Abstract
Methicillin-resistant Staphylococcus aureus (MRSA) is able to persist not only in hospitals (with a high level of antimicrobial agent use) but also in the community (with a low level of antimicrobial agent use). The former is called hospital-acquired MRSA (HA-MRSA) and the latter community-acquired MRSA (CA-MRSA). It is believed MRSA clones are generated from S. aureus through insertion of the staphylococcal cassette chromosome mec (SCCmec), and outbreaks occur as they spread. Several worldwide and regional clones have been identified, and their epidemiological, clinical, and genetic characteristics have been described. CA-MRSA is likely able to survive in the community because of suitable SCCmec types (type IV or V), a clone-specific colonization/infection nature, toxin profiles (including Pantone-Valentine leucocidin, PVL), and narrow drug resistance patterns. CA-MRSA infections are generally seen in healthy children or young athletes, with unexpected cases of diseases, and also in elderly inpatients, occasionally surprising clinicians used to HA-MRSA infections. CA-MRSA spreads within families and close-contact groups or even through public transport, demonstrating transmission cores. Re-infection (including multifocal infection) frequently occurs, if the cores are not sought out and properly eradicated. Recently, attention has been given to CA-MRSA (USA300), which originated in the US, and is growing as HA-MRSA and also as a worldwide clone. CA-MRSA infection in influenza season has increasingly been noted as well. MRSA is also found in farm and companion animals, and has occasionally transferred to humans. As such, the epidemiological, clinical, and genetic behavior of CA-MRSA, a growing threat, is focused on in this study.
Similar content being viewed by others
Introduction
Staphylococcus aureus colonizes the nasal mucosa of approximately 30% of individuals. It is associated with a wide range of infections including skin and soft tissue infections (SSTIs, such as bullous impetigo, abscesses, furunculosis, and staphylococcal scalded skin syndrome), systemic or fatal infections [such as blood stream infection (BSI), endocarditis, and pneumonia], and toxin-associated diseases (such as toxic shock syndrome and food poisoning) [1–4]. Historically, several classes of drug-resistant S. aureus have emerged [5–7]. They include penicillin G-resistant S. aureus, which produced penicillinase (PCase, penicillin-hydrolyzing enzyme) in the mid 1940s, and multidrug-resistant S. aureus (which was resistant to penicillin G, chloramphenicol, tetracycline, and erythromycin) in the late 1950s. Methicillin-resistant S. aureus (MRSA), isolated in 1961, continues to be a life-threatening multidrug-resistant bacterium, changing its resistance patterns by acquiring resistance to each new antimicrobial agent. Such MRSA has been a major cause of nosocomial infections, and thus is alternatively called hospital-acquired MRSA (HA-MRSA). In addition, since the period from 1997 to 1999, another class of MRSA has become a major concern worldwide as an emerging pathogen in the community [8–11]. This new class of MRSA is called community-acquired MRSA (CA-MRSA).
Ideally, MRSA in the community should impose no biological cost, i.e., it carries a certain type of staphylococcal cassette chromosome mec (SCCmec) and exhibits resistance to only limited numbers of antimicrobial agents. In turn, it can be selected by low levels of antimicrobial agent use in the community [12–14]. By contrast, MRSA in hospitals carries some other SCCmec types or exhibits multiple drug resistance, i.e., it imposes a high biological cost. Therefore, it has to be selected by high levels of antimicrobial agent use in hospitals to overcome the biological cost (in turn, transfer to the community is restricted) [12, 15, 16]. Moreover, CA-MRSA must have high colonization and infection abilities to distribute among healthy individuals in the community.
