Abstract
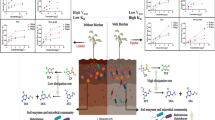
This study was performed to investigate the petroleum hydrocarbon (PH) degradative potential of indigenous microorganisms in ozonated soil to better develop combined pre-ozonation/bioremediation technology. Diesel-contaminated soils were ozonated for 0–900 min. PH and microbial concentrations in the soils decreased with increased ozonation time. The greatest reduction of total PH (TPH, 47.6%) and aromatics (11.3%) was observed in 900-min ozonated soil. The number of total viable heterotrophic bacteria decreased by three orders of magnitude in the soil. Ozonated soils were incubated for 9 weeks for bioremediation. The number of microorganisms in the soils increased during the incubation period, as monitored by culture- and nonculture-based methods. The soils showed additional PH-removal during incubation, supporting the presence of PH-degraders in the soils. The highest removal (25.4%) of TPH was observed during the incubation of 180-min ozonated soil during the incubation while a negligible removal was shown in 900-min ozonated soil. This negligible removal could be explained by the existence of relatively few or undetected PH-degraders in 900-min ozonated soil. After a 9-week incubation of the ozonated soils, 180-min ozonated soil showed the lowest TPH concentration, suggesting that appropriate ozonation and indigenous microorganisms survived ozonation could enhance remediation of PH-contaminated soil. Microbial community composition in 9-week incubated soils revealed a slight difference between 900-min ozonated and unozonated soils, as analyzed by whole cell hybridization. Taken together, this study provided insight into indigenous microbial potential to degrade PH in ozonated soils.
Similar content being viewed by others
References
Ahn Y, Sanseverino J & Sayler GS (1999) Analyses of polycyclic aromatic hydrocarbon-degrading bacteria isolated from contaminated soils. Biodegradation 10: 149–157
Amann RI, Krumholz L & Stahl DA (1990) Fluorescentoligonucleotide probing of whole cells for determinative, phylogenetic, and environmental studies in microbiology. J. Bacteriol. 172: 762–770
Amann RI, Ludwig W & Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 59: 143–169
Applegate BM, Matrabutjam U, Sanseverino J & Sayler GS (1995) Biodegradation genes as maker genes in microbial ecosystems. Mol. Microb. Ecol. Manual 6.1.8: 1–14
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA & Struhl K (1989) Current Protocols in Molecular Biology. John Wiley & Sons Ltd, New York
Casida LE Jr (1971) Microorganisms in unamended soil as observed by various forms of microscopy and staining. Appl. Microbiol. 21: 1040–1045
Cerniglia CE (1993) Biodegradation of polycyclic aromatic hydrocarbons. Curr. Opin. Biotechnol. 4: 331–338
Choi H, Lim HN, Kim J, Hwang TM & Kang JW (2002) Transport characteristics of gas phase ozone in unsaturated porous media for in situ chemical oxidation. J. Contam. Hydrol. 57: 81–98
Foght JM, Fedorak PM & Westlake DW (1990) Mineralization of [14C]hexadecane and [14C]phenanthrene in crude oil: specificity among bacterial isolates. Can. J. Microbiol. 36: 169–175
Gilbert E (1983) Investigations on the changes of biological degradability of single substances induced by ozonation. Ozone Sci. Eng. 5: 137–149
Gilbert E (1987) Biodegradability of ozonation products as a function of COD and DOC elimination by example of substituted aromatic substances. Water Res. 21: 1273–1278
Harayama S, Kok M & Neidle EL (1992) Functional and evolutionary relationships among diverse oxygenases. Ann. Rev. Microbiol. 46: 565–601
Hus I & Masten SJ (1997) The kinetics of the reaction of ozone with phenanthrene in unsaturated soils. Environ. Eng. Sci. 14: 207–217
Khadre MA, Yousef AE & Kim J-G (2001) Microbiological aspects of ozone applications in food: a review. J. Food Sci. 66: 1242–1253
Klappenbach JA, Saxman PR, Cole JT & Schmidt TM (2001) rrndb: the ribosomal RNA operon copy number database. Nucleic Acids Res. 29: 181–184
Kok M, Oldenhuis R, van der Linden MPG, Raatjes P, Kingma J, van Lelyveld PH & Witholt B (1989) The Pseudomonas oleovorans alkane dehydrogenase gene: sequence and expression. J. Biol. Chem. 264: 5435–5441
Kowalski WJ, Bahnfleth WP & Whittam TS (1998) Bactericidal effects of high airborne concentrations of Escherichia coli and Staphylococcus aureus. Ozone Sci. Eng. 20: 205–221
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E & Goodfellow M (Eds) Nucleic Acid Techniques in Bacterial Systematics (pp 115–147). John Wiley & Sons Ltd, New York
Legube B, Langlais B, Sohm B & Dore M (1981) Identification of ozonation products of aromatic hydrocarbon micropollutants: effect on chlorination and biological filtration. Ozone Sci. Eng. 3: 33–48
Lim HN, Choi H, Hwang TM & Kang JW (2002) Characterization of ozone decomposition in a soil slurry: kinetics and mechanism. Water Res. 36: 219–229
