Abstract
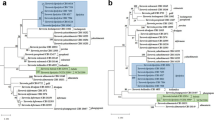
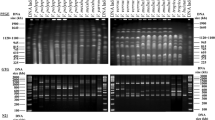
Eleven yeast strains representing two hitherto undescribed species were isolated from different kinds of meat samples in Hungary and one from the sediment of a tropical freshwater river in Southeastern Brazil. The analysis of the sequences of their large subunit rRNA gene D1/D2 domain and the internal transcribed spacer (ITS) regions placed the two new species in the Yarrowia clade. Some of the seven strains representing the first new species can mate and give rise to asci and form ascospores embedded in capsular material, which qualifies it as the third teleomorph species of the Yarrowia clade. The name Yarrowia porcina sp. nov. (type strain: NCAIM Y.02100T = CBS 12935T = NRRL Y-63669T, allotype strain UFMG-RD131A = CBS 12932A) is proposed for this new yeast species, which, based on physiological characters, is indistinguishable from Yarrowia lipolytica and some other species of the genus. Considerable intraspecific variability was detected among the sequences of the large subunit rRNA gene D1/D2 domains of the seven strains. The variability among the D1/D2 sequences exceeded the divergence observed among the ITS sequences and in some cases more than 1 % substitution among the D1/D2 sequences was detected. The conspecificity of these strains was supported by the low (0–3 substitutions) sequence divergence among their ITS sequences, the result of a parsimony network analysis utilizing the concatenated ITS and D1/D2 sequences and also by the fingerprint patterns generated by microsatellite primed PCR. No ascospore formation was observed in the group of the other five strains representing the second new species. These strains shared identical D1/D2 and ITS sequences. Yarrowia bubula f.a., sp. nov. (type strain: NCAIM Y.01998T = CBS 12934T = NRRL Y-63668T) is proposed to accommodate these strains.







Similar content being viewed by others
References
Atlas RM (1993) In: Parks LC (ed) Handbook of microbiological media. CRC Press, Boca Raton, p 968
Chang C-F, Chen C-C, Lee C-F, Liu S-M (2013) Identifying and characterizing Yarrowia keelungensis sp. nov., an oil-degrading yeast isolated from the sea surface microlayer. Antonie van Leeuwenhoek. doi:10.1007/s10482-013-0033-z
Chen Y-C, Eisner JD, Kattar MM, Rassoulian-Barrett SL, LaFe K, Yarfitz SL, Limaye AP, Cookson BT (2000) Identification of medically important yeasts using PCR-based detection of DNA sequence polymorphisms in the internal transcribed spacer 2 region of the rRNA genes. J Clin Microbiol 38:2302–2310
Clement M, Posada D, Crandall K (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Coelho MAZ, Amaral PFF, Belo I (2010) Yarrowia lipolytica: an industrial workhorse. In: Méndes-Vilas A (ed) Current research, technology and education topics in applied microbiology and microbial biotechnology. Formatex Research Center, Badajoz, pp 930–944
Daniel H-M, Vrancken G, Takrama JF, Camu N, De Vos P, De Vuyst L (2009) Yeast diversity of Ghanaian cocoa bean heap fermentations. FEMS Yeast Res 9:774–783
de Barros Lopes M, Soden A, Martens AL, Henschke PA, Langridge P (1998) Differentiation and species identification on yeasts using PCR. Int J Syst Bacteriol 48:279–286
Deák T (2008) Detection and enumeration. Handbook of food spoilage yeasts, 2nd edn. CRC Press, Boca Raton, pp 203–220
Encinas JP, López-Díaz TM, García-López ML, Otero A, Moreno B (2000) Yeast populations on Spanish fermented sausages. Meat Sci 54:203–208
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Ferreira AD, Viljoen BC (2003) Yeasts as adjunct starters in matured cheddar cheese. Int J Food Microbiol 86:131–140
Fonseca Á, Scorzetti G, Fell JW (2000) Diversity in the yeast Cryptococcus albidus and related species as revealed by ribosomal DNA sequence analysis. Can J Microbiol 46:7–27
Groenewald M, Smith MTh (2013) The teleomorph state of Candida deformans Langeron and Guerra and description of Yarrowia yakushimensis comb. nov. Antonie Van Leeuwenhoek 103:1023–1028
Groenewald M, Boekhout T, Neuvéglise C, Gaillardin C, van Dijck PWM, Wyss M (2013) Yarrowia lipolytica: safety assessment of an oleaginous yeast with a great industrial potential. Crit Rev Microbiol. doi:10.3109/1040841X.2013.770386
Hart MW, Sunday J (2007) Things fall apart: biological species form unconnected parsimony networks. Biol Lett 3:509–512
Ismail SAS, Deák T, Abd El-Rahman HA, Yassien MA, Beuchat LR (2000) Presence and changes in populations of yeasts on raw and processed poultry products stored at refrigeration temperature. Int J Food Microbiol 62:113–121
Knutsen AK, Robert V, Poot GA, Epping W, Figge M, Holst-Jensen A, Skaar I, Smith MTh (2007) Polyphasic re-examination of Yarrowia lipolytica strains and the description of three novel Candida species: Candida oslonensis sp. nov., Candida alimentaria sp. nov. and Candida hollandica sp. nov. Int J Syst Evol Microbiol 57:2426–2435
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 73:331–371
