Abstract
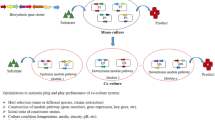
Natural products (NPs) produced by bacteria and fungi are often used as therapeutic agents due to their complex structures and wide range of bioactivities. Enzymes that build NPs are encoded by co-localized biosynthetic gene clusters (BGCs), and genome sequencing has recently revealed that many BGCs are “silent” under standard laboratory conditions. There are numerous methods used to activate “silent” BGCs that rely either upon altering culture conditions or genetic modification. In this review, we discuss several recent microbial cultivation methods that have been used to expand the scope of NPs accessible in the laboratory. These approaches are divided into three categories: addition of a physical scaffold, addition of small molecule elicitors, and co-cultivation with another microbe.

















Similar content being viewed by others
References
Afiyatullov SS, Zhuravleva OI, Antonov AS et al (2018) Prenylated indole alkaloids from co-culture of marine-derived fungi Aspergillus sulphureus and Isaria felina. J Antibiot 71:846–853. https://doi.org/10.1038/s41429-018-0072-9
Akhter N, Liu Y, Auckloo BN et al (2018) Stress-driven discovery of new angucycline-type antibiotics from a marine Streptomyces pratensis NA-ZhouS1. Mar Drugs 16:331. https://doi.org/10.3390/md16090331
Akone SH, Mándi A, Kurtán T et al (2016) Inducing secondary metabolite production by the endophytic fungus Chaetomium sp. through fungal–bacterial co-culture and epigenetic modification. Tetrahedron 72:6340–6347. https://doi.org/10.1016/j.tet.2016.08.022
Alonso S, Rendueles M, Díaz M (2017) Tunable decoupled overproduction of lactobionic acid in Pseudomonas taetrolens through temperature-control strategies. Process Biochem 58:9–16. https://doi.org/10.1016/j.procbio.2017.04.034
Ancheeva E, Mándi A, Király SB et al (2018) Chaetolines A and B, Pyrano[3,2-f]isoquinoline alkaloids from cultivation of Chaetomium sp. in the presence of autoclaved Pseudomonas aeruginosa. J Nat Prod 81:2392–2398. https://doi.org/10.1021/acs.jnatprod.8b00373
Ancheeva E, Küppers L, Akone SH et al (2017) Expanding the metabolic profile of the fungus Chaetomium sp. through co-culture with autoclaved Pseudomonas aeruginosa. Eur J Org Chem 2017:3256–3264. https://doi.org/10.1002/ejoc.201700288
Anjum K, Sadiq I, Chen L et al (2018) Novel antifungal janthinopolyenemycins A and B from a co-culture of marine-associated Janthinobacterium spp. ZZ145 and ZZ148. Tetrahedron Lett 59:3490–3494. https://doi.org/10.1016/j.tetlet.2018.08.022
Auckloo BN, Pan C, Akhter N et al (2017) Stress-driven discovery of novel cryptic antibiotics from a marine fungus Penicillium sp. BB1122. Front Microbiol 8:1450. https://doi.org/10.3389/fmicb.2017.01450
Bai J, Mu R, Dou M et al (2018) Epigenetic modification in histone deacetylase deletion strain of Calcarisporium arbuscula leads to diverse diterpenoids. Acta Pharm Sin B 8:687–697. https://doi.org/10.1016/j.apsb.2017.12.012
Ballouz S, Francis AR, Lan R, Tanaka MM (2010) Conditions for the evolution of gene clusters in bacterial genomes. PLoS Comput Biol 6:e1000672. https://doi.org/10.1371/journal.pcbi.1000672
Baltz RH (2017) Gifted microbes for genome mining and natural product discovery. J Ind Microbiol Biotechnol 44:573–588. https://doi.org/10.1007/s10295-016-1815-x
