Abstract
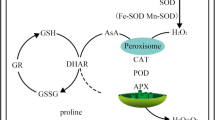
The genus Methylobacterium is composed of pink-pigmented methylotrophic bacterial species that are widespread in natural environments, such as soils, stream water and plants. When in association with plants, this genus colonizes the host plant epiphytically and/or endophytically. This association is known to promote plant growth, induce plant systemic resistance and inhibit plant infection by phytopathogens. In the present study, we focused on evaluating the colonization of soybean seedling-roots by Methylobacterium mesophilicum strain SR1.6/6. We focused on the identification of the key genes involved in the initial step of soybean colonization by methylotrophic bacteria, which includes the plant exudate recognition and adaptation by planktonic bacteria. Visualization by scanning electron microscopy revealed that M. mesophilicum SR1.6/6 colonizes soybean roots surface effectively at 48 h after inoculation, suggesting a mechanism for root recognition and adaptation before this period. The colonization proceeds by the development of a mature biofilm on roots at 96 h after inoculation. Transcriptomic analysis of the planktonic bacteria (with plant) revealed the expression of several genes involved in membrane transport, thus confirming an initial metabolic activation of bacterial responses when in the presence of plant root exudates. Moreover, antioxidant genes were mostly expressed during the interaction with the plant exudates. Further evaluation of stress- and methylotrophic-related genes expression by qPCR showed that glutathione peroxidase and glutathione synthetase genes were up-regulated during the Methylobacterium-soybean interaction. These findings support that glutathione (GSH) is potentially a key molecule involved in cellular detoxification during plant root colonization. In addition to methylotrophic metabolism, antioxidant genes, mainly glutathione-related genes, play a key role during soybean exudate recognition and adaptation, the first step in bacterial colonization.




Similar content being viewed by others
References
Abanda-Nkpwatt D, Musch M, Tschiersch J, Boettner M, Schwab W (2006) Molecular interaction between Methylobacterium extorquens and seedlings: growth promotion, methanol consumption, and localization of the methanol emission site. J Exp Bot 57:4025–4032
Almeida DM, Dini-Andreote, F, Camargo Neves AA, Jucá Ramos RT, Andreote FD, Carneiro AR, Oliveira de Souza Lima A, Caracciolo Gomes de Sá PH, Ribeiro Barbosa MS, Araújo WL, Silva A (2013) Draft Genome Sequence of Methylobacterium mesophilicum Strain SR1.6/6, Isolated from Citrus sinensis.Genome Announc 20 1(3):13
Allocati N, Federici L, Masulli M, Di Ilio C (2008) Glutathione transferases in bacteria. FEBS J 276:58–75
Alquéres S, Meneses C, Rouws L, Rothballer M, Baldani I, Schmid M, Hartmann A (2013) The bacterial superoxide dismutase and glutathione reductase are crucial for endophytic colonization of rice roots by Gluconacetobacter diazotrophicus PAL5. Mol Plant Microbe Interact 26:937–945
Alscher RG, Donahue JL, Cramer CL (1997) Reactive oxygen species and antioxidants: relationships in green cells. Physiol Plantarum 100:224–233
Anda M, Ikeda S, Eda S, Okubo T, Sato S, Tabata S, Mitsui H, Minamisawa K (2011) Isolation and genetic characterization of Aurantimonas and Methylobacterium strains from stems of hypernodulated soybeans. Microbes Environ 26:172–180
Andreote FD, Lacava PT, Gai CS, Araújo WL, Maccheroni W Jr, Overbeek LSV, Elsas JDV, Azevedo JL (2006) Model plants for studying the interaction between Methylobacterium mesophilicum and Xylella fastidiosa. Can J Microbiol 52:419–426