Lute JR, Skladany GJ & Nelson CH (1998) Evaluating the effectiveness of ozonation and combined ozonation/bioremediation technologies. In: Wickramanayake GB & Hinchee RE (Eds) Designing and Applying Treatment Technologies: Remediation of Chlorinated and Recalcitrant Compounds. Battelle Press, Columbus, Ohio
Madigan MT, Martinko JM & Parker J (2000) Biology of Microorganisms, 9th edn. Prentice Hall, Inc. New York
Manz W, Amann RI & Ludwig W (1992) Phylogenic oligonucleotide probes for major subclass of Proteobacteria: problems and solutions. System. Appl. Microbiol. 15: 593–600
Masten SJ & Davies SHR (1997) Efficacy of in-situ ozonation for the remediation of PAH contaminated soils. J. Contam. Hydrol. 28: 327–335
Nam K & Kukor JJ (2000) Combined ozonation and biodegradation for remediation of mixtures of polycyclic aromatic hydrocarbons in soil. Biodegradation 11: 1–9
Nelson CH & Brown RA (1994) Adapting ozonation for soil and groundwater cleanup. Environ. Eng., a Supplement to Chem. Eng. 20: EE18–EE24
Rabus R, Fukui M, Wilkes H & Widdel F (1996) Degradative capacities and 16S rRNA-targeted whole cell hybridization of sulfate-reducing bacteria in an anaerobic enrichment culture utilizing alkylbenzenes from crude oil. Appl. Environ. Microbiol. 62: 3605–3613
Ripp S, Nivens DE, Ahn Y, Werner C, Jarrell IV J, Easter JP, Cox CD, Burlage R & Sayler GS (2000) Controlled field release of a bioluminescent genetically engineered microorganism for bioremediation process monitoring and control. Environ. Sci. Technol. 34: 846–853
Roller C, Wagner M, Amann R, Ludwig W & Schleifer K-H (1994) In situ probing of gram-positive bacteria with high G +C content using 23S rRNA-targeted oligonucleotides. Microbiology 140: 2849–2858
Sanger F, Coulson AR, Hong GF, Hill DF & Peterson GB (1982) Nucleotide sequence of bacteriophage ? DNA. Mol. Biol. 162: 729–733
Sanseverino J, Werne C, Fleming JT, Applegate BM, King JHM & Sayler GS (1993) Molecular diagnostics of polycyclic aromatic hydrocarbon degradation in manufactured gas plant soils. Biodegradation 4: 303–321
Sayler GS, Cox CD, Burlage R, Ripp S, Nivens DE, Werner C, Ahn Y & Matrubutham U (1999) Field application of a genetically engineered microorganism for polycyclic aromatic hydrocarbon bioremediation process monitoring and control. In: Fass et al. (Eds) Novel approaches for Bioremediation of Organic Pollution (pp 241–254). Plenum Press, New York
Simon MJ, Osslund TD, Saunder R, Ensley BTD, Suggs S, Harcourt A, Suen W, Cruden DL, Gibson DT & Zylstra GJ (1993) Sequence of genes encoding naphthalene dioxygenase in Pseudomonas putida strains G7 and NCIB 9816–4.Gene 127: 31–37
Soil Science Society of America (1986) Part 1, physical and mineralogical methods. In: Klute A (Ed) Methods of Soil Analysis, 2nd edn. Soil Science Society of America, Inc. Publisher. Madison, Wisconsin
Song HG & Bartha R (1990) Effects of jet fuel spills on the microbial community of soil. Appl. Environ. Microbiol. 56: 646–651
Sotsky JB, Greer CW & Atlas RM (1994) Frequency of genes in aromatic and aliphatic hydrocarbon biodegradation pathways within bacterial populations from Alaskan sediments. Can. J. Microbiol. 40: 981–985
Stahl DA, Flescher B, Mansfield HR & Montgomery L (1988) Use of phylogenetically based hybridization probes for studies of ruminal microbial ecology. Appl. Environ. Microbiol. 54: 179–184
Stapleton RD & Sayler GS (1998) Assessment of the microbiological potential for the natural attenuation of petroleum hydrocarbons in a shallow aquifer system. Microb. Ecol. 36: 349–361
Stehr J, Muller T, Svensson K, Kamnerdpetch C & Scheper T (2001) Basic examinations on chemical pre-oxidation by ozone for enhancing bioremediation of phenanthrene contaminated soils. Appl. Microbiol. Biotechnol. 57: 803–809
Sung M & Huang CP (2002) In situ removal of 2-chlorophenol from unsaturated soils by ozonation. Environ. Sci. Technol. 36: 2911–2918
US EPA (1996a) Test methods for evaluating solid waste, physical/chemical methods (SW-846), Method 3545, pressurized fluid extraction (PFE). Office of solid waste and emergency response, Washington DC
US EPA (1996b) Test methods for evaluating solid waste, physical/chemical methods (SW-846), Method 3630C, silica gel cleanup, Office of solid waste and emergence response, Washington DC
van Veen JA, van Overbeek LS & van Elsas JD (1997) Fate and activity of microorganisms introduced into soil. Microbiol. Mol. Biol. Rev. 61: 121–135
Vomberg A & Klinner U (2000) Distribution of alkB genes within n-alkane-degrading bacteria. J. Appl. Microbiol. 89: 339–348
Zylstra GJ & Gibson DT (1989) Toluene degradation by Pseudomonas putida F1. J. Biol. Chem. 264: 14940–14946
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ahn, Y., Jung, H., Tatavarty, R. et al. Monitoring of petroleum hydrocarbon degradative potential of indigenous microorganisms in ozonated soil. Biodegradation 16, 45–56 (2005). https://doi.org/10.1007/s10531-004-0428-2
Issue Date:
DOI: https://doi.org/10.1007/s10531-004-0428-2