Abstract
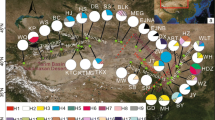
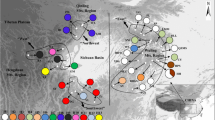
Genetic variation in the atpB–rbcL intergenic spacer region of chloroplast DNA (cpDNA) was investigated in Hygrophila pogonocalyx Hayata (Acanthaceae), an endangered and endemic species in Taiwan. In this aquatic species, seed dispersal from capsules via elasticity is constrained by gravity and is thereby confined within populations, resulting in limited gene flow between populations. In this study, a total of 849 bp of the cpDNA atpB–rbcL spacer were sequenced from eight populations of H. pogonocalyx. Nucleotide diversity in the cpDNA is low (θ=0.00343±0.00041). The distribution of genetic variation among populations agrees with an “isolation-by-distance” model. Two geographically correlated groups, the western and eastern regions, were identified in a neighbor-joining tree and a minimum-spanning network. Phylogeographical analyses based on the cpDNA network suggest that the present-day differentiation between western and eastern groups of H. pogonocalyx resulted from past fragmentation. The differentiation between eastern and western populations may be ascribed to isolation since the formation of the Central Mountain Range about 5 million years ago, which is consistent with the rate estimates based on a molecular clock of cpDNA.




Similar content being viewed by others
References
Avise JC (2000) Phylogeography: the history and formation of species. Harvard University Press, Cambridge
Avise JC, Arnold J, Ball RM, Bermingham E, Lamb T, Neigel JE, Reeb CA, Saunders NC (1987) Intraspecific phylogeography: the mitochondrial DNA bridge between population genetics and systematics. Annu Rev Ecol Syst 18:489–522
Bermingham E, Avise JC (1986) Molecular zoogeography of freshwater fishes in the southeastern United States. Genetics 113:939–965
Birky CW Jr (1995) Uniparental inheritance of mitochondrial and chloroplast genes: mechanisms and evolution. Proc Natl Acad Sci USA 92:11331–11338
Bossart JL, Prowell DP (1998) Genetic estimates of population structure and gene flow: limitations, lessons and new directions. Trends Ecol Evol 13:202–206
Chiang TY (2000) Lineage sorting accounting for the disassociation between chloroplast and mitochondrial lineages in oaks of southern France. Genome 43:1090–1094
Chiang TY, Schaal BA (2000a) The internal transcribed spacer 2 region of the nuclear ribosomal DNA and the phylogeny of the moss family Hylocomiaceae. Plant Syst Evol 224:127–137
Chiang TY, Schaal BA (2000b) Molecular evolution of the atpB–rbcL noncoding spacer of chloroplast DNA in the moss family Hylocomiaceae. Bot Bull Acad Sin 41:85–92
Chiang TY, Schaal BA, Peng CI (1998) Universal primers for amplification and sequencing a noncoding spacer between atpB and rbcL genes of chloroplast DNA. Bot Bull Acad Sin 39:245–250
Chiang TY, Chiang YC, Chen YJ, Chou CH, Havanond S, Hong TN, Huang S (2001) Phylogeography of Kandelia candel in East Asiatic mangroves based on nucleotide variation of chloroplast and mitochondrial DNAs. Mol Ecol 10:2697–2710
Chiang TY, Hung KH, Hsu TW, Wu WL (2004) Lineage sorting and phylogeography in Lithocarpus formosanus and L. dodonaeifolius (Fagaceae) from Taiwan. Ann MO Bot Gard 91:207–222
Cruzan MB, Templeton AR (2000) Paleoecology and coalescence: phylogeographic analysis of hypotheses from the fossil record. Trends Ecol Evol 15:491–496
Doyle JJ, Doyle JL (1987) A rapid isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Excoffier L, Smouse PE (1994) Using allele frequencies and geographic subdivision to reconstruct gene trees within a species: molecular variance parsimony. Genetics 136:343–359
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Felsenstein J (1993) PHYLIP (Phylogeny Inference Package) version 3.6a2. Distributed by the author. Department of Genetics, University of Washington, Seattle
Frankham R, Ballou JD, Briscoe DA (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge, pp 175–196
Futuyma DJ (1998) Evolutionary biology, 3rd edn. Sinauer, Sunderland
Ge XJ, Chiang YC, Chou CH, Chiang TY (2002) Nested clade analysis of Dunnia sinensis (Rubiaceae), a monotypic genus from China based on organelle DNA sequences. Conserv Genet 3:351–362