Current MRSA typing
MRSA is typed based on genetic characteristics such as multilocus sequence type (ST) [17], protein A gene (spa) type [18–20], agr type [21, 22], coagulase type [23, 24], and SCCmec type [25–28]. Generally, ST typing [and analysis of the clonal complex (CC) to which each ST type belongs] is a standard for genotyping (Fig. 1). Typing is based on the seven housekeeping gene sequences, with the allelic profile (allele no. representing the ST type) and CC obtained from the MLST website (http://www.mlst.net/) and eBURST software (http://eburst.mlst.net/), respectively. In contrast, spa typing targets a single gene (spa), and the spa type is given through a public spa type database (http://tools.egenomics.com/) and the Ridom Spa Server (http://spaserver.ridom.de/), but with different assignment (e.g., spa1 for the former and spat008 for the latter). Each ST type clone contains some spa variants. The agr type is determined by PCR; two methods are reported, demonstrating types I–IV. There are subtypes in agrI. Coagulase typing is conducted using an antiserum kit (Denka Seiken, Tokyo, Japan), determining types I–VIII. Attempts have been made for coagulase gene (coa) typing by PCR, but with incomplete results. Coagulase types well correlate with ST types. SCCmec typing is performed by multiplex PCR targeting the mec and ccr (recombinase gene) complexes or targeting even the junk regions, determining types I–VIII. There are subtypes in SCCmecII, III, and IV. Generally speaking, SCCmec I, II, and III are of HA-MRSA, while SCCmecIV and V are of CA-MRSA [9, 12, 15, 16, 29–32].
eBURST analysis of S. aureus. Data from the MLST website (http://www.mlst.net/) and eBURST software (http://eburst.mlst.net/), 23 December 2009. Major CC and ST types are indicated
For molecular epidemiology of MRSA epidemics due to certain clones, SmaI-digestion of MRSA DNA is used in pulsed-field gel electrophoresis (PFGE), aiming at PFGE typing (i.e., identification of epidemic strains and analysis of their divergence) [33, 34]. Indeed, in the US, MRSA clones have been classified according to SmaI PFGE types, such as USA100 (for ST5 HA-MRSA), USA300 (ST8 CA-MRSA), USA400 (ST1 CA-MRSA), and USA1000 (ST59 CA-MRSA) [33, 34]. Moreover, when an epidemic MRSA clone possesses a unique virulence gene or a combination of unique genes and unique DNA sequences, it can serve as a target for rapid and less-laborious screening of epidemic clones in PCR, multiplex PCR [35–38], or real-time PCR assay.
HA-MRSA
HA-MRSA is typically defined as MRSA isolated from inpatients that had been MRSA-negative at the beginning of hospitalization or MRSA isolated from inpatients 48 h or more after hospitalization [9, 39–41]. There are several types in HA-MRSA [29, 30, 42]. It is believed that MRSA is generated from S. aureus (methicillin-susceptible S. aureus, MSSA) by acquisition of SCCmec and that this has occurred only a limited number of times. Yet such limited numbers of MRSA clones, emerged, have disseminated worldwide, resulting in the current epidemics in hospital settings [42, 43]. Invasive MRSA epidemics were especially noted in the US and Japan in the late 1970s to early 1990s [5–7], when some second-generation cephems (such as cefoxitin and latamoxef) were widely used. These pandemics were followed by MRSA episodes in Europe. Some are still continuing.
Well-known MRSA clones [11, 42, 44–48] include the archaic clone (ST250/SCCmecI; reported in the UK from the 1960s), Iberian clone (ST247/SCCmecIA; reported in Spain from 1989), New York/Japan clone (ST5/SCCmecII; reported in the US from 1998), EMRSA-16 clone (ST36/SCCmecII; reported in the UK from 1993), Brazilian clone (ST239/SCCmecIIIA; reported in Brazil from 1992), Hungarian clone (ST239/SCCmecIII; reported in Hungary from 1993), EMRSA-15 clone (ST22/SCCmecIV; reported in the UK from 1993), pediatric clone (ST5/SCCmecIV or IVa; reported in Portugal from 1992), and Berlin clone (ST45/SCCmecIVa; reported in Germany from 1998) (Fig. 2) [11, 42, 44–57]. The Russian clone (ST239/SCCmecIIIR) can also be noted [57]. They are all Pantone-Valentine leucocidin (PVL)-negative. Among them, six clones (Iberian, Brazilian, Hungarian, EMRSA-15, New York/Japan, and pediatric) have disseminated more internationally, and are referred to as pandemic clones [42].