Barnhart MM, Lynem J, Chapman MR (2006) GlcNAc-6P levels modulate the expression of Curli fibers by Escherichia coli. J Bacteriol 188:5212–5219. https://doi.org/10.1128/JB.00234-06
Biggins JB, Ternei MA, Brady SF (2012) Malleilactone, a polyketide synthase-derived virulence factor encoded by the cryptic secondary metabolome of Burkholderia pseudomallei group pathogens. J Am Chem Soc 134:13192–13195. https://doi.org/10.1021/ja3052156
Boffa LC, Vidali G, Mann RS, Allfrey VG (1978) Suppression of histone deacetylation in vivo and in vitro by sodium butyrate. J Biol Chem 253:3364–3366
Boruta T, Bizukojc M (2019) Application of aluminum oxide nanoparticles in Aspergillus terreus cultivations: evaluating the effects on lovastatin production and fungal morphology. Biomed Res Int 2019:1–11. https://doi.org/10.1155/2019/5832496
Bosello M, Zeyadi M, Kraas FI et al (2013) Structural Characterization of the heterobactin siderophores from Rhodococcus erythropolis PR4 and elucidation of their biosynthetic machinery. J Nat Prod 76:2282–2290. https://doi.org/10.1021/np4006579
Brinkmann C, Kearns P, Evans-Illidge E, Kurtbӧke D (2017) Diversity and bioactivity of marine bacteria associated with the sponges Candidaspongia flabellata and Rhopaloeides odorabile from the Great Barrier Reef in Australia. Diversity 9:39. https://doi.org/10.3390/d9030039
Craney A, Ozimok C, Pimentel-Elardo SM et al (2012) Chemical perturbation of secondary metabolism demonstrates important links to primary metabolism. Chem Biol 19:1020–1027. https://doi.org/10.1016/j.chembiol.2012.06.013
Dashti Y, Grkovic T, Abdelmohsen UR et al (2017) Actinomycete metabolome induction/suppression with N-acetylglucosamine. J Nat Prod 80:828–836. https://doi.org/10.1021/acs.jnatprod.6b00673
Davie JR (2003) Inhibition of histone deacetylase activity by butyrate. J Nutr 133:2485S–2493S. https://doi.org/10.1093/jn/133.7.2485S
Davies J, Spiegelman GB, Yim G (2006) The world of subinhibitory antibiotic concentrations. Curr Opin Microbiol 9:445–453. https://doi.org/10.1016/j.mib.2006.08.006
de Oca-Mejía MM, Castillo-Juárez I, Martínez-Vázquez M et al (2015) Influence of quorum sensing in multiple phenotypes of the bacterial pathogen Chromobacterium violaceum. Pathog Dis 73:1–4. https://doi.org/10.1093/femspd/ftu019
Du C, van Wezel GP (2018) Mining for microbial gems: integrating proteomics in the postgenomic natural product discovery pipeline. Proteomics 18:1700332. https://doi.org/10.1002/pmic.201700332
El-Hawary SS, Sayed AM, Mohammed R et al (2018) New Pim-1 kinase inhibitor from the co-culture of two sponge-associated Actinomycetes. Front Chem 6:538. https://doi.org/10.3389/fchem.2018.00538
Everest GJ, Meyers PR (2011) Evaluation of the antibiotic biosynthetic potential of the genus Amycolatopsis and description of Amycolatopsis circi sp. nov., Amycolatopsis equina sp. nov. and Amycolatopsis hippodromi sp. nov. J Appl Microbiol 111:300–311. https://doi.org/10.1111/j.1365-2672.2011.05058.x
Feng X, He C, Jiao L et al (2019) Analysis of differential expression proteins reveals the key pathway in response to heat stress in Alicyclobacillus acidoterrestris DSM 3922T. Food Microbiol 80:77–84. https://doi.org/10.1016/j.fm.2019.01.003
Gilmore SA, Naseem S, Konopka JB, Sil A (2013) N-Acetylglucosamine (GlcNAc) triggers a rapid, temperature-responsive morphogenetic program in thermally dimorphic fungi. PLoS Genet. https://doi.org/10.1371/journal.pgen.1003799