Grassly NC, Holmes EC (1997) A likelihood method for the detection of selection and recombination using nucleotide sequences. Mol Biol Evol 9:366–369
Hamrick JL, Godt MJ (1989) Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding and genetic resources. Sinauer, Sunderland, pp 43–63
Hanski IA, Gilpin ME (1997) Metapopulation biology: ecology, genetics, and evolution. Academic, Orlando
Hawkins JA, Harris SA (1998) RAPD characterization of two neotropical hybrid legumes. Plant Syst Evol 213:11–34
Hedrick PW (1983) Genetics of populations. Science Book International, Boston
Hewitt GM (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hoelzer GA, Wallman J, Melnick DJ (1998) The effects of social structure, geographical structure, and population size on the evolution of mitochondrial DNA. II. Molecular clocks and the lineage sorting period. J Mol Evol 47:21–31
Hsieh CF, Huang TC (1974) The Acanthaceous plants of Taiwan. Taiwania 19:19–57
Huang JC, Wang WK, Chiang TY (2001a) Population differentiation and phylogeography of Hygrophila pogonocalyx based on RAPDs fingerprint. Aquat Bot 70:269–280
Huang S, Chiang YC, Schaal BA, Chou CH, Chiang TY (2001b) Organelle DNA phylogeography of Cycas taitungensis, a relict species in Taiwan. Mol Ecol 10:2669–2681
Hudson RR (1990) Gene genealogies and the coalescent process. In: Futuyma DJ, Antonovices J (eds) Oxford survey of evolutionary biology. Oxford University Press, New York, pp 1–44
IUCN (International Union for Conservation of Nature and Natural Resources) (1997) Red list of threatened plants. IUCN, Gland, Switzerland
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munroled HN (ed) Mammalian protein metabolism. Academic, New York, pp 31–132
Kanno M, Yokoyama J, Suyama Y, Ohyama M, Itoh T, Suzuki M (2003) Geographical distribution of two haplotypes of chloroplast DNA in four oak species (Quercus) in Japan. J Plant Res 116:311–317
Kim HN, Harada K, Yamazaki T (1996) Isozyme polymorphism and genetic structure of a liverwort Conocephallum conicum. Genes Genet Syst 71:225–235
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kubo T, Satoh Y, Muro T, Kinoshita T, Mikami T (1995) Physical and gene organization of mitochondrial DNA from the fertile cytoplasm of sugarbeet (Beta vulgaris L). Curr Genet 28:235–241
Kumar S, Tamura K, Jakobsen IB, Nei M (2001) MEGA2: molecular evolutionary genetics analysis software. Bioinformatics 17:1244–1245
Li WH (1997) Molecular evolution. Sinauer, Sunderland
Liu TK, Chen YG, Chen WS, Jiang SH (2000) Rates of cooling and denudation of the Early Penglai Orogeny, Taiwan, as assessed by fission track constraints. Tectonophysics 320:69–82
Lonsdale DM, Brears T, Hodge TP, Melville SE, Rottmann WH (1988) The plant mitochondrial genome: homologous recombinations as a mechanism for generating heterogeneity. Phil Trans R Soc Lond B Biol Sci 319:49–163
Lu SY, Peng CI, Cheng YP, Hong KH, Chiang TY (2001) Chloroplast DNA phylogeography of Cunninghamia konishii (Cupressaceae), an endemic conifer of Taiwan. Genome 44:797–807
Manen JF, Natali A (1995) Comparison of the evolution of ribulose-1, 5-biphosphate carboxylase (rbcL) and atpB–rbcL noncoding spacer sequences in a recent plant group, the tribe Rubieae (Rubiaceae). J Mol Evol 41:920–927
Maskas SD, Cruzan MB (2000) Patterns of intraspecific diversification in the Piriqueta caroliniana complex in southeastern North America and Bahamas. Evolution 54:815–827
McCauley DE (1994) Contrasting the distribution of chloroplast DNA and allozyme polymorphisms among local populations of Silene alba: implications for studies of gene flow in plants. Proc Natl Acad Sci USA 91: 8127–8131
McCauley DE (1995) The use of chloroplast DNA polymorphism in studies of gene flow in plants. Trends Ecol Evol 10:198–202
Moore WS (1995) Inferring phylogenies from mtDNA variation: mitochondrial-gene tree versus nuclear-gene trees. Evolution 49:718–726
Moritz C (1994) Defining ‘evolutionarily significant units’ for conservation. Trends Ecol Evol 9:373–375