Worldwide distribution of major HA-MRSA clones. Data from Ref. [11, 42, 44–57]. The colored areas indicate the areas of each spreading clone. Yellow represents the region where the archaic clone was isolated. Orange represents the region where the Iberian clone was isolated. Pink represents the region where the New York/Japan clone was isolated. Lime green represents the region where the EMRSA-16 clone was isolated. Blue represents the region where the ST239 clone, including the Hungarian clone, Brazilian clone, and Russian clone, was isolated. Purple represents the region where the Berlin clone was isolated. Green represents the region where EMRSA-15 was isolated. Gray represents the region where the pediatric clone was isolated. Graphs indicate the rate of MRSA in all S. aureus isolates from the hospitals in each country
Due to the replacement nature of MRSA clones, distribution patterns in each country or area may change. In Hungary, southern Germany (ST228-I) and New York/Japan (ST5) clones have appeared recently, instead of the Hungarian clone (ST239) [47]. In Hong Kong, three major clones, the Hungarian clone (ST239), New York/Japan clone (ST5), and ST45 clone, have appeared recently [58]. The TW clone, a variant of ST239 MRSA, was recently noted as the cause of an outbreak in an intensive care unit (ICU) in London [59, 60]. The association with bacteremia of TW was four times higher than that of other common MRSA clones (such as EMRSA-15 or EMRSA-16). Therefore, it was described as highly transmissible MRSA [59, 60]. ST239 HA-MRSA is a growing MRSA group, which includes at least nine members (EMRSA-1, EMRSA-4, EMRSA-7, EMRSA-11, and the Brazilian, Portuguese, Hungarian, Viennese, and Russian clones), and is widely spreading in Asia, South America, and Eastern Europe [44, 61–66].
MRSA incidence among S. aureus varies country by country, with a low frequency (around 1%) in some countries in Europe (e.g., the Netherlands and Sweden) and a high frequency (>60%) in the US and Japan (Fig. 2) [51, 55, 67]. In the US, invasive MRSA infections have become a serious problem, with an estimated patient number of 94,360 and 18,650 deaths in 2005. MRSA infections generally occur most frequently among inpatients who undergo invasive medical procedures, who have weakened immune systems (or are immunocompromised hosts) or whose age is 50 or 65 years and older [67–70].
Criteria and risk factors for CA-MRSA
Epidemiologically, CA-MRSA is defined as MRSA isolated from outpatients with no history of hospitalization within the past 1 year, and who presented no other established risk factors for MRSA infection, such as surgery, residence in a long-term care facility, dialysis, or indwelling percutaneous medical devices or catheters [9, 39–41].
Since some MRSA clones have been well established as CA-MRSA, and CA-MRSA has spread into hospital settings, CA-MRSA may also be defined bacteriologically [9, 32–34, 71, 72]. This is probably the case when it is genetically identified (based on, e.g., ST type, spa type, and SCCmec type), when it is identified by PFGE typing, when it produces PVL, when it possesses SCCmecIV, or when it is resistant only to β-lactam antimicrobial agents (in some regions or countries). Low MIC values for oxacillin (MICs, around 32 μg/ml or less) or imipenem (MICs, around 1 μg/ml or less) may also serve as a marker of CA-MRSA; HA-MRSA exhibits higher MIC values for oxacillin (MICs, ≥128 μg/ml) or imipenem (MICs, 32 μg/ml) in many cases [73]. Such a bacteriological definition would be needed to investigate CA-MRSA spread within hospital settings.
CA-MRSA infection occurs in healthy individuals, especially children and adolescents, through skin-to-skin contact. Reflecting this, risk factors for CA-MRSA infection includes (e.g.) university and school students, adolescent athletes, military trainees, jail inmates, men who have sex with men (MSM), members of an infected family, hurricane evacuees, and people with tattoos [10, 33, 74–76].
Clinical aspects of CA-MRSA infections
Clinical aspects of CA-MRSA infection are summarized in Fig. 3 [4, 40, 75, 77–95]. CA-MRSA has emerged as a cause of serious infections. The vast majority (70–80%) of infections with CA-MRSA manifest as SSTI including pyogenic skin infection [40]. The incidence of SSTI caused by CA-MRSA has been increasing (e.g., 24 cases per 100,000 persons in 2000 to 164.2 cases per 100,000 persons in 2005 in the US) [96]. Most people who have developed CA-MRSA skin infection have been healthy individuals (such as children, prisoners, and athletes) without risk factors such as hospitalization [97].