Gonciarz J, Bizukojc M (2014) Adding talc microparticles to Aspergillus terreus ATCC 20542 preculture decreases fungal pellet size and improves lovastatin production. Eng Life Sci 14:190–200. https://doi.org/10.1002/elsc.201300055
Hegemann JD, Zimmermann M, Xie X, Marahiel MA (2015) Lasso peptides: an intriguing class of bacterial natural products. Acc Chem Res 48:1909–1919. https://doi.org/10.1021/acs.accounts.5b00156
Jiang J, Sun Y-F, Tang X et al (2018) Alkaline pH shock enhanced production of validamycin A in fermentation of Streptomyces hygroscopicus. Bioresour Technol 249:234–240. https://doi.org/10.1016/j.biortech.2017.10.012
Ju K-S, Zhang X, Elliot MA (2017) New kid on the block: LmbU expands the repertoire of specialized metabolic regulators in Streptomyces. J Bacteriol. https://doi.org/10.1128/JB.00559-17
Kawai K, Wang G, Okamoto S, Ochi K (2007) The rare earth, scandium, causes antibiotic overproduction in Streptomyces spp. FEMS Microbiol Lett 274:311–315. https://doi.org/10.1111/j.1574-6968.2007.00846.x
Keilhofer N, Nachtigall J, Kulik A et al (2018) Streptomyces AcH 505 triggers production of a salicylic acid analogue in the fungal pathogen Heterobasidion abietinum that enhances infection of Norway spruce seedlings. Antonie Van Leeuwenhoek 111:691–704. https://doi.org/10.1007/s10482-018-1017-9
Kharel MK, Pahari P, Shepherd MD et al (2012) Angucyclines: biosynthesis, mode-of-action, new natural products, and synthesis. Nat Prod Rep 29:264–325. https://doi.org/10.1039/c1np00068c
Kim SD, Park SK, Cho JY et al (2006) Surfactin C inhibits platelet aggregation. J Pharm Pharmacol 58:867–870. https://doi.org/10.1211/jpp.58.6.0018
Lehmann LH, Jebessa ZH, Kreusser MM et al (2018) A proteolytic fragment of histone deacetylase 4 protects the heart from failure by regulating the hexosamine biosynthetic pathway. Nat Med 24:62–72. https://doi.org/10.1038/nm.4452
Ligon BL (2004) Penicillin: its discovery and early development. Semin Pediatr Infect Dis 15:52–57. https://doi.org/10.1053/j.spid.2004.02.001
Luger K, Richmond TJ (1998) The histone tails of the nucleosome. Curr Opin Genet Dev 8:140–146
Manczinger M, Bocsik A, Kocsis GF et al (2015) The absence of N-acetyl-d-glucosamine causes attenuation of virulence of Candida albicans upon interaction with vaginal epithelial cells in vitro. Biomed Res Int. https://doi.org/10.1155/2015/398045
Mao D, Okada BK, Wu Y et al (2018) Recent advances in activating silent biosynthetic gene clusters in bacteria. Curr Opin Microbiol 45:156–163. https://doi.org/10.1016/j.mib.2018.05.001
Matilla MA, Daddaoua A, Chini A et al (2018) An auxin controls bacterial antibiotics production. Nucleic Acids Res 46:11229–11238. https://doi.org/10.1093/nar/gky766
Mo S, Kim J, Oh C-H (2013) Different effects of acidic pH shock on the prodiginine production in Streptomyces coelicolor M511 and SJM1 mutants. J Microbiol Biotechnol 23:1454–1459. https://doi.org/10.4014/jmb.1307.07067
Montaser R, Luesch H (2011) Marine natural products: a new wave of drugs? Future Med Chem 3:1475–1489. https://doi.org/10.4155/fmc.11.118
Moore JM, Bradshaw E, Seipke RF et al (2012) Chapter eighteen—use and discovery of chemical elicitors that stimulate biosynthetic gene clusters in Streptomyces bacteria. In: Hopwood DA (ed) methods in enzymology. Academic Press, Cambridge, pp 367–385
