Abstract
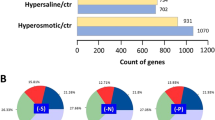
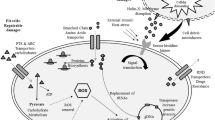
Sulfate-reducing bacteria such as Desulfovibrio vulgaris have developed a set of responses that allow them to survive in hostile environments. To obtain further knowledge of the protective mechanisms employed by D.␣vulgaris in response to oxidative stress and heat shock, we performed a genome-wide transcriptomic analysis to determine the cellular responses to both stimuli. The results showed that 130 genes were responsive to oxidative stress, while 427 genes were responsive to heat-shock. Functional analyses suggested that the genes regulated were involved in a variety of cellular functions. Amino acid biosynthetic pathways were induced by both oxidative stress and heat shock treatments, while fatty acid metabolism, purine and cofactor biosynthesis were induced by heat shock only. The rubrerythrin gene (rbr) was up-regulated in response to oxidative stress, suggesting an important role for this protein in the oxidative damage resistance response in D. vulgaris. In addition, thioredoxin reductase (trxB) was also responsive to oxidative stress, suggesting that the thiol-specific redox system might also be involved in oxidative protection in this organism. In contrast, the expression of rubredoxin oxidoreductase (rbo), superoxide dismutase (sodB) and catalase (katA) genes were not regulated in response to oxidative stress. Comparison of cellular responses to oxidative stress and heat-shock allowed the identification of 66 genes that showed a similar drastic response to both environmental perturbations, implying that these genes might be part of the general stress response (GSR) network in D. vulgaris. This hypothesis was further supported by the identification of a conserved motif upstream of these stress-responsive genes.
Similar content being viewed by others
References
Arsene F., Tomoyasu T., Bukau B. (2000) The heat shock response of Escherichia coli. Int. J. Food Microbiol. 55:3–9
Baumgarten A., Redenius I., Kranczoch J., Cypionka H. (2001) Periplasmic oxygen reduction by Desulfovibrio species. Arch. Microbiol. 176: 306–309
Carty S.M., Colbeau A., Vignais P.M., Larson T.J. (1994) Identification of the rpmF-plsX-fabH genes of Rhodobacter capsulatus. FEMS Microbiol. Lett. 118: 227–231
Crooks G.E., Hon G., Chandonia J.M., Brenner S.E. (2004) WebLogo: a sequence logo generator. Genome Res. 14: 1188–1190
Cypionka H. (2000) Oxygen respiration by Desulfovibrio species. Ann. Rev. Microbiol. 54: 827–848
Darmon E., Noone D., Masson A., Bron S., Kuipers O.P., Devine K.M., van Dijl J.M. (2002) A novel class of heat and secretion stress-responsive genes is controlled by the autoregulated CssRS two-component system of Bacillus subtilis. J. Bacteriol. 184: 5661–5671
Dos Santos W.G., Paheco I., Liu M.-Y., Texeira M., Xavier A.V., Legall J. (2000) Purification and characterization of an iron superoxide dismutase and a catalase from the sulfate-reducing bacterium Desulfovibrio gigas. J. Bacteriol. 182: 796–804
Fareleira P., Santos B.S., Antonio C., Moradas-Ferreira P., LeGall J., Xavier A.V., Santos H. (2003) Response of a strict anaerobe to oxygen: survival strategies in Desulfovibrio gigas. Microbiology 149: 1513–1522
Fournier M., Zhang Y., Wildschut J.D., Dolla A., Voordouw J.K., Schriemer D.C. and Voordouw G. (2003) Function of oxygen resistance proteins in the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris hildenborough. J. Bacteriol. 185:71–79
Fournier M., Dermoun Z., Durand M.C. and Dolla A. (2004) A new function of the Desulfovibrio vulgaris Hildenborough [Fe] hydrogenase in the protection against oxidative stress. J. Biol. Chem. 279: 1787–1793
Fournier M., Aubert C., Dermoun Z., Durand M.C., Moinier D., Dolla A. (2005) Response of the anaerobe Desulfovibrio vulgaris Hildenborough to oxidative conditions: proteome and transcript analysis. Biochimie 88: 85–94
