Abstract
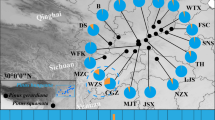
Range-wide genetic variation of Korean pine (Pinus koraiensis) was assessed using maternally inherited mtDNA and paternally inherited cpDNA for 16 natural populations throughout northeast Asia in order to study its phylogeographical history during the Quaternary. The cpDNA variation indicated that there was no difference between populations on the Asian continent and those in the Japanese archipelago. In contrast, the mtDNA variation indicated that there was significant difference between the populations from the two regions, with each region having a different lineage. The continental populations exhibited no diversity in the mtDNA examined despite the species’ current extensive range and large populations. Conversely, while the Korean pine is rare in Japan, the Japanese populations exhibited greater levels of mtDNA diversity (H T = 0.502). The higher mtDNA diversity and evidence from numerous Korean pine macrofossil remains dated to the Pleistocene and recovered various sites in Japan suggest that the Japanese archipelago once served as a refugium to a much larger Korean pine population with a more extensive range than is the case today. The presence of the single mtDNA haplotype across the Asian continent suggests that the present widespread populations could have expanded from a single refugium population after the last glacial periods.

Similar content being viewed by others
References
Aizawa M, Yoshimaru H, Saito H, Katsuki T, Kawahara T, Kitamura K, Shi F, Kaji M (2007) Phylogeography of a northeast Asian spruce, Picea jezoensis, inferred from genetic variation observed in organelle DNA markers. Mol Ecol 16:3393–3405
Artyukova EV, Kozyrenko MM, Gorovoy PG, Zhuravlev YN (2009) Plastid DNA variation on highly fragmented populations of Microbiota decussata Kom. (Cupressaceae), an endemic to Sikhote Alin Mountains. Genetica 137:201–212
Bai WN, Liao WJ, Zhang DY (2010) Nuclear and chloroplast DNA phylogeography reveal two refuge areas with asymmetrical gene flow in a temperate walnut tree from East Asia. New Phytol 188:892–901
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Burban C, Petit RJ (2003) Phylogeography of maritime pine inferred with organelle markers having contrasted inheritance. Mol Ecol 12:1487–1495
Chen K, Abbott RJ, Milne RI, Tian XM, Liu J (2008) Phylogeography of Pinus tabulaeformis Carr. (Pinaceae), a dominant species of coniferous forest in northern China. Mol Ecol 17:4276–4288
Chen MM, Feng F, Sui X, Li MH, Zhao D, Han S (2010) Constructing of a framework map for Pinus koraiensis Sieb. et Zucc. using SRAP, SSR and ISSR markers. Trees 24:685–693
Demesure B, Sodzi N, Petit RJ (1995) A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Mol Ecol 4:129–131
Dobson M, Kawamura Y (1998) Origin of the Japanese land mammal fauna: allocation of extant species to historically based categories. Quat Res 37:385–395
Du FK, Petit RJ, Liu JQ (2009) More introgression with less gene flow: chloroplast vs. mitochondrial DNA in the Picea asperata complex in China, and comparison with other conifers. Mol Ecol 18:1396–1407
Duff RJ, Nickrent DL (1999) Phylogenetic relationships of land plants using mitochondrial small-subunit rDNA sequences. Amer J Bot 86:372–386
Excoffier L, Ray N (2008) Surfing during population expansions promotes genetic revolutions and structuration. Trends Ecol Evol 23:347–351
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Feng F, Han S, Wang H (2006) Genetic diversity and genetic differentiation of natural Pinus koraiensis population. J For Res 17:21–24
Feng F, Chen M, Zhang D, Sui X, Han S (2009) Application of SRAP in the genetic diversity of Pinus koraiensis of different provenances. Afr J Biotech 8:1000–1008
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hallatschek O, Nelson DR (2008) Gene surfing in expanding populations. Theor Popul Biol 73:158–170