Motoyama T, Osada H (2016) Biosynthetic approaches to creating bioactive fungal metabolites: pathway engineering and activation of secondary metabolism. Bioorg Med Chem Lett 26:5843–5850. https://doi.org/10.1016/j.bmcl.2016.11.013
Moussian B (2008) The role of GlcNAc in formation and function of extracellular matrices. Comp Biochem Physiol B Biochem Mol Biol 149:215–226. https://doi.org/10.1016/j.cbpb.2007.10.009
Munshi A, Shafi G, Aliya N, Jyothy A (2009) Histone modifications dictate specific biological readouts. J Genet Genom 36:75–88. https://doi.org/10.1016/S1673-8527(08)60094-6
Nah H-J, Pyeon H-R, Kang S-H et al (2017) Cloning and heterologous expression of a large-sized natural product biosynthetic gene cluster in Streptomyces species. Front Microbiol 8:1–10. https://doi.org/10.3389/fmicb.2017.00394
Naseem S, Parrino SM, Buenten DM, Konopka JB (2012) Novel roles for GlcNAc in cell signaling. Commun Integr Biol 5:156–159. https://doi.org/10.4161/cib.19034
Ochi K, Hosaka T (2013) New strategies for drug discovery: activation of silent or weakly expressed microbial gene clusters. Appl Microbiol Biotechnol 97:87–98. https://doi.org/10.1007/s00253-012-4551-9
Okada BK, Seyedsayamdost MR (2017) Antibiotic dialogues: induction of silent biosynthetic gene clusters by exogenous small molecules. FEMS Microbiol Rev 41:19–33. https://doi.org/10.1093/femsre/fuw035
Özkaya FC, Peker Z, Camas M et al (2017) Marine fungi against aquaculture pathogens and induction of the activity via co-culture: biotechnology. CLEAN Soil Air Water 45:1700238. https://doi.org/10.1002/clen.201700238
Pacwa-Płociniczak M, Płaza GA, Piotrowska-Seget Z, Cameotra SS (2011) Environmental applications of biosurfactants: recent advances. Int J Mol Sci 12:633–654. https://doi.org/10.3390/ijms12010633
Pál C, Hurst LD (2004) Evidence against the selfish operon theory. Trends Genet 20:232–234. https://doi.org/10.1016/j.tig.2004.04.001
Park HB, Kwon HC, Lee C-H, Yang HO (2009) Glionitrin A, an antibiotic–antitumor metabolite derived from competitive interaction between abandoned mine microbes. J Nat Prod 72:248–252. https://doi.org/10.1021/np800606e
Pimentel-Elardo SM, Sørensen D, Ho L et al (2015) Activity-independent discovery of secondary metabolites using chemical elicitation and cheminformatic inference. ACS Chem Biol 10:2616–2623. https://doi.org/10.1021/acschembio.5b00612
Rateb ME, Hallyburton I, Houssen WE et al (2013) Induction of diverse secondary metabolites in Aspergillus fumigatus by microbial co-culture. RSC Adv 3:14444–14450. https://doi.org/10.1039/c3ra42378f
Ren H, Wang B, Zhao H (2017) Breaking the silence: new strategies for discovering novel natural products. Curr Opin Biotechnol 48:21–27. https://doi.org/10.1016/j.copbio.2017.02.008
Rigali S, Anderssen S, Naômé A, van Wezel GP (2018) Cracking the regulatory code of biosynthetic gene clusters as a strategy for natural product discovery. Biochem Pharmacol 153:24–34. https://doi.org/10.1016/j.bcp.2018.01.007
Romano S, Jackson S, Patry S, Dobson A (2018) Extending the “One Strain Many Compounds” (OSMAC) principle to marine microorganisms. Mar Drugs 16:244. https://doi.org/10.3390/md16070244
Ross AC, Gulland LES, Dorrestein PC, Moore BS (2015) Targeted capture and heterologous expression of the Pseudoalteromonas alterochromide gene cluster in Escherichia coli represents a promising natural product exploratory platform. ACS Synth Biol 4:414–420. https://doi.org/10.1021/sb500280q