Some CA-MRSA strains produce PVL, which lyses white blood cells. PVL-positive CA-MRSA has been especially isolated from “deep” skin infection and SSTI, such as furuncle, carbuncle, and cellulitis [4]. These patients complain of a severe pain due to unusually large abscesses. On the other hand, impetigo, the most common SSTI disease, which occurs in superficial skin, is related to PVL-negative CA-MRSA. A study of nasal MRSA carriers (53% positive for USA300) among American soldiers has shown that 29% of nasal MRSA carriers developed SSTI 1 year later. Of the nasal MRSA carriers who developed SSTI, 97% were positive for USA300, indicating that more than 50% of nasal USA300 carriers developed SSTI [98]. This strongly suggests that nasal carriage of MRSA is a risk factor for SSTI; however, the significance of nasal carriage may be varied among the clones. Especially, nasal USA300 carriers may be the high-risk group for SSTI.
CA-MRSA can also cause wound infections. Bacterial chronic wounds (diabetic ulcer, venous stasis ulcer) have increased over the past years, and the majority of these are due to CA-MRSA [80]. CA-MRSA has also been detected from surgical site infection, urinary tract infections (UTI) [82], infections of the eye and orbit [83], meningitis [85], and sinusitis.
Severe necrotizing fasciitis cases caused by CA-MRSA have been described as well. Necrotizing fasciitis is a rapidly progressive infection of adjacent subcutaneous tissue and superficial fascia. A 36-year-old Hispanic male with human immunodeficiency virus (HIV) infection suffered necrotizing fasciitis caused by CA-MRSA. In that case, the strain had the USA300 genotype, which is the most common CA-MRSA clone in the US and now spreading worldwide [99]. In another report, five cases of necrotizing fasciitis caused by USA300 were also described [87].
Nasal carriage and SSTI caused by CA-MRSA are also the risk factors for bloodstream infections (BSIs). USA300-associated BSIs have been described [100, 101]. Nasal carriage of S. aureus associates with higher incidence of BSIs, compared to non-carriers. Previous studies have reported that in approximately 80% of the patients who were initially positive for nasal S. aureus at admission and then developed BSIs, the PFGE genotypes of isolates from blood cultures were identical to those of the isolates from nasal swabs taken at admission, indicating endogenous infection [79, 81].
Management of SSTI caused by CA-MRSA includes incision and drainage of infected tissue and appropriate antimicrobial therapy. Most people with CA-MRSA primary infections can be cured with appropriate treatment. On the contrary, some patients with CA-MRSA primary infection will unfortunately develop severe life-threatening infections, so-called CA-MRSA-related secondary infection, probably through BSIs.
As has been reported, the mortality rates of community-acquired and hospital-acquired BSIs were 10 and 15%, respectively [78]. Malani et al. [102] have reported that advanced age, chronic renal insufficiency, catheter-related infection, Charlson weighted index of comorbidity score, and presence of MRSA were associated with in-hospital and 6-month mortality of S. aureus BSI. A case of BSI with CA-MRSA in four drug users was described in the UK [103].
Secondary infection with CA-MRSA includes pelvic abscess [75], lung abscess [84], osteomyelitis [86], arthritis [88], brain abscess [89], myositis [90], and endocarditis [91]. The prevalence of S. aureus secondary infection is very rare, but severe. Severe systemic infection due to CA-MRSA represents as an initial cutaneous abscess progressing to bacteremia, osteomyelitis, and simultaneous iliopsoas and piriformis abscesses [75]. Lung abscesses due to CA-MRSA are also commonly associated with SSTI [104]. Septic arthritis caused by USA300 isolates has been described. The arthritis was associated with longer duration of fever [104]. Although rare, CA-MRSA causes purpura fulminans and Waterhouse-Friderichsen syndrome, which are associated with a very high mortality [93].
