den Berg, P.M., Stam, P., Van Eeuwijk, F.A.: Linkage disequilibrium mapping of yield and yield stability in modern spring barley cultivars. — Genetics 168: 435–446, 2004.
Kraakman, A.T.W., Martinez, F., Mussiraliev, B., van Eeuwijk, F.A., Niks, R.E.: Linkage disequilibrium mapping of morphological, resistance, and other agronomically relevant traits in modern spring barley cultivars. — Mol. Breed. 17: 41–58, 2006.
Kulwal, P., Ishikawa, G., Benscher, D., Feng, Z., Yu, L.-X., Jadhav, A., Mehetre, S., Sorrells, M.E.: Association mapping for pre-harvest sprouting resistance in white winter wheat. — Theor. appl. Genet. 125: 793–805, 2012.
Kulwal, P.L., Kumar, N., Gaur, A., Khurana, P., Khurana, J.P., Tyagi, A.K., Balyan, H.S., Gupta, P.K.: Mapping of a major QTL for pre-harvest sprouting tolerance on chromosome 3A in bread wheat. — Theor. appl. Genet. 111: 1052–1059, 2005.
Kulwal, P.L., Mir, R.R., Kumar, S., Gupta, P.K.: QTL analysis and molecular breeding for seed dormancy and pre-harvest sprouting tolerance in bread wheat. — J. Plant Biol. 37: 59–74, 2010.
Kuroda, M., Kiyosaki, T., Matsumoto, I., Misaka, T., Arai, S., Abe, K.: Molecular cloning, characterization, and expression of wheat cystatins. — Biosci. Biotechnol. Biochem. 65: 22–28, 2001.
Lane, B.G.: Oxalate oxidases and differentiating surface structure in wheat: germins. — Biochem. J. 349: 309–321, 2000.
Li, C., Ni, P., Francki, M., Hunter, A., Zhang, Y., Schibeci, D., Li, H., Tarr, A., Wang, J., Cakir, M., Yu, J., Bellgard, M., Lance, R., Appels, R.: Genes controlling seed dormancy and pre-harvest sprouting in a rice-wheat-barley comparison. — Funct. Integr. Genomics 4: 84–93, 2004.
Liu, S., Bai, G., Cai, S., Chen, C.: Dissection of genetic components of preharvest sprouting resistance in white wheat. — Mol. Breed. 27: 511–523, 2011.
Lohwasser, U., Röder, M.S., Börner, A.: QTL mapping of the domestication traits pre-harvest sprouting and dormancy in wheat (Triticum aestivum L.). — Euphytica 143: 247–249, 2005.
Mares, D., Rathjen, J., Mrva, K., Cheong, J.: Genetic and environmental control of dormancy in white-grained wheat (Triticum aestivum L.). — Euphytica 168: 311–318, 2009.
McFadden, E.S., Sears, E.R.: The genome approach in radical wheat breeding. — J. amer. Soc. Agron. 39: 1011–1026, 1947.
Mori, M., Uchino, N., Chono, M., Kato, K., Miura, H.: Mapping QTLs for grain dormancy on wheat chromosome 3A and the group 4 chromosomes, and their combined effect. — Theor. appl. Genet. 110: 1315–1323, 2005.
Munkvold, J.D., Tanaka, J., Benscher, D. Sorrells, M.E.: Mapping quantitative trait loci for preharvest sprouting resistance in white wheat. — Theor. appl. Genet. 119: 1223–1235, 2009.
Nagel, M., Vogel, H., Landjeva, S., Buck-Sorlin, G., Lohwasser, U., Scholz, U., Börner, A.: Seed conservation in ex situ genebanks — genetic studies on longevity in barley. — Euphytica 170: 5–14, 2009.
Nelson, J.C.: QGENE: software for mapping — based genomic analysis and breeding. — Mol. Breed. 3: 239–245, 1997.
Neumann, K., Kobiljski, B., Dencic, S., Varshney, R.K., Börner, A.: Genome-wide association mapping: a case study in bread wheat (Triticum aestivum L.). — Mol. Breed. 27: 37–58, 2011.
Noda, K., Matsuura, T., Maekawa, M., Taketa, S.: Chromosomes responsible for sensitivity of embryo to abscisic acid and dormancy in wheat. — Euphytica 123: 203–209, 2002.
