Abstract
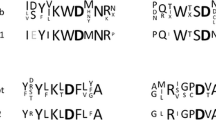
A putative recombinant β-galactosidase from Deinococcus geothermalis was purified as a single 79 kDa band of 42 U activity/mg using His-Trap affinity chromatography. The molecular mass of the native enzyme was a 158 kDa dimer. The catalytic residues E151 and E325 of β-galactosidase from D. geothermalis were conserved in all aligned GH family 42 β-galactosidases, indicating that this enzyme is also a GH family 42 β-galactosidase. Maximal activity of the enzyme was at pH 6.5 and 60°C. It has a unique hydrolytic activity for p-nitrophenyl(pNP)-β-d-galactopyranoside (k cat/K m = 69 s−1 mM−1), pNP-β-d-fucopyranoside (13), oNP-β-d-galactopyranoside (9.5), oNP-β-d-fucopyranoside (2.6), lactose (0.97), and pNP-α-l-arabinopyranoside (0.78), whereas no activity, or less than 2% of the pNP-β-d-galactopyranoside activity, for other pNP- and oNP- glycosides.




Similar content being viewed by others
References
Coker JA, Sheridan PP, Loveland-Curtze J, Gutshall KR, Auman AJ, Brenchley JE (2003) Biochemical characterization of a beta-galactosidase with a low temperature optimum obtained from an Antarctic Arthrobacter isolate. J Bacteriol 185:5473–5482
Copeland A, Lucas S, Lapidus A et al (2006) Complete sequence of chromosome 1 of Deinococcus geothermalis DSM 11300. EMBL/GenBank/DDBJ databases
Davies G, Henrissat B (1995) Structures and mechanisms of glycosyl hydrolases. Structure 3:853–859
Durand P, Lehn P, Callebaut I, Fabrega S, Henrissat B, Mornon JP (1997) Active-site motifs of lysosomal acid hydrolases: invariant features of clan GH-A glycosyl hydrolases deduced from hydrophobic cluster analysis. Glycobiology 7:277–284
Ferreira AC, Nobre MF, Rainey FA, Silva MT, Wait R, Burghardt J, Chung AP, da Costa MS (1997) Deinococcus geothermalis sp. nov. and Deinococcus murrayi sp. nov., two extremely radiation-resistant and slightly thermophilic species from hot springs. Int J Syst Bacteriol 47:939–947
Henrissat B (1991) A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 280(Pt 2):309–316
Henrissat B, Callebaut I, Fabrega S, Lehn P, Mornon JP, Davies G (1995) Conserved catalytic machinery and the prediction of a common fold for several families of glycosyl hydrolases. Proc Natl Acad Sci USA 92:7090–7094
Hidaka M, Fushinobu S, Ohtsu N, Motoshima H, Matsuzawa H, Shoun H, Wakagi T (2002) Trimeric crystal structure of the glycoside hydrolase family 42 beta-galactosidase from Thermus thermophilus A4 and the structure of its complex with galactose. J Mol Biol 322:79–91
Hinz SW, van den Brock LA, Beldman G, Vincken JP, Voragen AG (2004) beta-Galactosidase from Bifidobacterium adolescentis DSM20083 prefers beta(1,4)-galactosides over lactose. Appl Microbiol Biotechnol 66:276–284
Holmes ML, Scopes RK, Moritz RL, Simpson RJ, Englert C, Pfeifer F, Dyall-Smith ML (1997) Purification and analysis of an extremely halophilic beta-galactosidase from Haloferax alicantei. Biochim Biophys Acta 1337:276–286
Hu JM, Li H, Cao LX, Wu PC, Zhang CT, Sang SL, Zhang XY, Chen MJ, Lu JQ, Liu YH (2007) Molecular cloning and characterization of the gene encoding cold-active beta-galactosidase from a psychrotrophic and halotolerant Planococcus sp. L4. J Agric Food Chem 55:2217–2224
Kang SK, Cho KK, Ahn JK, Bok JD, Kang SH, Woo JH, Lee HG, You SK, Choi YJ (2005) Three forms of thermostable lactose-hydrolase from Thermus sp. IB-21: cloning, expression, and enzyme characterization. J Biotechnol 116:337–346
Li L, Zhang M, Jiang Z, Tang L, Cong Q (2009) Characterisation of a thermostable family 42 beta-galactosidase from Thermotoga maritima. Food Chem 112:844–850
Pan Q, Zhu J, Liu L, Cong Y, Hu F, Li J, Yu X (2010) Functional identification of a putative beta-galactosidase gene in the special lac gene cluster of Lactobacillus acidophilus. Curr Microbiol 60:172–178
Park AR, Oh DK (2010) Effects of galactose and glucose on the hydrolysis reaction of a thermostable beta-galactosidase from Caldicellulosiruptor saccharolyticus. Appl Microbiol Biotechnol 85:1427–1435
Puri M, Gupta S, Pahuja P, Kaur A, Kanwar JR, Kennedy JF (2009) Cell disruption optimization and covalent immobilization of beta-d-galactosidase from Kluyveromyces marxianus YW-1 for lactose hydrolysis in milk. Appl Biochem Biotechnol 160:98–108
Shaikh SA, Khire JM, Khan MI (1999) Characterization of a thermostable extracellular beta-galactosidase from a thermophilic fungus Rhizomucor sp. Biochim Biophys Acta 1472:314–322
Yuan T, Yang P, Wang Y, Meng K, Luo H, Zhang W, Wu N, Fan Y, Yao B (2008) Heterologous expression of a gene encoding a thermostable beta-galactosidase from Alicyclobacillus acidocaldarius. Biotechnol Lett 30:343–348
Acknowledgment
This study was carried out with the support of the 21C Frontier Project for Microbial Genomics, Ministry of Education, Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lee, JH., Kim, YS., Yeom, SJ. et al. Characterization of a glycoside hydrolase family 42 β-galactosidase from Deinococcus geothermalis . Biotechnol Lett 33, 577–583 (2011). https://doi.org/10.1007/s10529-010-0459-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-010-0459-6