Araújo WL, Saridakis HO, Barroso PAV, Aguilar-Vildoso CI, Azevedo JL (2001) Variability and interactions between endophytic bacteria and fungi isolated from leaf tissues of citrus rootstocks. Can J Microbiol 47:229–236
Araújo WL, Marcon J, MaccheroniJr W, Elsas JDV, Azevedo JL (2002) Diversity of endophytic bacterial populations and their interaction with Xylella fastidiosa in citrus plant. Appl Environ Microbiol 68:4906–4914
Barnard AML, Bowdwn SD, Burr T, Coulthurst SJ, Monson RE, Salmond GPC (2007) Quorum sensing, virulence and secondary metabolite production in plant soft-rotting bacteria. Philos Trans R Soc B 362:1165–1183
Blankenberg D, Von Kuster G, Coraor N, Ananda G, Lazarus R, Mangan M, Nekrutenko A, Taylor J (2010) Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol Chapter 19:Unit 19.10.1–21
Camilios-Neto D, Bonato P, Wassem R, Tadra-Sfeir MZ, Brusamarello-Santos LC, Valdameri G, Donatti L, Faoro H, Weiss VA, Chubatsu LS, Pedrosa FO, Souza EM (2014) Dual RNA-seq transcriptional analysis of wheat roots colonized by Azospirillum brasilense reveals up-regulation of nutrient acquisition and cell cycle genes. BMC Genom 15:378
Cervantes-Martínez J, Lopez-Díaz S, Rodriguez-Garay B (2004) Detection of the effects of in Weber var. azul by laser-induced fluorescence. Plant Sci 166:889–892
Conesa A, Götz S, García-Gomez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Davis MJ, French WJ, Schaad NW (1981) Axenic culture of the bacteria associated with phony disease of peach and plum leaf scald. Curr Microbiol 5:311–316
Dourado MN, Andreote FD, Dini-Andreote F, Conti R, Araújo JM, Araújo WL (2012a) Analysis of 16S rRNA and mxaF genes revealing insights into Methylobacterium niche-specific plant association. Genet Mol Biol 35:142–148
Dourado MN, Ferreira A, Araújo WL, Azevedo JL, Lacava PT (2012b) The diversity of endophytic methylotrophic bacteria in an oil-contaminated and an oil-free mangrove ecosystem and their tolerance to heavy metals. Biotechnol Res Int 2012:759865
Dourado MN, Bogas AC, Pomini AM, Andreote FD, Quecine MC, Marsaioli AJ, Araújo WL (2013) Methylobacterium-plant interaction genes regulated by plant exudate and quorum sensing molecules. Braz J Microbiol 44:1331–1339
Fernandez D, Tisserant E, Talhinhas P, Azinheira H, Vieira A, Petitot AS, Loureiro A, Poulain J, Da Silva C, Silva Mdo C, Duplessis S (2012) 454-pyrosequencing of Coffea arabica leaves infected by the rust fungus Hemileia vastatrix reveals in planta-expressed pathogen-secreted proteins and plant functions in a late compatible plant-rust interaction. Mol Plant Pathol 13(1):17–37
Frendo P, Matamoros MA, Alloing G, Becana M (2013) Thiol-based redox signaling in the nitrogen-fixing symbiosis. Front Plant Sci 26(4):376
Gai CS, Lacava PT, Quecine MC, Auriac MC, Lopes JRS, Araújo WL, Miller TA, Azevedo JL (2009) Transmission of Methylobacterium mesophilicum by Bucephalogonia xanthophis for paratansgenic control strategy of citrus variegated chlorosis. J Microbiol 47:448–454
Ghelfi A, Gaziola SA, Cia MC, Chabregas SM, Falco MC, Kuser-Falcão PR, Azevedo RA (2011) Cloning, expression, molecular modelling and docking analysis of glutathione transferase from Saccharum officinarum. Ann Appl Biol 159:267–280
Giardine B, Riemer C, Hardison RC, Burhans R, Elnitski L, Shah P, Zhang Y, Blankenberg D, Albert I, Taylor J, Miller W, Kent WJ, Nekrutenko A (2005) Galaxy: a platform for interactive large-scale genome analysis. Genome Res 15:1451–1455
Goecks J, Nekrutenko A, Taylor J (2010) The galaxy team. galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol 11(8):R86
Gonzalez-Pasayo R, Martinez-Romero E (2000) Multiresistance genes of Rhizobium etli CFN42. Mol Plant Microbe In 13:572–577