Fu R., Wall J.D., Voordouw G. (1994) DcrA, a c-type heme-containing methyl-accepting protein from Desulfovibrio vulgaris Hildenborough, senses the oxygen concentration or redox potential of the environment. J. Bacteriol. 176: 344–350
Gasch A.P., Spellman P.T., Kao C.M., Carmel-Harel O., Eisen M.B., Storz G., Botstein D., Brown P.O. (2000) Genomic expression programs in the response of yeast cells to environmental changes. Mol. Biol. Cell 11:4241–4257
Gilmore M.E., Bandyopadhyay D., Dean A.M., Linnstaedt S.D., Popham D.L. (2004) Production of muramic delta-lactam in Bacillus subtilis spore peptidoglycan. J. Bacteriol. 186: 80–89
Greenberg J.T., Demple B. (1989) A global response induced in Escherichia coli by redox-cycling agents overlaps with that induced by peroxide stress. J. Bacteriol. 171: 3933–3939
Grovc J., Busby S., Cole J. (1996) The role of the genes nrf EFG and ccmFH in cytochrome c biosynthesis in Escherichia coli. Mol. Gen. Genet. 252: 332–341
Guedon E., Moore C.M., Que Q., Wang T., Ye R.W., Helmann J.D. (2003) The global transcriptional response of Bacillus subtilis to manganese involves the MntR, Fur, TnrA and sigmaB regulons. Mol. Microbiol. 49: 1477–1491
Gustavsson N., Kokke B.P., Harndahl U., Silow M., Bechtold U., Poghosyan Z., Murphy D., Boelens W.C., Sundby C. (2002) A peptide methionine sulfoxide reductase highly expressed in photosynthetic tissue in Arabidopsis thaliana can protect the chaperone-like activity of a chloroplast-localized small heat shock protein. Plant J. 29:545–553
Hecker M., Schumann W., Volker U. (1996) Heat-shock and general stress response in Bacillus subtilis. Mol. Microbiol. 19:417–428
Heidelberg J.F., Seshadri R., Haveman S.A., Hemme C.L., Paulsen I.T., Kolonay J.F., Eisen J.A., Ward N., Methe B., Brinkac L.M., Daugherty S.C., Deboy R.T., Dodson R.J., Durkin A.S., Madupu R., Nelson W.C., Sullivan S.A., Fouts D., Haft D.H., Selengut J., Peterson J.D., Davidsen T.M., Zafar N., Zhou L., Radune D., Dimitrov G., Hance M., Tran K., Khouri H., Gill J., Utterback T.R., Feldblyum T.V., Wall J.D., Voordouw G., Fraser C.M. (2004) The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. Nat. Biotechnol. 22:554–559
Helmann J.D., Wu M.F., Kobel P.A., Gamo F.J., Wilson M., Morshedi M.M., Navre M., Paddon C. (2001) Global transcriptional response of Bacillus subtilis to heat shock. J. Bacteriol. 183:7318–7328
Hughes J.D., Estep P.W., Tavazoie S., Church G.M. (2000) Computational identification of cis-regulatory elements associated with groups of functionally related genes in Saccharomyces cerevisiae. J. Mol. Biol. 296:1205–1214
Jenney F.E., Verhagen M.F.J.M., Cui X., Adams M.W.W. (1999) Anaerobic microbes: oxygen detoxification without superoxide dismutase. Science 286:306–309
Keon R.G., Fu R., Voordouw G. (1997) Deletion of two downstream genes alters expression of the hmc operon of Desulfovibrio vulgaris subsp. vulgaris Hildenborough. Arch. Microbiol. 167:376–383
Kobayashi N., McEntee K. (1993) Identification of cis and trans components of a novel heat shock stress regulatory pathway in Saccharomyces cerevisiae. Mol. Cell Biol. 13:248–256
Krekeler D., Teske A., Cypionka H. (1997) Strategies of sulfate-reducing bacteria to escape oxygen stress in a cyanobacterial mat. FEMS Microbiol. Ecol. 25:89–96
Laplace J.M., Hartke A., Giard J.C., Auffray Y. (2000) Cloning, characterization and expression of an Enterococcus faecalis gene responsive to heavy metals. Appl. Microbiol. Biotechnol. 53:685–689
Laport M.S., de Castro A.C., Villardo A., Lemos J.A., Bastos M.C., Giambiagi-deMarval M. (2001) Expression of the major heat shock proteins DnaK and GroEL in Streptococcus pyogenes: a comparison to Enterococcus faecalis and Staphylococcus aureus. Curr. Microbiol. 42:264–268
Lawrence C.E., Altschul S.F., Bogouski M.S., Liu J.S., Neuwald A.F., Wooten J.C. (1993) Detecting subtle sequence signals: a Gibbs Sampling Strategy for multiple alignment. Science 262:208–214
Lemos R.S., Gomes C.M., Santana M., LeGall J., Xavier A.V., Teixeira M. (2001) The ‘strict’ anaerobe Desulfovibrio gigas contains a membrane-bound oxygen-reducing respiratory chain. FEBS Lett. 496:40–43