CA-MRSA infections of the respiratory tract are less common in comparison with HA-MRSA, but can be severe. Typical symptoms of CA-MRSA-associated community-acquired pneumonia (MRSA-CAP) include high temperature, arterial hypotension, hemoptysis, rapid aggravation to the point of septic shock, and the need for ventilatory support [105]. In the US, two pediatric fatal cases (ages of 13 years and 12 months) due to necrotizing pneumonia of CA-MRSA (USA400) were reported in 1997 and 1998, respectively [8]. Fatal MRSA-CAP caused by PVL-positive ST30 CA-MRSA in a 1-year-old boy has been reported in Japan [92]. Most importantly, co-infection of influenza patients with CA-MRSA is associated with severe pneumonia with high fatality. In the US during the 2003–2004 influenza season, USA300- and USA400- caused pneumonia among patients with seasonal influenza or influenza-like disease (mean age 21 years, range 3 months–62 years) showed approximately 30% fatality [94]. In Consensus Guidelines, the American Thoracic Society and Infectious Diseases Society of America emphasized that CA-MRSA pneumonia, as well as SARS and novel influenza, should be paid attention to as life-threatening CAP [95]. In 2009, outbreak of the novel swine origin influenza A (H1N1) occurred worldwide. Fatal co-infection of a 42-year-old sailor with swine origin influenza virus A (H1N1) and PVL-positive ST30 CA-MRSA has already been reported in Hong Kong [106]. Therefore, for all age groups, pneumonia due to co-infection with influenza virus (including novel swine origin influenza virus A) and CA-MRSA should be given serious attention.
Molecular characteristics and epidemiology of major PVL-positive CA-MRSA
Well-known and well-characterized CA-MRSA clones include CA-MRSA belonging to ST1 and ST8 (USA400 and USA300, respectively), which are mostly found in the US [33, 34] and Canada [107, 108]. The USA300 clone associates with SSTI, and also invasive infections such as sepsis, pneumonia, and necrotizing fasciitis. It has the ability to replace preexisting MRSA clones, and now is the predominant CA-MRSA clone in the US [33, 34]. It is also expected to become a major clone even in hospital settings.
As for the evolution of USA300 (type strain USA300-0114), based on MLST data, seven cell surface protein genes, and virulence genes including (e.g.) enterotoxins, PVL, and the arginine catabolic mobile element (ACME) [34, 109], it has been proposed that HA-MRSA USA500 (ST8/SCCmecIV; negative for enterotoxins, PVL, and ACME) initially emerged from ST8 MSSA through the acquisition of SCCmecIV; subsequently, USA300 emerged from USA500 through mutation in two cell surface protein genes, and subsequent acquisition of staphylococcal pathogenicity island 5 (SaPI5, carrying sek and seq), PVL prophage, and ACME.
Drug resistance of USA300 (USA300-0114) has been diverging. Analysis of early 2000s USA300 strains revealed that 90% of the strains carried a 3.1-kb cryptic plasmid and 30-kb plasmid carrying a macrolide transporter gene (msrA) and blaZ gene, resulting in macrolide resistance, as well as β-lactam resistance (“core resistance”) [33]. Some of those strains further showed additional resistance to i) tetracycline through acquisition of a 4.3-kb plasmid with a tetK gene; ii) clindamycin through acquisition of a 2.6-kb plasmid with an ermC gene; or iii) levofloxacin [33] through gyrA mutation. Genomic analysis of an early 2000s strain (TCH1516) also revealed another β-lactamase plasmid (accession no. CP000731; 27 kb in size) with an aminoglycoside-resistant gene (aphA-3) and cadmium-resistant gene (cadD), in addition to msrA and blaZ [110].
Since the mid 2000s, clindamycin resistance and mupirocin resistance have increasingly been noted [111, 112]. Particularly, mupirocin resistance has been highly associated with multiple drug-resistant strains that frequently show resistance to fluoroquinolone, tetracycline, macrolide, lincosamide, and mupirocin. Genomic analysis of a multiple drug-resistant strain of USA300 [USA300-0114 (strain FPR3757), accession no. NC_007793] revealed carriage of a 37-kb plasmid with an ermC and mupirocin resistant gene (mupA), as well as gyrA mutation and a 4.4-kb tetK plasmid, but not a β-lactamase plasmid [72].
More recently, vancomycin-intermediate strains of USA300 (designated as VISA) have been noted [113, 114]. Although mutations associated with vancomycin-intermediate phenotypes have been proposed [115], the exact responsible genes for the vancomycin-intermediate phenotypes of USA300 are still unknown.
