Abstract
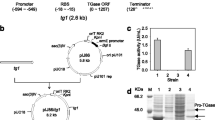
Inulinase gene (Kcinu) derived from Kluyveromyces cicerisporus was expressed extracellularly in Kluyveromyces lactis using an episomal vector directed by Kcinu promoter. The influence of hap1 gene disruption on the expression of inulinase was studied. Inulinase activity in the supernatant of the recombinant Klhap1Δ strain was 391 U ml−1 after cultured 120 h, which was 2.2-fold that of the wild type host. The relative inulinase mRNA level of the Klhap1Δ strain was 11.3-fold that of the wild type strain, and the expression plasmid was more stable in the mutant host. Based on these results, the disruption of hap1 facilitated the high and stable expression of inulinase controlled by Kcinu promoter in K. lactis.





Similar content being viewed by others
References
Adams A, Gottschling DE, Kaiser CA, Stearns T (1997) Methods in yeast genetics, 1997 edition. Cold Spring Harbor Laboratory Press, New York, USA
Bao WG, Guiard B, Fang ZA, Donnini C, Gervais M, Passos FML, Ferrero I, Fukuhara H, Bolotin-Fukuhara M (2008) Oxygen-dependent transcriptional regulator Hap1p limits glucose uptake by repressing the expression of the major glucose transporter gene RAG1 in Kluyveromyces lactis. Eukaryot Cell 7:1895–1905
Cai XP, Zhang J, Yuan HY, Fang ZA, Li YY (2005) Secretory expression of heterologous protein in Kluyveromyces cicerisporus. Appl Microbiol Biotechnol 67:364–369
Jayaram M, Li YY, Broach JR (1983) The yeast plasmid 2 μ circle encodes components required for its high copy propagation. Cell 34:95–104
Lamas-Maceiras M, Nunez L, Rodriguez-Belmonte E, Gonzalez-Siso MI, Cerdan ME (2007) Functional characterization of KlHAP1: a model to foresee different mechanisms of transcriptional regulation by Hap1p in yeasts. Gene 405:96–107
Muller S, Sandal T, Kamp-Hansen P, Dalboge H (1998) Comparison of expression systems in the yeasts Saccharomyces cerevisiae, Hansenula polymorpha, Klyveromyces lactis, Schizosaccharomyces pombe and Yarrowia lipolytica. Cloning of two novel promoters from Yarrowia lipolytica. Yeast 14:1267–1283
Ohta K, Akimoto H, Moriyama S (2004) Fungal inulinases: enzymology, molecular biology and biotechnology. J Appl Glycosci 51:247–254
Pandey A, Soccol CR, Selvakumar P, Soccol VT, Krieger N, Fontana JD (1999) Recent developments in microbial inulinases—its production, properties, and industrial applications. Appl Biochem Biotechnol 81:35–52
Swinkels BW, Vanooyen AJJ, Bonekamp FJ (1993) The yeast Kluyveromyces-lactis as an efficient host for heterologous gene-expression. Antonie Van Leeuwenhoek 64:187–201
Ter Linde JJM, Steensma HY (2002) A microarray-assisted screen for potential Hap1 and Rox1 target genes in Saccharomyces cerevisiae. Yeast 19:825–840
van Ooyen AJJ, Dekker P, Huang M, Olsthoorn MMA, Jacobs DI, Colussi PA, Taron CH (2006) Heterologous protein production in the yeast Kluyveromyces lactis. FEMS Yeast Res 6:381–392
Yuan XL, Roubos JA, van den Hondel C, Ram AFJ (2008) Identification of InuR, a new Zn(II)2Cys6 transcriptional activator involved in the regulation of inulinolytic genes in Aspergillus niger. Mol Genet Genomics 279:11–26
Zhang L, Hach A (1999) Molecular mechanism of heme signalling in yeast: the transcriptional activator Hap1 serves as the key mediator. Cell Mol Life Sci 56:415–426
Acknowledgments
This work was supported by grants from Chinese 863 Program (No. 2007AA02Z107) and Shanghai Major Basic Research Program (No. 08JC1401100) .
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, J., Jiang, J., Fang, Z. et al. Enhanced expression of heterologous inulinase in Kluyveromyces lactis by disruption of hap1 gene. Biotechnol Lett 32, 507–512 (2010). https://doi.org/10.1007/s10529-009-0182-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-009-0182-3