Abstract
Objective
To construct effective genetic expression parts controlling transcription and translation initiation for synthetic biology and heterologous expression in Corynebacterium glutamicum.
Result
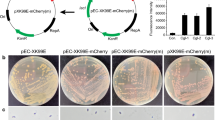
Twelve highly expressed genes were identified from the proteomic data of C. glutamicum. Their related sequences were used to construct bicistronic genetic expression parts. Each part contain promoter, 5′-UTR, N-terminal sequence of the source gene and a conserved SD sequence, associated with target gene, forming the bicistronic expression cassette. The enhanced green fluorescent protein (EGFP) expression levels controlled by these novel parts have 1.4 to 790-fold increase in C. glutamicum compared with corresponding promoter-5′-UTR part. One of the bicistronic parts is 1.35 times the EGFP expression of the constitutive-expression pXMJ19. These bicistronic parts had expression advantage compared with conventional promoter-5′-UTR parts.
Conclusion
Various genetic parts for efficient gene expression can be quickly obtained via this new method.






Similar content being viewed by others
References
Becker J, Wittmann C (2012) Bio-based production of chemicals, materials and fuels—Corynebacterium glutamicum as versatile cell factory. Curr Opin Biotechnol 23:631–640
Dai Z, Nielsen J (2015) Advancing metabolic engineering through systems biology of industrial microorganisms. Curr Opin Biotechnol 36:8–15
Engler C, Kandzia R, Marillonnet S (2008) A one pot, one step, precision cloning method with high throughput capability. PLoS One 3:e3647
Estrem ST, Gaal T, Ross W, Gourse RL (1998) Identification of an UP element consensus sequence for bacterial promoters. Proc Natl Acad Sci USA 95:9761–9766
Goodman DB, Church GM, Kosuri S (2013) Causes and effects of N-terminal codon bias in bacterial genes. Science 342:475–479
Grunberg-Manago M (1999) Messenger RNA stability and its role in control of gene expression in bacteria and phages. Ann Rev Genet 33:193–227
Gu Q, Wang W, Bower AGW, Collins CH, Koffas MAG (2013) Modular optimization of multi-gene pathways for fatty acids production in E. coli. Nat Commun 4:1409
Hu YD, Wan H, Li JG, Zhou JW (2015) Enhanced production of L-sorbose in an industrial Gluconobacter oxydans strain by identification of a strong promoter based on proteomics analysis. J Ind Microbiol Biotechnol 42:1039–1047
Huang JF et al (2011) The patterns and expression of KDR in normal tissues of human internal organs. J Mol Histol 42:597–603
Jakoby M, Ngouoto-Nkili CE, Burkovski A (1999) Construction and application of new Corynebacterium glutamicum vectors. Biotechnol Tech 13:437–441
Jang SA, Sung BH, Cho JH, Kim SC (2009) Direct expression of antimicrobial peptides in an intact form by a translationally coupled two-cistron expression system. Appl Environ Microbiol 75:3980–3986
Kosuri S et al (2013) Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc Natl Acad Sci USA 110:14024–14029
Kudla G, Murray AW, Tollervey D, Plotkin JB (2009) Coding-sequence determinants of gene expression in Escherichia coli. Science 324:255–258
Levin-Karp A et al (2013) Quantifying translational coupling in E. coli synthetic operons using RBS modulation and fluorescent reporters. ACS Synth Biol 2:327–336
Liu X et al (2015) Expression of recombinant protein using Corynebacterium glutamicum: progress, challenges and applications. Crit Rev Biotechnol. doi:10.3109/07388551.2015.1004519
Martin JF, Barreiro C, Gonzalez-Lavado E, Barriuso M (2003) Ribosomal RNA and ribosomal proteins in corynebacteria. J Biotechnol 104:41–53
Ming YM, Wei ZW, Lin CY, Sheng GY (2010) Development of a Bacillus subtilis expression system using the improved Pglv promoter. Microb Cell Fact 9:1
Mutalik VK et al (2013a) Precise and reliable gene expression via standard transcription and translation initiation elements. Nat Methods 10:354
Mutalik VK et al (2013b) Quantitative estimation of activity and quality for collections of functional genetic elements. Nat Methods 10:347
Nesvera J, Patek M (2011) Tools for genetic manipulations in Corynebacterium glutamicum and their applications. Appl Microbiol Biotechnol 90:1641–1654
Pfeifer-Sancar K, Mentz A, Rueckert C, Kalinowski J (2013) Comprehensive analysis of the Corynebacterium glutamicum transcriptome using an improved RNAseq technique. BMC Genom 14:1. doi:10.1186/1471-2164-14-888
Qin XL, Qian JC, Yao GF, Zhuang YP, Zhang SL, Chu J (2011) GAP promoter library for fine-tuning of gene expression in Pichia pastoris. Appl Environ Microbiol 77:3600–3608
Rytter JV, Helmark S, Chen J, Lezyk MJ, Solem C, Jensen PR (2014) Synthetic promoter libraries for Corynebacterium glutamicum. Appl Microbiol Biotechnol 98:2617–2623
Schaffer S et al (2001) A high-resolution reference map for cytoplasmic and membrane-associated proteins of Corynebacterium glutamicum. Electrophoresis 22:4404–4422
van der Rest ME, Lange C, Molenaar D (1999) A heat shock following electroporation induces highly efficient transformation of Corynebacterium glutamicum with xenogeneic plasmid DNA. Appl Microbiol Biotechnol 52:541–545
Wang J, Ai X, Mei H, Fu Y, Chen B, Yu Z, He J (2013) High-throughput identification of promoters and screening of highly active promoter-5′-UTR DNA region with different characteristics from Bacillus thuringiensis. PLoS One 8:e62960. doi:10.1371/journal.pone.0062960
Wendisch VF, Bott M, Eikmanns BJ (2006) Metabolic engineering of Escherichia coli and Corynebacterium glutamicum for biotechnological production of organic acids and amino acids. Curr Opin Microbiol 9:268–274
Woo HM, Park J-B (2014) Recent progress in development of synthetic biology platforms and metabolic engineering of Corynebacterium glutamicum. J Biotechnol 180:43–51
Yim SS, An SJ, Kang M, Lee J, Jeong KJ (2013) Isolation of fully synthetic promoters for high-level gene expression in Corynebacterium glutamicum. Biotechnol Bioeng 110:2959–2969
Acknowledgment
This work was financially supported by the National Basic Research Program of China (973 Program) (Grant No. 2013CB733602), the Fundamental Research Funds for the Central Universities (Grant No. JUSRP51401A) and the Natural Science Foundation of Jiangsu Province (Grant No. BK20150148).
Supporting information
Supplementary Table 1—Primers used.
Supplementary Table 2—Information about highly expressed genes in C. glutamicum.
Supplementary Fig. 1—Construction process of the probe vector pE-0.
Supplementary Fig. 2—Insertion of fragments guided by Golden Gate method to seamlessly connect genetic expression parts to reporter gene.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, Z., Liu, X., Zhang, W. et al. Construction of genetic parts from the Corynebacterium glutamicum genome with high expression activities. Biotechnol Lett 38, 2119–2126 (2016). https://doi.org/10.1007/s10529-016-2196-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-016-2196-y