In the US, there has been recent clonal expansion of a subset of USA300 isolates, originating from the common ancestor [116]. Those USA300 isolates are identified primarily based on i) PFGE patterns, but also based on other genetic markers such as ii) ST type (ST8), iii) spa types [major type: spa1 (t008)], iv) PVL genes, v) ACME, vi) SCCmecIVa, vii) the 40-bp direct repeat unit (dru) region, which is located adjacent to IS431 within SCCmecIVa, and viii) a signature AT repeat sequence within the hypothetical SACOL0058 gene, which is located 1.4-kb downstream of the 3′ end of SCCmecIVa [33, 34, 37, 72, 117]. A subset of USA300 isolates with modest genetic changes, however, may differ in their virulence potentials. Major types in the US are USA300-0114 (spa1) and USA300-0247 (spa1), and minor types include (e.g.) USA300-0114 (spa168). In addition, USA300-0114 (spa168) exhibited low virulence in a mouse sepsis/bacteremia model [116].
The USA300 clone has already spread across the Atlantic and Pacific oceans [108, 111, 112, 116, 118–130]. It has been isolated in, e.g., Canada (North America), the UK, Germany, Denmark, Switzerland, Austria, and Italy (Europe), Colombia (South America), Australia (Oceania), and Japan, Korea, and Taiwan (Asia), as well as the US. Among USA300 clones, clones with spa1 (t008) have been widely identified in North America and Europe (Table 1). However, hospital-associated USA300 with spa363 (t024), as well as a community-associated clone with spa1 (t008), has been isolated in Denmark. Other variants with spa163 and spat063 have also been isolated in the US and Australia, respectively. In Japan, USA300 clones with spa363 (t024) and spa985 (t711) have been isolated.
In the US, multidrug-resistant USA300 clones, which were resistant to levofloxacin, tetracycline, erythromycin, clindamycin, and mupirocin, have been isolated [72, 111, 112]. On the other hand, USA300 clones isolated in the other countries have shown resistance to limited drugs (Table 1).
ST80 CA-MRSA is mostly found in Europe [9], though also in North Africa (Algeria) [131] and the Middle East [132]. Fusidic acid resistance, mediated by fusA, fusB, or fusC genes, is one of the genetic and phenotypic characteristics [9, 133, 134]. Currently, high-level mupirocin-resistant strains, whose MIC levels were >1,024 μg/ml, emerged in Kuwait, suggesting the difficulty of eradication of ST80 CA-MRSA [135]. This was mediated by different sizes (21 and 26 kb) of plasmids with mupA (high-level mupirocin resistance gene).
ST59 CA-MRSA is spreading in Taiwan [32] and has also been isolated from Hong Kong [136], the US (USA1000) [33] and Japan. ST59 CA-MRSA from Taiwan carries a unique SCCmec structure (initially designated as SCCmecVII and currently reclassified as SCCmecV) that involves two distinct ccrC genes [32, 137]. In contrast to the initial common understanding that CA-MRSA is generally resistant to only β-lactam agents, it was multiple drug-resistant to tetracycline, kanamycin, streptomycin, erythromycin, clindamycin, and chloramphenicol. ST59 CA-MRSA carries a PCase plasmid with a tetracycline resistance gene (tetK), composite transposon-bearing streptomycin resistance gene (aadE), kanamycin resistance gene [aph(3′)-III], and erythromycin/clindamycin resistance gene (ermB) [32]. In contrast to many other cases [in which the chloramphenicol resistance gene (cat) is carried by a plasmid], the cat gene in the Taiwanese clone is located in the chromosome. PVL-negative ST59/SCCmecIVx strains with the same multiple drug resistance pattern are present in Taiwanese hospital settings [32]. Recently, a very similar PVL-positive ST59/SCCmecVII strain was found in Japan [138].
PVL-positive ST30 CA-MRSA [9, 33, 89, 139–141] is a global clone, which is spreading worldwide including Japan. Nosocomial outbreak-derived MRSA in Japan in the 1980s was largely PVL-positive ST30 MRSA with spa43 and SCCmecIVa, IVc, or IVx [142], while current PVL-positive ST30 strains exhibit spa19 and SCCmecIVa or IVc. Some PVL-positive ST30 strains acquired a plasmid (pGKT1) conferring resistance to gentamicin, kanamycin, and tetracycline [141].