Leverrier P., Vissers J.P., Rouault A., Boyaval P., Jan G. (2004) Mass spectrometry proteomic analysis of stress adaptation reveals both common and distinct response pathways in Propionibacterium freudenreichii. Arch. Microbiol. 181:215–230
Li B., Nierras C.R., Warner J.R. (1999) Transcriptional elements involved in the repression of ribosomal protein synthesis. Mol. Cell Biol. 19:5393–5404
Lombard M., Fontecave M., Touati D., Niviere V. (2000) Reaction of the desulfoferrodoxin from Desulfoarculus baarsii with superoxide anion. Evidence for a superoxide reductase activity. J. Biol. Chem. 275:115–121
Lumppio H.L., Shenvi N.V., Garg R.P., Summers A.O., Kurtz Jr. D.M. (1997) A rubrerythrin operon and nigerythrin gene in Desulfovibrio vulgaris (Hildenborough). J. Bacteriol. 179:4607–4615
Lumppio H.L., Shenvi N.V., Summers A.O., Voordouw G., Kurtz D.M. Jr. (2001) Rubrerythrin and rubredoxin oxidoreductase in Desulfovibrio vulgaris: a novel oxidative stress protection system. J. Bacteriol. 183:101–108
Martinez-Pastor M.T., Marchler G., Schuller C., Marchler-Bauer A., Ruis H., Estruch F. (1996) The Saccharomyces cerevisiae zinc finger proteins Msn2p and Msn4p are required for transcriptional induction through the stress response element (STRE). EMBO J. 15: 2227–2235
McCluskey J., Hinds J., Husain S., Witney A. and Mitchell T.J. (2004) A two-component system that controls the expression of pneumococcal surface antigen A (PsaA) and regulates virulence and resistance to oxidative stress in Streptococcus pneumoniae. Mol. Microbiol. 51:1661–1675
Morbidoni H.R., de Mendoza D., Cronan J.E. Jr. (1996) Bacillus subtilis acyl carrier protein is encoded in a cluster of lipid biosynthesis genes. J. Bacteriol. 178: 4794–4800
Moskovitz J., Rahman M.A., Strassman J., Yancey S.O., Kushner S.R., Brot N., Weissbach H. (1995) Escherichia coli peptide methionine sulfoxide reductase gene: regulation of expression and role in protecting against oxidative damage. J. Bacteriol. 177: 502–507
Moskovitz J., Berlett B.S., Poston J.M., Stadtman E.R. (1997) The yeast peptide-methionine sulfoxide reductase functions as an antioxidant in vivo. Proc. Natl. Acad. Sci. USA 94:9585–9589
Moskovitz J., Flescher E., Berlett B.S., Azare J., Poston J.M., Stadtman E.R. (1998) Over-expression of peptide-methionine sulfoxide reductase in Saccharomyces cerevisiae and human T cells provides them with high resistance to oxidative stress. Proc. Natl. Acad. Sci. USA 95:14071–14075
Muro-Pastor A.M., Herrero A., Flores E. (2001) Nitrogen-regulated group 2 sigma factor from Synechocystis sp. strain PCC 6803 involved in survival under nitrogen stress. J. Bacteriol. 183: 1090–1095
Ng W.L., Kazmierczak K.M., Robertson G.T., Gilmour R., Winkler M.E. (2003) Transcriptional regulation and signature patterns revealed by microarray analyses of Streptococcus pneumoniae R6 challenged with sublethal concentrations of translation inhibitors. J. Bacteriol. 185:359–370
Nuwaysir E.F., Huang W., Albert T.J., Singh J., Nuwaysir K., Pitas A., Richmond T., Gorski T., Berg J.P., Ballin J., McCormick M., Norton J., Pollock T., Sumwalt T., Butcher L., Porter D., Molla M., Hall C., Blattner F., Sussman M.R., Wallace R.L., Cerrina F., Green R.D. (2002) Gene expression analysis using oligonucleotide arrays produced by maskless photolithography. Genome Res. 12: 1749–1755
Ohmori S., Nawata Y., Kiyono K., Murata H., Tsuboi S., Ikeda M., Akagi R., Morohashi K.I., Ono B. (1999) Saccharomyces cerevisiae cultured under aerobic and anaerobic conditions: air-level oxygen stress and protection against stress. Biochim. Biophys. Acta 1472:587–594
Page M.D., Pearce D.A., Norris H.A., Ferguson S.J. (1997) The Paracoccus denitrificans ccmA, B and C genes: cloning and sequencing, and analysis of the potential of their products to form a haem or apo- c-type cytochrome transporter. Microbiology 143:563–576