Moritz C (1995) Uses of molecular phylogenies for conservation. Phil Trans R Soc Lond B Biol Sci 349:113–118
Moritz C, Faith DP (1998) Comparative phylogeography and the identification of genetically divergent areas for conservation. Mol Ecol 7:419–429
Nei M, Tajima F (1983) Maximum likelihood estimation of the number of nucleotide substitutions from restriction sites data. Genetics 105:207–217
Noruis MJ (1994) SPSS for Windows, version 6.0. Prentice Hall, Englewood Cliffs
Ohsako T, Ohnishi O (2000) Intra- and interspecific phylogeny of wild Fagopyrum (Polygonaceae) species based on nucleotide sequences of noncoding regions in chloroplast DNA. Am J Bot 87:573–582
Orive ME, Asmussen MA (2000) The effects of pollen and seed migration on nuclear-dicytoplasmic systems. II. A new method for estimating plant gene flow from joint nuclear-cytoplasmic data. Genetics 155:833–854
Ouborg NJ, Piquot Y, Groenendael JMVan (1999) Population genetics, molecular markers and the study of dispersal of plants. J Ecol 87:551–568
Peng CI, Liu SL, Chiang TY (1999) Conservation of Ludwigia × taiwanensis (Onagraceae) in Taiwan. Endem Spec Res 1:73–78
Posada D, Crandall KA, Templeton AR (2000) GeoDis: a program for the cladistic nested analysis of the geographical distribution of genetic haplotypes. Mol Ecol 9:487–488
Provan J, Powell W, Hollingsworth PM (2001) Chloroplast microsatellites: new tools for studies in plant ecology and evolution. Trends Ecol Evol 16:142–147
Ray N, Currat M, Excoffier L (2003) Intra-deme molecular diversity in spatially expanding populations. Mol Biol Evol 20:76–86
Rozas J, Rozas R (1999) DnaSP version 3.0: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15:174–175
Sanderson MJ (2002) r8s: Analysis of rates (“r8s”) of evolution (and other stuff), version 1.5. Section of Evolution and Ecology, University of California, Davis. Distributed at http://ginger.ucdavis.edu/r8s/
Sarich VM, Wilson AC (1973) Generation time and genomic evolution in primates. Science 179:1144–1147
Slatkin M (1993) Isolation by distance in equilibrium and nonequilibrium populations. Evolution 47:264–279
Templeton AR (1998) Nested clade analyses of phylogeographic data: testing hypotheses about gene flow and population history. Mol Ecol 7:381–397
Templeton AR, Routman E, Phillips CA (1995) Separating population structure from population history: a cladistic analysis of the geographical distribution of mitochondrial DNA haplotypes in the tiger salamander, Ambystoma tigrinum. Genetics 140:767–782
Teng LS (1990) Geotectonic evolution of late Cenozoic arc-continent collision in Taiwan. Tectonophysics 183:57–76
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Wang WK, Huang JC, Chiang TY (2000) Ecology and conservation of Hygrophila pogonocalyx, endemic species of Taiwan. Quart Nat Conserv 31:54–58
Whitlock MC, MaCauley D (1999) Indirect measures of gene flow and migration, Fst ≠ 1/(4 Nm + 1). Heredity 82:117–125
Wu CI, Li WH (1985) Evidence for higher rates of nucleotide substitution in rodents than in man. Proc Natl Acad Sci USA 82:1741–1745
Wu JC, Chiang TY, Shiue WK, Wang SY, Sheen IJ (1999) Recombination of hepatitis D virus RNA sequences and its implications. Mol Biol Evol 16:1622–1632
Zuckerkandl E, Pauling L (1965) Evolutionary divergence and convergence in proteins. In: Bryson V, Vogel HJ (eds) Evolving genes and proteins. Academic, New York, pp 97–166
Acknowledgments
Financial support for this project was provided by the National Science Council and the Council of Agriculture, Taiwan. The authors thank Mei-Jane Fang of Academia Sinica for the technical assistance with nucleotide sequencing. We are indebted to two anonymous reviewers for their critical comments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Huang JC and Wang WK equally contributed to this work.
Rights and permissions
About this article
Cite this article
Huang, JC., Wang, WK., Peng, CI. et al. Phylogeography and conservation genetics of Hygrophila pogonocalyx (Acanthaceae) based on atpB–rbcL noncoding spacer cpDNA. J Plant Res 118, 1–11 (2005). https://doi.org/10.1007/s10265-004-0185-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-004-0185-z