In the 1950s and 1960s, PVL-positive and -negative penicillin-resistant S. aureus (ST30/spa43) spread in hospitals and the community worldwide (e.g., Australia, Denmark, the UK, US, and Cuba) [143]. One of the possibilities is that PVL-positive ST30 CA-MRSA evolved from this worldwide ST30 MSSA. PVL-positive ST30/spa19/SCCmecIVc with the same PFGE pattern was isolated from Far Eastern Russia, while PVL-positive ST30/SCCmecV with the same PFGE pattern from Egypt exhibited spa251. PVL-positive CA-MRSA clones also include other ST types such as emerging ST22, spreading to Europe and Japan, [144, 145] and ST93 [146].
CA-MRSA factors associated with colonization and spread in the community
Some major CA-MRSA clones have unique colonization and infection ability, which may play a role in community transmission. The USA300 clone (ST8) possesses the ACME-SCCmecIV linkage as a merit of colonization and spread [72]. ACME-SCCmecIV linkage, which was first found in USA300, is considered to play a key role in the persistence of USA300 in the community. ACME (type I) in USA300 appears to originate in S. epidermidis, a representative skin-colonizing bacterium [12]. ACME contains two operons associated with pathogenesis, arc (encoding for the arginine deaminase pathway), and opp3 (encoding for the oligopeptide permease system) [12, 72]. Arginine deaminase is a virulence factor of Streptococcus pyogenes, and plays a role in (e.g.) survival at low pH, inhibition of human peripheral blood mononuclear cell proliferation, or invasion [147, 148]. Oligopeptide permease is associated with (e.g.) MRSA growth and infection [149]. ACME is also found in certain strains of ST5 (USA100) and ST59 (USA1000), but such ACME lacks one possible virulence component (the opp3 operon) found in USA300 [12].
The presence of SCCmecIV is an advantage for CA-MRSA, because low levels of antimicrobial agents use in the community could be sufficient to drive the spread of MRSA with SCCmecIV (imposing no biological cost) [12–14]. In contrast, SCCmecI, SCCmecII, or SCCmecIII (all of which impose biological cost) would need high levels of antimicrobial agent use (in hospitals) to overcome the biological cost, and may restrict such HA-MRSA from spreading into the community [12, 15, 16].
The PVL-positive ST30 clone carries the collagen adhesin gene (cna) and bone sialoprotein adhesin gene (bbp) [140], and the combination of PVL, cna and bbp would play a role in colonization to matrix within the body. The presence of the PVL gene and the cna gene correlates with pulmonary manifestations, probably due to viral infection-caused epithelial damage permitting binding of PVL-positive S. aureus to exposed collagens and laminin [150, 151]. In addition, presence of the bbp gene allows S. aureus adherence to bone sialoprotein, leading to hematogenously spread osteomyelitis, arthritis, and iliopsoas abscess [75, 152–154].
The PVL-positive ST22 clone exhibits heavy biofilm formation (Fig. 4), which results in heavy colonization among family members and their relatives [of the 12 members in 4 families, 9 (75%) were colonized with the same MRSA, Fig. 4] [144]. In these families, cutaneous abscess occurred every 2 months.
Scanning electron micrograph showing ST22 MRSA and its biofilm (a) and family pedigree with ST22 MRSA infection (b). Data are from Ref. [144]. a Strains were grown on trypticase soy agarII with 5% sheep blood at 37°C overnight. Bar is 1 μm. b Square and circle represent male and female, respectively. Closed mark represents the person who was colonized on infected with MRSA
CA-MRSA is also spreading among healthy (asymptomatic) people in the community (including families or public transport). CA-MRSA colonizes the nares or cheek in many cases, but also axilla, umbilicus, and groin (Fig. 5) [144]. Although methicillin-resistant strains (MR-CNS) of coagulase-negative staphylococci, a major pathogen of nosocomial BSI, colonize even the feet [155], no MRSA is isolated from the feet [144]. Cheeks and the nasal cavity can be highly colonized by CA-MRSA in comparison with other parts of possible colonization. So it is highly possible that CA-MRSA can be transmitted through close body contact (hugs, kisses, etc).