Hamilton M (1999) Four primer pairs for the amplification of chloroplast intergenic regions with intraspecific variation. Mol Ecol 8:521–523
Hamrick JL, Godt MLW, Sherman-Broyles SL (1992) Factors influencing levels of genetic diversity in woody plant species. New For 6:95–124
Harrison SP, Yu C, Takahara H, Prentice IC (2001) Palaeovegetation: diversity of temperate plants in East Asia. Nature 413:129–130
Hewitt G (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hu LJ, Uchiyama K, Shen HL, Saito Y, Tsuda Y, Ide Y (2008) Nuclear DNA microsatellites reveal genetic variation but a lack of phylogeographical structure in an endangered species, Fraxinus mandshurica, across North-east China. Ann Bot 102:195–205
Hutchins HE, Hutchins SA, Liu B (1996) The role of birds and mammals in Korean pine (Pinus koraiensis) regeneration dynamics. Oecologia 107:120–130
Ishikawa Y, Krestov PV, Namikawa K (1999) Disturbance history and tree establishment in old-growth Pinus koraiensis-hardwood forests in the Russian Far East. J Veg Sci 10:439–448
Jaramillo-Correa JP, Bousquet J, Beaulieu J, Isabel N, Perron M, Bouillé M (2003) Cross-species amplification of mitochondrial DNA sequence-tagged-site markers in conifers: the nature of polymorphism and variation within and among species in Picea. Theor Appl Genet 106:1353–1367
Jaramillo-Correa JP, Beaulieu J, Bousquet J (2004) Variation in mitochondrial DNA reveals multiple distant glacial refugia in black spruce (Picea mariana), a transcontinental North American conifer. Mol Ecol 13:2735–2747
Jaramillo-Correa JP, Beaulieu J, Ledig FT, Bousquet J (2006) Decoupled mitochondrial and chloroplast DNA population structure reveals Holocene collapse and population isolation in a threatened Mexican-endemic conifer. Mol Ecol 15:2787–2800
Kim ZS, Lee SW, Lim JH, Hwang JW, Kwon KW (1994) Genetic diversity and structure of natural populations of Pinus koraiensis (Sieb. et Zucc.) in Korea. For Genet 1:41–49
Kim ZS, Hwang JW, Lee SW, Yang C, Gorovoy PG (2005) Genetic variation of Korean Pine (Pinus koraiensis Sieb. et Zucc.) at allozyme and RAPD markers in Korea, China and Russia. Silvae Genet 54:235–246
Kong WS, Watts D (1993) Evergreen arboreal arctic-alpine and alpine plants and their environments. In: Kong WS, Watts D (eds) The plant geography of Korea with an emphasis on the alpine zones. Geobotany, vol 19. Kluwer, Dordrecht, pp 105–140
Kremenetski CV, Liu K, MacDonald GM (1998) The late Quaternary dynamics of pines in northeast Asia. In: Richardson DM (ed) Ecology and biogeography of Pinus. Cambridge University Press, Cambridge
Krestov PV (2003) Forest vegetation of easternmost Russia (Russian Far East). In: Kolbek J, Šrůtek M, Box EO (eds) Forest vegetation of northeast Asia. Geobotany, vol 28, Kluwer, Dordrecht, pp 93–180
Lascoux M, Palmé AE, Cheddadi R, Latta RG (2004) Impact of ice ages on the genetic structure of trees and shrubs. Phil Trans R Soc Lond B 359:197–207
Liepelt S, Bialozyt R, Ziegenhagen B (2002) Wind-dispersed pollen mediates postglacial gene flow among refugia. Proc Natl Acad Sci USA 100:10824–10829
Meng L, Yang R, Abbott RJ, Miehe G, Hu T, Liu J (2007) Mitochondrial and chloroplast phylogeography of Picea crassifolia Kom. (Pinaceae) in the Qinghai-Tibetan Plateau and adjacent highlands. Mol Ecol 16:4128–4137
Miki S (1956) Remains of Pinus koraiensis S. et Z. and associated remains in Japan. Bot Mag Tokyo 69:446–454
Miyaki M (1987) Seed dispersal of the Korean pine, Pinus koraiensis, by the red squirrel, Sciurus vulgaris. Ecol Res 2:147–157
Momohara A, Mizuno K, Okitsu S (1997) Plant macrofossil assemblages in a latest early Pleistocene cold stage from upper member of the Shobudani formation, southwestern Japan. Jpn J Hist Bot 5:29–37 (in Japanese with English summary)