Gourion B, Rossignol M, Vorholt JA (2006) A proteomic study of Methylobacterium extorquens reveals a response regulator essential for epiphytic growth. Proc Natl Acad Sci USA 103(35):13186–13191
Gourion B, Francez-Charlot A, Vorholt JA (2008) PhyR is involved in the general stress response of Methylobacterium extorquens AM1. J Bacteriol 190(3):1027–1035
Gourion B, Sulser S, Frunzke J, Francez-Charlot A, Stiefel P, Pessi G, Vorholt JA, Fischer HM (2009) The PhyR-sigma (EcfG) signalling cascade is involved in stress response and symbiotic efficiency in Bradyrhizobium japonicum. Mol Microbiol 73:291–305
Gratão PL, Polle A, Lea PJ, Azevedo RA (2005) Making the life of heavy metal-stressed plants a little easier. Funct Plant Biol 32:481
Holmes DE, Nevin KP, O’neil RA, Ward JE, Adams LA, Woodard TL, Vrionis HA, Lovley DR (2005) Potential for quantifying expression of the Geobacteraceae citrate synthase gene to assess the activity of Geobacteraceae in the subsurface and on current-harvesting electrodes. Appl Environ Microbiol 71:6870–6877
Jacobs JM, Babujee L, Meng F, Milling A, Allen C (2012) The in planta transcriptome of Ralstonia solanacearum: conserved physiological and virulence strategies during bacterial wilt of tomato. mBio 3:e00114–e00116
Jourand P, Renier A, Rapior S, Miana de Faria S, Prin Y, Galiana A, Giraud E, Dreyfus B (2005) Role of methylotrophy during symbiosis between Methylobacterium nodulans and Crotalaria podocarpa. Mol Plant Microbe Interact 18:1061–1068
Lacava PT, Araújo WL, Marcon J, Maccheroni W Jr, Azevedo JL (2004) Interaction between endophytic bacteria from citrus plants and the phytopathogenic bacteria Xylella fastidiosa, casual agent of citrus-variegated clorosis. Lett Appl Microbiol 39:55–59
Lacava PT, Li WB, Araújo WL, Azevedo JL, Hartung JS (2006) Rapid, specific and quantitative assays for the detection of the endophytic bacterium Methylobacterium mesophilicum in plants. J Microbiol Methods 65:535–541
Lacava PT, Silva-Stenico ME, Araújo WL, Simionato AVC, Carrilho E, Tsai SM, Azevedo JL (2008) Detection of siderophores in endophytic bacteria Methylobacterium spp. associated with Xylella fastidiosa subsp. Pauca. Pesq Agropec Bras 43:521–528
Langlois P, Bourassa S, Poirier GG, Beaulieu C (2003) Identification of Streptomyces coeli color proteins that are differentially expressed in the presence of plant material. Appl Environ Microbiol 69:1884–1889
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Larrainzar E, Wienkoop S, WeckwerthW Ladrera R, Arrese-Igor C, Esther M et al (2007) Medicago truncatula root nodule proteome analysis reveals differential plant and bacteroid responses to drought stress. Plant Physiol 144:1495–1507
Lassmann T, Hayashizaki Y, Daub CO (2011) SAMStat: monitoring biases in next generation sequencing data. Bioinformatics 27:130–131
LeFevre GH, Hozalski RM, Novak PJ (2013) Root exudate enhanced contaminant desorption: an abiotic contribution to the rhizosphere effect. Environ Sci Technol 47:11545–11553
Leo JC, Grin I, Linke D (2012) Type V secretion: mechanism(s) of autotransport through the bacterial outer membrane. Philos Trans R SocB 367:1088–1101
Li XG, Zhang TL, Wang XX, Hua K, Zhao L, Han ZM (2013) The composition of root exudates from two different resistant peanut cultivars and their effects on the growth of soil-borne pathogen. Int J Biol Sci 9:164–173
Lindemann A, Koch M, Pessi G, Müller AJ, Balsiger S, Hennecke H, Fischer HM (2010) Host-specific symbiotic requirement of BdeAB, a RegR-controlled RND-type efflux system in Bradyrhizobium japonicum. FEMS Microbiol Lett 312(2):184–191