Ogbonnaya, F.C., Imtiaz, M., Ye, G., Hearnden, P.R., Hernandez, E., Eastwood, R.F., van Ginkel, M., Shorter, S.C., Winchester, J.M.: Genetic and QTL analyses of seed dormancy and preharvest sprouting resistance in wheat germplasm CN10955. — Theor. appl. Genet. 116: 891–902, 2008.
Osa, M., Kato, K., Mori, M., Shindo, C., Torada, A., Miura, H.: Mapping QTLs for seed dormancy and the Vp1 homologue on chromosome 3A in wheat. — Theor. appl. Genet. 106: 1491–1496, 2003.
Pestsova, E.G., Börner, A., Röder, M.S.: Development of a set of Triticum aestivum — Aegilops tauschii introgression lines. — Hereditas 135: 139–143, 2001.
Pestsova, E., Börner, A., Röder, M.S.: Development and QTL assessment of Triticum aestivum-Aegilops tauschii introgression lines. — Theor. appl. Genet. 112: 634–647, 2006.
Prada, D., Ullrich, S.E., Molina-Cano, J.L., Cistue, L., Clancy, J.A., Romagosa, I.: Genetic control of dormancy in a Triumph/Morex cross in barley. — Theor. appl. Genet. 109: 62–70, 2004.
Pulido, P., Dominguez, F., Cejudo, F.J.: A hydrogen peroxide detoxification system in the nucleus of wheat seed cells. — Plant Signal Behav. 4: 23–25, 2009.
Rehman Arif, M.A.: Seed longevity and dormancy in wheat (Triticum aestivum L.) — phenotypic variation and genetic mapping. — PhD Thesis, Martin-Luther-Universität Halle and Wittenberg Institut für Agrar- und Ernährungswissenschaften der Naturwissenschaftlichen Fakultät III, Halle 2012.
Rehman Arif, M.A., Neumann, K., Nagel, M., Kobiljski, B., Lohwasser, U., Börner, A.: An association mapping analysis of dormancy and pre-harvest sprouting in wheat. — Euphytica 188: 409–417, 2012.
Romero, A., Alamillo, J.M., Garcia-Olmedo, F.: Processing of thionin precursors in barley leaves by a vacuolar proteinase. — Eur. J. Biochem. 243: 202–208, 1997.
Sanchez-Monge, R., Delibes, A., Hernandez-Lucas, C., Carbonero, P., Garcia-Olmedo, F.: Homoeologous chromosomal location of the genes encoding thionins in wheat and rye. — Theor. appl. Genet. 54: 31–63, 1979.
Sato, K., Matsumoto, T., Ooe, N., Takeda, K.: Genetic analysis of seed dormancy QTL in barley. — Breed. Sci. 59: 645–650, 2009.
Stein, N., Prasad, M., Scholz, U., Thiel, T., Zhang, H.N., Wolf, M., Kota, R., Varshney, R.K., Perovic, D., Grosse, I., Graner, A.: A 1,000-loci transcript map of the barley genome: new anchoring points for integrative grass genomics. — Theor. appl. Genet. 114: 823–839, 2007.
Strand, E.: Studies on seed dormancy in barley. — Meldinger fra Norges Landbrukshogskole Hoegskole 44: 1–23, 1965.
Torada, A., Amano, Y.: Effect of seed coat color on seed dormancy in different environments. — Euphytica 126: 99–105, 2002.
Ullrich, S.E., Lee, H., Claney, J.A., Del Blanco, I.A., Jitkov, V.A., Kleinhofs, A., Han, F., Prada, D., Romagosa, I., Molina-Cano, J.L.: Genetic relationships between preharvest sprouting and dormancy in barley. — Euphytica 168: 331–345, 2009.
Xia, L.Q., Ganal, M.W., Shewry, P.R., He, Z.H., Yang, Y., Röder, M.S.: Exploiting the diversity of Viviparous-1 gene associated with pre-harvest sprouting tolerance in European wheat varieties. — Euphytica 159: 411–417, 2008.
Author information
Authors and Affiliations
Corresponding author
Additional information
Acknowledgements: The authors thank Marion Röder and Nils Stein for providing the map data and Annette Marlow, Stefanie Thumm, and Renate Voss for excellent technical assistance.
Rights and permissions
About this article
Cite this article
Lohwasser, U., Rehman Arif, M.A. & Börner, A. Discovery of loci determining pre-harvest sprouting and dormancy in wheat and barley applying segregation and association mapping. Biol Plant 57, 663–674 (2013). https://doi.org/10.1007/s10535-013-0332-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10535-013-0332-2