Morita Y (2000) The vegetation history of the subalpine zone in Japan since Last Glacial period–Were the forest zones higher than they are at present during the Climatic Optimum period?–. Jpn J Hist Bot 9:3–20 (in Japanese with English summary)
Muir G, Schlötterer C (2005) Evidence for shared ancestral polymorphism rather than recurrent gene flow at microsatellite loci differentiating two hybridizing oaks (Quercus spp.). Mol Ecol 14:549–561
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Ohshima K (1990) The history of straits around the Japanese islands in the late-Quaternary. Quat Res 29:193–208 (in Japanese with English summary)
Okitsu S, Momohara A (1997) Distribution of Pinus koraiensis Sieb. et Zucc. in Japan. Tech Bull Fac Hort Chiba Univ 51:137–145 (in Japanese with English summary)
Pleines T, Jakob SS, Blattner FR (2009) Application of non-coding DNA regions in intraspecific analyses. Plant Syst Evol 282:281–294
Polezhaeva MA, Lascoux M, Semerikov VL (2010) Cytoplasmic DNA variation and biogeography of Larix Mill. in Northeast Asia. Mol Ecol 19:1239–1252
Pons O, Petit RJ (1996) Measuring and testing genetic differentiation with ordered vs. unordered alleles. Genetics 144:1237–1245
Potenko VV, Velikov AV (1998) Genetic diversity and differentiation of natural populations of Pinus koraiensis (Sieb. et Zucc.) in Russia. Silvae Genet 47:202–208
Qiu YX, Fu CX, Comes HP (2011) Plant molecular phylogeography in China and adjacent regions: tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol Phylogenet Evol 59:225–244
Ren G, Zhang L (1998) A preliminary mapped summary of Holocene pollen data for Northeast China. Quat Sci Rev 17:669–688
Stebich M, Mingram J, Han J, Liu J (2009) Late Pleistocene spread of (cool-)temperate forests in Northeast China and climate changes synchronous with the North Atlantic region. Global Planet Change 65:56–70
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tsutsui K, Suwa A, Sawada K, Kato T, Ohsawa TA, Watano Y (2009) Incongruence among mitochondrial, chloroplast and nuclear gene trees in Pinus subgenus Strobus (Pinaceae). J Plant Res 122:509–521
Wagner DB (1992) Nuclear, chloroplast and mitochondrial DNA polymorphisms as biochemical markers in population genetic analysis of forest trees. New For 6:373–390
Wang XQ, Tank DC, Sang T (2000) Phylogeny and divergence times in Pinaceae: evidence from three genomes. Mol Biol Evol 17:773–781
Yano M, Ishikari Teichitai Research Group (1967) Outline of the plants remains from the Quaternary deposits in the Ishikari Plain Hokkaido. Quat Res 7:41–48 (in Japanese with English summary)
Zhou YF, Abbott RJ, Jiang ZY, Du FK, Milne RI, Liu JQ (2010) Gene flow and species delimitation: a case study of two pine species with overlapping distributions in southeast China. Evolution 64:2342–2352
Acknowledgments
We wish to thank Y. Tsuda (Uppsala University), Y. Tsumura, S. Kikuchi, Y. Taguchi and Y. Moriguchi (Department of Forest Genetics, Forestry and Forest Products Research Institute, FFPRI) for their technical advice on experimental design and data analyses. We are grateful to Y.-P. Hong and K.N. Hong (Korea Forest Research Institute) and H. Sugita (FFPRI) for fruitful discussion. We also thank the regional offices of the Forestry Agency, Japan, and prefectural offices for permitting sampling. This study was financially supported by a Grant-in-Aid for Scientific Research (#19780111 and #21780141) from the Ministry of Education, Culture, Sports, Science and Technology.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aizawa, M., Kim, ZS. & Yoshimaru, H. Phylogeography of the Korean pine (Pinus koraiensis) in northeast Asia: inferences from organelle gene sequences. J Plant Res 125, 713–723 (2012). https://doi.org/10.1007/s10265-012-0488-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-012-0488-4