Rutledge PJ, Challis GL (2015) Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat Rev Microbiol 13:509–523. https://doi.org/10.1038/nrmicro3496
Seyedsayamdost MR (2014) High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc Natl Acad Sci USA 111:7266–7271. https://doi.org/10.1073/pnas.1400019111
Sharma R, Jamwal V, Singh VP et al (2017) Revelation and cloning of valinomycin synthetase genes in Streptomyces lavendulae ACR-DA1 and their expression analysis under different fermentation and elicitation conditions. J Biotechnol 253:40–47. https://doi.org/10.1016/j.jbiotec.2017.05.008
Shi Y, Pan C, Auckloo BN et al (2017) Stress-driven discovery of a cryptic antibiotic produced by Streptomyces sp. WU20 from Kueishantao hydrothermal vent with an integrated metabolomics strategy. Appl Microbiol Biotechnol 101:1395–1408. https://doi.org/10.1007/s00253-016-7823-y
Shu C-H, Tseng K, Jaiswal R (2018) Effects of light intensity and wavelength on the production of lactobionic acid from whey by Pseudomonas taetrolens in batch cultures: effects of light intensity and wavelength on the production of lactobionic acid. J Chem Technol Biotechnol 93:1595–1600. https://doi.org/10.1002/jctb.5528
Skellam E (2018) Strategies for engineering natural product biosynthesis in fungi. Trends in Biotechnol. https://doi.org/10.1016/j.tibtech.2018.09.003
Slawson C, Housley MP, Hart GW (2006) O-GlcNAc cycling: how a single sugar post-translational modification is changing the way we think about signaling networks. J Cell Biochem 97:71–83. https://doi.org/10.1002/jcb.20676
Smetanina OF, Yurchenko AN, Afiyatullov SS et al (2012) Oxirapentyns B–D produced by a marine sediment-derived fungus Isaria felina (DC.) Fr. Phytochem Lett 5:165–169. https://doi.org/10.1016/j.phytol.2011.12.002
Stierle AA, Stierle DB, Decato D et al (2017) The berkeleylactones, antibiotic macrolides from fungal coculture. J Nat Prod 80:1150–1160. https://doi.org/10.1021/acs.jnatprod.7b00133
Sung A, Gromek S, Balunas M (2017) Upregulation and identification of antibiotic activity of a marine-derived Streptomyces sp. via co-cultures with human pathogens. Mar Drugs 15:250. https://doi.org/10.3390/md15080250
Tanaka Y, Hosaka T, Ochi K (2010) Rare earth elements activate the secondary metabolite–biosynthetic gene clusters in Streptomyces coelicolor A3(2). J Antibiot 63:477–481. https://doi.org/10.1038/ja.2010.53
Timmermans M, Paudel Y, Ross A (2017) Investigating the biosynthesis of natural products from marine Proteobacteria: a survey of molecules and strategies. Mar Drugs 15:235. https://doi.org/10.3390/md15080235
Timmermans ML, Picott KJ, Ucciferri L, Ross AC (2018) Culturing marine bacteria from the genus Pseudoalteromonas on a cotton scaffold alters secondary metabolite production. Microbiol Open. https://doi.org/10.1002/mbo3.724
Tong Y, Charusanti P, Zhang L et al (2015) CRISPR-Cas9 based engineering of actinomycetal genomes. ACS Synth Biol 4:1020–1029. https://doi.org/10.1021/acssynbio.5b00038
van der Meij A, Willemse J, Schneijderberg MA et al (2018) Inter- and intracellular colonization of Arabidopsis roots by endophytic actinobacteria and the impact of plant hormones on their antimicrobial activity. Antonie Van Leeuwenhoek 111:679–690. https://doi.org/10.1007/s10482-018-1014-z
