Abstract
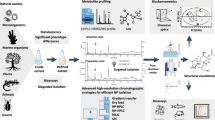
Mass spectral resolution for clean mass spectra is a very challenging task in non-targeted hyphenated techniques, especially for massive amounts of such data from herbs. In this work, the regions of interest (ROI) strategy was used to compress liquid chromatography–high-resolution mass spectrometry (LC–HRMS) data. For single or overlapped peaks, a rule of “first in, first out” was intended for visual recognition of features that could be of relevance in ROI curves. In the light of the two embedded peaks in a local segment, an improved selective ion analysis after ROI compression (ROI–SIA) possesses the main advantages of being rapid, automatic and parameter free. From the clean mass spectra above, we can estimate their correct feature sets that belong to the same compound. Tandem MS spectra were further determined, and peak annotations were rediscovered in known natural products by the interpretation of known MS/MS fragmentations. These procedures were tested in the analysis of the LC–HR–MS data coming from phytochemicals in Ligusticum chuanxiong with satisfactory results. The in vivo prototypes were also characterized in mice plasma after oral administration of the herbal extract. These analytical results provide an important basis for the identification of active ingredients, also used for a further investigation on the efficacy of different technologies.






Similar content being viewed by others
References
Cheng X, Chen B, Pan Y, Guo L, Feng W, Dong Y (2018) Chromatographia 81:157–165. https://doi.org/10.1007/s10337-017-3446-4
Gorrochategui E, Jaumot J, Lacorte S, Tauler R (2016) TrAC Trend Anal Chem 82:425–442. https://doi.org/10.1016/j.trac.2016.07.004
He M, Lv HY, Li YP, Vicente Gonçalves CM, Dong NP, Pan LS, Liu PL, Liang YZ (2014) Anal Methods 6:2239–2246. https://doi.org/10.1039/c3ay41861h
Tao J, Zhao M, Jiang S, Zhang W, Xu B, Duan J (2018) J Pharmaceut Biomed 161:254–261. https://doi.org/10.1016/j.jpba.2018.08.051
Yang Y, Yin XJ, Guo HM, Wang RL, Song R, Tian Y, Zhang ZJ (2014) Chin J Nat Med 12:542–553. https://doi.org/10.1016/S1875-5364(14)60084-4
Forsberg EM, Huan T, Rinehart D, Benton HP, Warth B, Hilmers B, Siuzdak G (2018) Nat Protoc 13:633–651. https://doi.org/10.1038/nprot.2017.151
Lommen A (2012) Metabolomics 8:719–726. https://doi.org/10.1007/s11306-011-0369-1
Pluskal T, Castillo S, Villar-Briones A, Orešič M (2010) BMC Bioinform 11:395. https://doi.org/10.1186/1471-2105-11-395
Lima KMG, Bedia C, Tauler R (2014) Microchem J 117:255–261. https://doi.org/10.1016/j.microc.2014.07.010
Dalmau N, Bedia C, Tauler R (2018) Anal Chim Acta 1025:80–91. https://doi.org/10.1016/j.aca.2018.04.003
Gorrochategui E, Jaumot J, Lacorte S, Tauler R (2016) TrAC Trend Anal Chem 82:425–442. https://doi.org/10.1016/j.trac.2016.07.004
Jaumot J, Juan A, Tauler R (2015) Chemom Intell Lab Sys 140:1–12. https://doi.org/10.1016/j.chemolab.2014.10.003
He M, Wu H, Nie J, Yan P, Yang T, Yang Z, Pei R (2017) J Pharm Biomed Anal 146:37–47. https://doi.org/10.1016/j.jpba.2017.07.065
Li CM, Guo YQ, Dong XL, Li H, Wang B, Wu JH, Wong MS, Chan SW (2014) Food Funct 5:2475–2485. https://doi.org/10.1039/c4fo00211c
Zeng Z, Xie R, Tan L, Zhang T (2011) Chin J Appl Chem 28:956–962. https://doi.org/10.1090/s0002-9939-2011-10775-5
Chi Y, Zhang Z, Li C, Liu Q, Yan P, Welz-Biermann U (2011) Green Chem 13:666–670. https://doi.org/10.1039/c0gc00864h
Li HB, Chen F (2004) J Chromatogr A 1047:249–253. https://doi.org/10.1016/s0021-9673(04)01103-3
Sun YY, Li SF, Song HT, Tian SJ (2006) Chem Eng 34:60–63. https://doi.org/10.1007/s10589-005-3075-y
Gorrochategui E, Jaumot J, Tauler R (2015) Protocol Exchange. https://doi.org/10.1038/protex.2015.102
Kind T, Fiehn O (2007) BMC Bioinform 8:105. https://doi.org/10.1186/1471-2105-8-105
Kind T, Fiehn O (2010) Bioanal Rev 2:23–60. https://doi.org/10.1007/s12566-010-0015-9
Zhang Q, Huo M, Zhang Y, Qiao Y, Gao X (2018) J Chromatogra A 1552:17–28. https://doi.org/10.1016/j.chroma.2018.03.055
Chen Z, Zhang C, Gao F, Fu Q, Fu C, He Y, Zhang J (2018) Food Chem Toxicol 119:309–325. https://doi.org/10.1016/j.fct.2018.02.050
Yan R, Ko NL, Li SL, Tam YK, Lin G (2008) Pharmacokinetics and metabolism of ligustilide, a major bioactive component in Rhizoma Chuanxiong in the rat. Drug Metab Dispos 36:400–408. https://doi.org/10.1124/dmd.107.017707
He M, Yan P, Yang ZY, Zhang ZM, Yang TB, Hong L (2018) J Chromatogra B 1079:41–50. https://doi.org/10.1016/j.jchromb.2018.01.040
Acknowledgements
This work is financially supported by Hunan 2011 Collaborative Innovation Center of Chemical Engineering & Technology with Environmental Benignity and Effective Resource Utilization, Hunan Province Natural Science Fund (no. 2016JJ4085), the key project of Hunan Provincial Education Department (18A055). The study met the approval of the university’s review board.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No potential conflict of interest was reported by the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
He, M., Peng, G., Xie, F. et al. Liquid Chromatography–High-Resolution Mass Spectrometry with ROI Strategy for Non-targeted Analysis of the In Vivo/In Vitro Ingredients Coming from Ligusticum chuanxiong hort. Chromatographia 82, 1069–1077 (2019). https://doi.org/10.1007/s10337-019-03740-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10337-019-03740-x