Kurtzman CP, Fell JW, Boekhout T, Robert V (2011) Methods for isolation, phenotypic characterization and maintenance of yeasts. In: Kurtzman CP, Fell JW, Boekhout T (eds) The yeasts, a taxonomic study, 5th edn. Elsevier, Amsterdam, pp 97–107
Lachance MA (2012) In defense of yeast sexual life cycles: the forma asexualis: An informal proposal. Yeast Newsl 61:24–25
Lachance MA, Bowles JM, Starmer WT, Barker JSF (1999) Kodamaea kakaduensis and Candida tolerans, two new ascomycetous yeast species from Australian Hibiscus flowers. Can J Microbiol 45:172–177
Lachance MA, Daniel HM, Meyer W, Prasad GS, Gautam SP, Boundy-Mills K (2003) The D1/D2 domain of the large-subunit rDNA of the yeast species Clavispora lusitaniae is unusually polymorphic. FEMS Yeast Res 4:253–258
Lachance MA, Dobson J, Wijayanayaka DN, Smith AM (2010) The use of parsimony network analysis for the formal delineation of phylogenetic species of yeasts: Candida apicola, Candida azyma, and Candida parazyma sp. nov., cosmopolitan yeasts associated with floricolous insects. Antonie Van Leeuwenhoek 97:155–170
Lachance MA, Wijayanayaka TM, Bundus JD, Wijayanayaka DN (2011) Ribosomal DNA sequence polymorphism and the delineation of two ascosporic yeast species: Metschnikowia agaves and Starmerella bombicola. FEMS Yeast Res 11:324–333
Mayoral MB, Martin R, Sanz A, Hernández PE, González I, Garcia T (2005) Detection of Kluyveromyces marxianus and other spoilage yeasts in yoghurt using a PCR-culture technique. Int J Food Microbiol 105:27–34
McIlvaine TC (1921) A buffer solution for colorimetric comparison. J Biol Chem 49:183–186
Mcneill J, Barrie FR, Buck WR, Demoulin V, Greuter W, Hawksworth DL, Herendeen PS, Knapp S, Marhold K, Prado J, Prud’homme Van Reine WF, Smith GF, Wiersema JH, Turland NJ (2012) International code of nomenclature for algae, fungi, and plants (Melbourne Code). Koeltz scientific books. http://www.iapt-taxon.org/nomen/main.php. Accessed 15 Sept 2013
Medeiros AO, Kohler LM, Hamdan JS, Missagia BS, Barbosa FAR, Rosa CA (2008) Diversity and antifungal susceptibility of yeasts from tropical freshwater environments in Southeastern Brazil. Water Res 42:3921–3929
Mirbagheri M, Nahvi I, Emtiazi G, Mafakher L, Darvishi F (2012) Taxonomic characterization and potential biotechnological applications of Yarrowia lipolytica isolated from meat and its products. Jundishapur J Microbiol 5:346–351
Nagy E, Niss M, Dlauchy D, Arneborg N, Nielsen DS, Péter G (2013) Yarrowia divulgata f.a., sp. nov., a yeast species from animal related and marine sources. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.057208-0
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Patrignani F, Iucci L, Vallicelli M, Guerzoni ME, Gardini F, Lanciotti R (2007) Role of surfaceinoculated Debaryomyces hansenii and Yarrowia lipolytica strains in dried fermented sausage manufacture. Part 1: evaluation of their effects on microbial evolution, lipolytic and proteolytic patterns. Meat Sci 75:676–686
Péter G, Tornai-Lehoczki J, Dlauchy D (2009) Candida ogatae sp. nov., an anamorphic member of the Kuraishia clade. FEMS Yeast Res 9:328–333
Pincus DH, Salkin IF, Hurd NJ, Levy IL, Kemna MA (1988) Modification of potassium nitrate assimilation test for identification of clinically important yeasts. J Clin Microbiol 26:366–368
Samson RA, Hoekstra ES, Frisvad JC (eds) (2004) Introduction to food- and airborne fungi, 7th edn. Centraalbureau voor Schimmelcultures, Utrecht
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Vasdinyei R, Deák T (2003) Characterization of yeast isolates originating from Hungarian dairy products using traditional and molecular identification techniques. Int J Food Microbiol 86:123–130
Wickerham LJ, Kurtzman CP, Herman AI (1970) Sexuality in Candida lipolytica. In: Ahearn DG (ed) Recent trends in yeast research, vol 1. Georgia State University, Atlanta, pp 81–92
Wyder MT, Bachmann HP, Puhan Z (1999) Role of selected yeasts in cheese ripening: an evaluation in foil wrapped raclette cheese. Lebensm Wiss Technol 32:333–343
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A greedy algorithm for aligning DNA sequences. J Comput Biol 7:203–214
Acknowledgments
This study was partly supported by the State Secretariat for Higher Education, Hungarian Ministry of Human Resources and the Doctoral School of Food Sciences, Corvinus University of Budapest. This work was also funded by the Conselho Nacional de Desenvolvimento Cientifico e Tecnológico (CNPq – Brazil), Fundação do Amparo a Pesquisa do Estado de Minas Gerais (FAPEMIG), the Financiadora de Estudos e Projetos (FINEP, Brazil, process number 2084/07).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nagy, E., Dlauchy, D., Medeiros, A.O. et al. Yarrowia porcina sp. nov. and Yarrowia bubula f.a. sp. nov., two yeast species from meat and river sediment. Antonie van Leeuwenhoek 105, 697–707 (2014). https://doi.org/10.1007/s10482-014-0125-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-014-0125-4