Wang C, Huang D, Liang S (2018) Identification and metabolomic analysis of chemical elicitors for tacrolimus accumulation in Streptomyces tsukubaensis. Appl Microbiol Biotechnol 102:7541–7553. https://doi.org/10.1007/s00253-018-9177-0
Watrous JD, Dorrestein PC (2011) Imaging mass spectrometry in microbiology. Nat Rev Microbiol 9:683–694. https://doi.org/10.1038/nrmicro2634
Wright GD (2017) Opportunities for natural products in 21st century antibiotic discovery. Nat Prod Rep 34:694–701. https://doi.org/10.1039/C7NP00019G
Xu D, Han L, Li C et al (2018) Bioprospecting deep-sea Actinobacteria for novel anti-infective natural products. Front Microbiol. https://doi.org/10.3389/fmicb.2018.00787
Xu D, Nepal KK, Chen J et al (2018) Nocardiopsistins A–C: new angucyclines with anti-MRSA activity isolated from a marine sponge-derived Nocardiopsis sp. HB-J378. Synth Syst Biotechnol 3:246–251. https://doi.org/10.1016/j.synbio.2018.10.008
Xu F, Nazari B, Moon K et al (2017) Discovery of a cryptic antifungal compound from Streptomyces albus J1074 using high-throughput elicitor screens. J Am Chem Soc 139:9203–9212. https://doi.org/10.1021/jacs.7b02716
Xu F, Wu Y, Zhang C et al (2019) A genetics-free method for high-throughput discovery of cryptic microbial metabolites. Nat Chem Biol 15:161. https://doi.org/10.1038/s41589-018-0193-2
Zeng Z, Cai X, Wang P et al (2017) Biofilm formation and heat stress induce pyomelanin production in deep-sea Pseudoalteromonas sp. SM9913. Front Microbiol 8:1822. https://doi.org/10.3389/fmicb.2017.01822
Zhang MM, Wong FT, Wang Y et al (2017) CRISPR–Cas9 strategy for activation of silent Streptomyces biosynthetic gene clusters. Nat Chem Biol 13:607–609. https://doi.org/10.1038/nchembio.2341
Zhang Z, He X, Zhang G et al (2017) Inducing secondary metabolite production by combined culture of Talaromyces aculeatus and Penicillium variabile. J Nat Prod 80:3167–3171. https://doi.org/10.1021/acs.jnatprod.7b00417
Zhu H, Sandiford SK, van Wezel GP (2014) Triggers and cues that activate antibiotic production by actinomycetes. J Ind Microbiol Biotechnol 41:371–386. https://doi.org/10.1007/s10295-013-1309-z
Zhuravleva OI, Afiyatullov SS, Vishchuk OS et al (2012) Decumbenone C, a new cytotoxic decaline derivative from the marine fungus Aspergillus sulphureus KMM 4640. Arch Pharmacal Res 35:1757–1762. https://doi.org/10.1007/s12272-012-1007-9
Zou A, Liu J, Garamus VM et al (2010) Interaction between the natural lipopeptide [Glu1, Asp5] surfactin-C15 and hemoglobin in aqueous solution. Biomacromolecules 11:593–599. https://doi.org/10.1021/bm9011453
Zuck KM, Shipley S, Newman DJ (2011) Induced production of N-formyl alkaloids from Aspergillus fumigatus by co-culture with Streptomyces peucetius. J Nat Prod 74:1653–1657. https://doi.org/10.1021/np200255f
Acknowledgements
Funding is gratefully acknowledged from the Natural Sciences and Engineering Research Council of Canada (RGPIN-2015-06078) (Discovery Grant to ACR) and Queen’s University (Research Initiation Grant to ACR and Graduate award to LU).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Tomm, H.A., Ucciferri, L. & Ross, A.C. Advances in microbial culturing conditions to activate silent biosynthetic gene clusters for novel metabolite production. J Ind Microbiol Biotechnol 46, 1381–1400 (2019). https://doi.org/10.1007/s10295-019-02198-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-019-02198-y