Abstract
Objectives
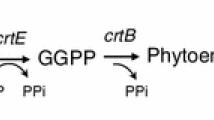
Putative genes crtE, crtB, and crtI from Deinococcus wulumiqiensis R12, a novel species, were identified by genome mining and were co-expressed using the optimized Shine-Dalgarno (SD) regions to improve lycopene yield.
Results
A lycopene biosynthesis pathway was constructed by co-expressing these three genes in Escherichia coli. After optimizing the upstream SD regions and the culture medium, the recombinant strain EDW11 produced 88 mg lycopene g−1 dry cell wt (780 mg lycopene l−1) after 40 h fermentation without IPTG induction, while the strain EDW without optimized SD regions only produced 49 mg lycopene g−1 dry cell wt (417 mg lycopene l−1).
Conclusion
Based on the optimization of the upstream SD regions and culture medium, the yield of the strain EDW11 reached a high level during microbial lycopene production until now.





Similar content being viewed by others
References
Ajikumar PK, Xiao WH, Tyo KEJ et al (2010) Isoprenoid pathway optimization for taxol precursor overproduction in Escherichia coli. Science 330:70–74
Bahieldin A, Gadalla NO, Al-Garni SM et al (2014) Efficient production of lycopene in Saccharomyces cerevisiae by expression of synthetic crt genes from a plasmid harboring the ADH2 promoter. Plasmid 72:18–28
Chen J, Zhu X, Tan Z et al (2014) Activating C4-dicarboxylate transporters DcuB and DcuC for improving succinate production. Appl Microbiol Biotechnol 98:2197–2205
Chenna R, Sugawara H, Koike T et al (2003) Multiple sequence alignment with the Clustal series of programs. Nucl Acid Res 31:3497–3500
Ji X, Liu L, Shen M et al (2015) Constructing a synthetic metabolic pathway in Escherichia Coli to produce the enantiomerically pure (R, R)-2,3-butanediol. Biotechnol Bioeng 112:1056–1059
Kim SH, Park YH, Schmidt-Dannert C, Lee PC (2010) Redesign, reconstruction, and directed extension of the Brevibacterium linens C40 carotenoid pathway in Escherichia coli. Appl Environ Microb 76:5199–5206
Kim Y, Lee J, Kim N et al (2011) Increase of lycopene production by supplementing auxiliary carbon sources in metabolically engineered Escherichia coli. Appl Microbiol Biot 90:489–497
Koch AL (1983) The protein burden of lac operon products. J Mol Evol 19:455–462
Kong K, Khoo H, Prasad KN et al (2010) Revealing the power of the natural red pigment lycopene. Molecules 15:959–987
Kosuri S, Goodman DB, Cambray G et al (2013) Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc Natl Acad Sci USA 110:14024–14029
Novick A, Weiner M (1957) Enzyme induction as an all-or-none phenomenon. Proc Natl Acad Sci USA 43:553
Pennacchio A, Pucci B, Secundo F et al (2008) Purification and characterization of a novel recombinant highly enantioselective short-chain NAD(H)-dependent alcohol dehydrogenase from Thermus thermophilus. Appl Environ Microbiol 74:3949–3958
Pfleger BF, Pitera DJ, Smolke CD et al (2006) Combinatorial engineering of intergenic regions in operons tunes expression of multiple genes. Nat Biotechnol 24:1027–1032
Sun T, Miao L, Li Q et al (2014) Production of lycopene by metabolically-engineered Escherichia coli. Biotechnol Lett 36:1515–1522
Thakker C, Zhu J, San K, Bennett G (2011) Heterologous pyc gene expression under various natural and engineered promoters in Escherichia coli for improved succinate production. J Biotechnol 155:236–243
Tian B, Xu Z, Sun Z et al (2007) Evaluation of the antioxidant effects of carotenoids from Deinococcus radiodurans through targeted mutagenesis, chemiluminescence, and DNA damage analyses. Biochim Biophys Acta 1770:902–911
Verwaal R, Jiang Y, Wang J et al (2010) Heterologous carotenoid production in Saccharomyces cerevisiae induces the pleiotropic drug resistance stress response. Yeast 27:983–998
Wang W, Mao J, Zhang Z et al (2010) Deinococcus wulumuqiensis sp. nov., and Deinococcus xibeiensis sp. nov., isolated from radiation-polluted soil. Int J Syst Evol Micr 60:2006–2010
Xu X, Jiang L, Zhang Z et al (2013) Genome sequence of a gamma- and UV-ray-resistant strain, Deinococcus wulumuqiensis R12. Genome Announc 1:e206–e213
Yoon S, Kim J, Lee S et al (2007) Engineering the lycopene synthetic pathway in E. coli by comparison of the carotenoid genes of Pantoea agglomerans and Pantoea ananatis. Appl Microbiol Biotechnol 74:131–139
Zhao J, Li Q, Sun T et al (2013) Engineering central metabolic modules of Escherichia coli for improving β-carotene production. Metab Eng 17:42–50
Zhou Y, Nambou K, Wei L et al (2013) Lycopene production in recombinant strains of Escherichia coli is improved by knockout of the central carbon metabolism gene coding for glucose-6-phosphate dehydrogenase. Biotechnol Lett 35:2137–2145
Acknowledgments
This work was supported by the National Science Foundation for Distinguished Young Scholars of China (21225626), the National High Technology Research and Development Program of China (2012AA021705, 2012AA022101), the National Natural Science Foundation of China for Young Scholars (21406111), and the Natural Science Foundation of Jiangsu Province, China (BK20131406).
Supporting information
Supplementary Table 1—Bacterial strains and plasmids used in this work.
Supplementary Table 2—Primers used in this work.
Supplementary Fig. 1—HPLC profiles of the isolated carotenoids from the strain EDWe, EDW, EDW11, EDW12, EDW21, EDW22, and the commercial lycopene.
Supplementary Fig. 2—The color of the cell pellets of the strains EDW11, EDW12, EDW21, and EDW22.
Supplementary Fig. 3—Comparison of the phytoene dehydrogenase expression and the growth of the recombinant strains E. coli EDW-I1 and EDW-I2.
Supplementary Fig. 4—Comparison of the phytoene synthase expression and the growth of the recombinant strains E. coli EDW-B1 and EDW-B2.
Author information
Authors and Affiliations
Corresponding author
Additional information
Weiyue Jin and Xian Xu have contributed to the work equally and should be regarded as co-first authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jin, W., Xu, X., Jiang, L. et al. Putative carotenoid genes expressed under the regulation of Shine–Dalgarno regions in Escherichia coli for efficient lycopene production. Biotechnol Lett 37, 2303–2310 (2015). https://doi.org/10.1007/s10529-015-1922-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1922-1