Petersohn A., Brigulla M., Haas S., Hoheisel J.D., Volker U., Hecker M. (2001) Global analysis of the general stress response of Bacillus subtilis. J. Bacteriol. 183:5617–5631
Rince A., Flahaut S., Auffray Y. (2000) Identification of general stress genes in Enterococcus faecalis. Int. J. Food Microbiol. 55:87–91
Rodionov D.A., Dubchak I., Arkin A., Alm E., Gelfand M.S. (2004) Reconstruction of regulatory and metabolic pathways in metal-reducing delta-proteobacteria. Genome Biol. 5:R90
Santos H., Fareleira P., Xavier A.V., Chen L., Liu M.Y., LeGall J. (1993) Aerobic metabolism of carbon reserves by the “obligate anaerobe” Desulfovibrio gigas. Biochem. Biophys. Res. Commun. 195:551–557
Saxena R.K., Pandey P.K., Bisen P.S. (2002) Physiological and biochemical alterations in Anabaena 7120 under iron stress. Indian J. Exp. Biol. 40:594–599
Saxild H.H., Nygaard P. (1988) Gene-enzyme relationships of the purine biosynthetic pathway in Bacillus subtilis. Mol. Gen. Genet. 211:160–167
Scharf C., Riethdorf S., Ernst H., Engelmann S., Volker U., Hecker M. (1998) Thioredoxin is an essential protein induced by multiple stresses in Bacillus subtilis. J. Bacteriol. 180:1869–1877
Schlindwein C., Giordano G., Santini C.L., Mandrand M.A. (1990) Identification and expression of the Escherichia coli fdhD and fdhE genes, which are involved in the formation of respiratory formate dehydrogenase. J. Bacteriol. 172:6112–6121
Silva G., Legall J., Xavier A.V., Texeira M., Rodriguez-Pousada C. (2001) Molecular characterization of Desulfovibrio gigas neelaredoxin, a protein involved in oxygen detoxification in anaerobes. J. Bacteriol. 183:4413–4420
Singh-Gasson S., Green R.D., Yue Y., Nelson C., Blattner F., Sussman M.R., Cerrina F. (1999) Maskless fabrication of light-directed oligonucleotide microarrays using a digital micromirror array. Nat. Biotechnol. 17:974–978
Tamburro A., Allocati N., Masulli M., Rotilio D., Di Ilio C., Favaloro B. (2001) Bacterial peptide methionine sulphoxide reductase: co-induction with glutathione S-transferase during chemical stress conditions. Biochem. J. 360: 675–681
Thompson W., Rouchka E.C., Lawrence C.E. (2003) Gibbs recursive sampler: finding transcription factor binding sites. Nucleic Acids Res. 31:3580–3585
Uziel O., Borovok I., Schreiber R., Cohen G., Aharonowitz Y. (2004) Transcriptional regulation of the Staphylococcus aureus thioredoxin and thioredoxin reductase genes in response to oxygen and disulfide stress. J. Bacteriol. 186:326–334
Voordouw G. (1995) The Genus Desulfovibrio: the Centennial. Appl. Environ. Microbiol. 61:2813–2819
Voordouw J.K., Voordouw G. (1998) Deletion of the rbo gene increases the oxygen sensitivity of the sulfate-reducing bacterium Desulfovibrio vulgaris Hildenborough. Appl. Environ. Microbiol. 64: 2882–2887
Wingender E., Dietze P., Karas H., Knuppel R. (1996) TRANSFAC: a database on transcription factors and their DNA binding sites. Nucleic Acids Res. 24:238–241
Zhang W., Culley D.E., Scholten J.C.M., Hogan M., Vitiritti L. and Brockman F.J. 2005. Global transcriptomic analysis of Desulfovibrio vulgaris on different electron donors. Antonie van Leeuwenhoek. In press
Acknowledgements
The research described in this paper was conducted under the Laboratory Directed Research and Development LDRD Program at the Pacific Northwest National Laboratory, a multi-program national laboratory operated by Battelle for the U.S. Department of Energy under Contract DE-AC056-76RLO1830.
Author information
Authors and Affiliations
Corresponding author
Additional information
The complete microarray data including ratio change for all genes in response to heat shock and oxidative stress could be provided upon request.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, W., Culley, D.E., Hogan, M. et al. Oxidative stress and heat-shock responses in Desulfovibrio vulgaris by genome-wide transcriptomic analysis. Antonie Van Leeuwenhoek 90, 41–55 (2006). https://doi.org/10.1007/s10482-006-9059-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-006-9059-9