Lohse M, Bolger AM, Nagel A, Fernie AR, Lunn JE, Stitt M, Usadel B (2012) RobiNA: a user-friendly, integrated software solution for RNA-Seq-based transcriptomics. Nucleic Acids Res 40:W622–W627
Madahaiyan M, Reddy BVS, Anandham R, Senthilkumar M, Poonguzhali S, Sundaram SP, Sa TM (2006) Plant growth-promoting Methylobacterium induces defense responses in groundnut (Arachis hypogaea L.) compared with rot pathogens. Curr Microbiol 53:270–276
Madhaiyan M, Poonguzhali S (2014) Methylobacterium pseudosasae sp. nov., a pink-pigmented, facultatively methylotrophic bacterium isolated from the bamboo phyllosphere. Anton Leeuw 105:367–376
Madhaiyan M, Poonguzhali S, Sa T (2007) Metal tolerating methylotrophic bacteria reduces nickel and cadmium toxicity and promotes plant growth of tomato (Lycopersicon esculentum L). Chemosphere 69:220–228
Madhaiyan M, Poonguzhali S, Senthilkumar M, Sundaram SP, Sa T (2009) Nodulation and plant-growth promotion by methylotrophic bacteria isolated from tropical legumes. Microbiol Res 164:114–120
Margis R, Dunand C, Teixeira FK, Margis-Pinheiro M (2008) Glutathione peroxidase family—an evolutionary overview. FEBS J 275:3959–3970
Masip L, Veeravalli K, Georgiou G (2006) The many faces of glutathione in bacteria. Antioxid Redox Signal 8:753–762
Mccord JM, Fridovich I (1969) Superoxide Dismutase an enzymic function for erythrocuprein (Hemocuprein). J Biol Chem 244:6049–6055
Meena KK, Kumar M, Kalyuzhnaya MG, Yandigeri MS, Singh DP, Saxena AK, Arora DK (2012) Epiphytic pink-pigmented methylotrophic bacteria enhance germination and seedling growth of wheat (Triticum aestivum) by producing phytohormone. Anton Leeuw 101:777–786
Morris CJ, Biville F, Turlin E, Lee E, Ellermann K, Fan WH, Ramamoorthi R, Springer AL, Lidstrom ME (1994) Isolation, phenotypic characterization, and complementation Analysis of Mutants of Methylobacterium extorquens AM1 unable to synthesize Pyrroloquinoline Quinone and Sequences of pqqD, pqqG, and pqqC. J Bacteriol 176:1746–1755
Pauly N, Pucciariello C, Mandon K, Innocenti G, Jamet A, Baudouin E, Hérouart D, Frendo P, Puppo A (2006) Reactive oxygen and nitrogen species and glutathione: key players in the legume-Rhizobium symbiosis. J Exp Bot 57:1769–1776
Petti C, Khan M, Doohan F (2010) Lipid transfer proteins and protease inhibitors as key factors in the priming of barley responses to Fusarium head blight disease by a biocontrol strain of Pseudomonas fluorescens. Funct Integr Genomics 10(4):619–627
Pomini AM, Cruz PLR, Gai C, Araújo WL, Marsaioli AJ (2009) Long-chain acyl-homoserine lactones from Methylobacterium mesophilicum: synthesis and absolute configuration. J Nat Prod 72:2130–2134
Rehman A, Anjum MS (2011) Multiple metal tolerance and biosorption of cadmium by Candida tropicalis isolated from industrial effluents: glutathione as detoxifying agent. Environ Monit Assess 174:585–595
Roberts A, Trapnell C, Donaghey J, Rinn JL, Pachter L (2011) Improving RNA-Seq expression estimates by correcting for fragment bias. Genome Biol 12:R22
Rose Figura JM, Puehringer S, Schwarzenbacher R, Toyama H, Klinman JP (2011) Characterization of a protein-generated O2 binding pocket in PqqC, a cofactor less oxidase catalyzing the final step in PQQ production. Biochemistry 50:1556–1566
Rossetto PB, Dourado MN, Quecine MC, Andreote FD, Araújo WL, Azevedo JL, Pizzirani-Kleiner AA (2011) Specific plant induced biofilm formation in Methylobacterium species. Braz J Microbiol 42:878–883
Sanches-Contreras M, Bauer WD, Gao M, Robinson JB, Downie A (2007) Quorum-sensing regulation in rhizobia and its role in symbiotic interactions with legumes. Philos Trans R Soc B362:1149–1163
Sarma AD, Emerich DW (2005) Global protein expression pattern of Bradyrhizobium japonicum bacteroids: aprelude to functional proteomics. Proteomics 5:4170–4184
Scrutton NS, Berry A, Perham RN (1987) Purification and characterization of glutathione reductase encoded by a cloned and over-expressed gene in Escherichia coli. Biochem J 245:875–880
Skovran E, Palmer AD, Rountree AM, Good NM, Lidstrom ME (2011) XoxF is required for expression of methanol dehydrogenase in Methylobacterium extorquens AM1. J Bacteriol 193:6032–6038
Sy A, Giraud E, Jourand P, Garcia N, Willems A, De Lajudie P, Prin Y, Neyra M, Gillis M, Bivin-Masson C, Dreyfus B (2001) Methylotrophic Methylobacterium bacteria nodulate and fix nitrogen in symbiosis with legumes. J Bacteriol 183:214–220
Sy A, Timmers AC, Knief C, Vorholt JA (2005) Methylotrophic metabolism is advantageous for Methylobacterium extorquens during colonization of Medicago truncatula under competitive conditions. Appl Environ Microbiol 71:7245–7252
Toyama H, Anthony C, Lidstrom ME (1998) Construction of insertion and deletion mxa mutants of Methylobacterium extorquens AM1 by electroporation. FEMS Microbiol Lett 166:1–7
Trapnell C, Williams BA, Pertea G, Mortazavi AM, Kwan G, van Baren MJ, Salzberg SL, Wold B, Pachter L (2010) Transcript assembly and abundance estimation from RNA-Seq reveals thousands of new transcripts and switching among isoforms. Nat Biotechnol 28:511–515
Vadassery J, Oelmüller R (2009) Calcium signaling in pathogenic and beneficial plant microbe interactions: what can we learn from the interaction between Piriformospora indica and Arabidopsis thaliana. Plant Signal Behav 4(11):1024–1027
Van Aken B, Yoon JM, Schnoor JL (2004) Biodegradation of nitro-substituted explosives 2,4,6-trinitrotoluene, hexahydro-1,3,5-trinitro-1,3,5-triazine, and octahydro-1,3,5,7-tetranitro-1,3,5-tetrazocine by a phytosymbiotic Methylobacterium sp. associated with poplar tissues (Populus deltoides). Appl Environ Microbiol 70:508–517
Van Dien SJ, Okubo Y, Hough MT, Korotkova N, Taitano T, Lidstrom ME (2003) Reconstruction of C3 and C4 metabolism in Methylobacterium extorquens AM1 using transposon mutagenesis. Microbiology 149:601–609
Vuilleumier M, Pagni S (2002) The elusive roles of bacterial glutathione S- transferases: new lessons from genomes. Appl Microbiol Biotechnol 58:138–146
White EW, Winans SC (2007) Cell-cell communication in the plant pathogen Agrobacterium tumefaciens. Philos Trans R Soc B362:1135–1148
Acknowledgments
This work was supported by a Grant from the FAPESP -Foundation for Research Assistance of São Paulo State, Brazil (Proc. 12/24217-6). We thank CAPES for the fellowship to J.K.S.L.; CNPq for the fellowship to D.S.S. (Proc. 167502/2013-1) and also FAPESP for the fellowship to M.N.D. (Proc. 2013/17314-8) and A.A.C.N. (Proc. 2011/14290-5) and CNPq by the productivity fellowship (1B) to W.L.A.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table 1
Gene expression values (FPKM) of the transcripts in Control (without plant) treatment by RNA Seq (DOCX 258 kb)
Table 2
Gene expression values (FPKM) of the transcripts in Planktonic (with plant) treatment by RNA Seq (DOCX 430 kb)
Fig. 1
M-SOY 8001 soybean seedling with root/shoot used in this work; detail of the root surface A) root without bacteria and B) root with M. mesophilicum SR1.6/6 72 h after inoculation (DOCX 3913 kb)
Rights and permissions
About this article
Cite this article
Araújo, W.L., Santos, D.S., Dini-Andreote, F. et al. Genes related to antioxidant metabolism are involved in Methylobacterium mesophilicum-soybean interaction. Antonie van Leeuwenhoek 108, 951–963 (2015). https://doi.org/10.1007/s10482-015-0548-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0548-6