Virulence factors
Adhesins common to MRSA
Adhesins involve not only microbial surface components recognizing adhesive matrix molecules (MSCRAMMs) [156, 157], which are spike-like protein structures anchored to the cell wall, but also biofilms [158], and signal peptides of PVL [159]. MSCRAMMs commonly found in MRSA are (e.g.) adhesins for laminin, fibronectin, elastin, fibrinogen, and protein A (Spa). Some adhesins play a role not only in cell adherence but also in cell invasion [160].
Immuno-modification factors at the initial stage of infection
Spa (protein A) commonly presents on the cell surface of most S. aureus clinical strains, and, in cases, is released in culture supernatants [161–163]. The N-terminus of Spa neutralizes immunoglobulins through direct binding to Fc and Fab fragments, indicating immunoresistant protein (which prevents efficient elimination of bacteria by immune cells). The C-terminus is the transmembrane domain. Short sequence repeats (SSRs), consisting of eight amino acids, are located between the Fc binding domain and transmembrane domain and are used for spa typing [18–20]. Structures of SSRs could vary the virulence of Spa by changing the distance of the Fc binding domain from the cell surface. It is also known that Spa activates tumor necrosis factor receptor 1 signaling of human airway epithelial cells through the Fc binding domain, resulting in release of IL-8 [164]. PVL and Spa synergistically induce severe damage of lung tissue and high mortality in murine necrotizing pneumonia models using PVL-positive S. aureus [163].
Toxic shock syndrome toxin-1 (TSST-1) and staphylococcal enterotoxin B (SEB) could suppress the motility of polymorphonuclear neutrophils (PMNs) through the inhibition of MRSA exoproteins expression [165, 166]. SEG, SEI, SEM, SEN, and SEO [all located on the enterotoxin gene cluster (egc)] [167] modify polyclonal T cell proliferation and induce immune anergy by T cell suppressor activity, B cell depletion, and inhibition of antibody responses [167–170]. And also, they may reduce S. aureus pathogenicity, and may contribute to bacterial colonization [171, 172].
PVL targets PMNs and induces cell death, resulting in bacterial evasion from bactericidal functions of PMNs [173]. ACME inhibits human peripheral blood mononuclear cell proliferation, and also inhibits nitric oxide production of the immune cells, which is important in both innate and adaptive immune responses [72]. Fibronectin-binding proteins play a role in adherence to extracellular matrix and invasion of host cells including epithelial cells and fibroblasts [174]. Therefore, possession of those factors could be an advantage in colonization and/or invasion of MRSA.
Cytolysins
Hemolysins include α-hemolysin (α-toxin; gene, hla), β-hemolysin (β-toxin; gene, hlb), γ-hemolysin (γ-toxin; genes, hlgA, hlgB, hlgC), and δ-hemolysin (δ-toxin; gene, hld) [174, 175]. Of those, at least α-, γ-, and δ- hemolysin exhibit cytolytic activity even for non-erythrocytes. α-Hemolysin is a strong cytolytic toxin in murine models. γ-Hemolysin consists of three component proteins, HlgA, HlgB, and HlgC. Combination of HlgA and HlgB acts as hemolysin, and combination of HlgB and HlgC acts as a leucocidin [173]. δ-Hemolysin is a phenol-soluble modulin that may be classified as a cytolytic peptide [176]. The δ-hemolysin gene is located within the RNAIII region in the agr locus.
Leucocidins are a family consisting of LukED, PVL, γ-hemolysin (HlgB and HlgC), and LukM [173, 175, 177]. The first three leucocidins are virulent for humans [173], while LukM is virulent for bovines (not for humans) [177]. PVL is unique to CA-MRSA. The PVL and LukM genes are carried by a phage.
Cytolytic peptides
Cytolytic peptides are 22-25 amino acids or 40 amino acids long [178]. S. aureus (including MRSA) harbors three distinct cytolytic peptide genes on three different loci on the chromosome. The three genes are the δ-hemolysin gene (as described above), the PSMα gene (4 tandemly-repeated genes, PSMα1 to PSMα4), and the PSMβ gene (2 tandemly-repeated genes, PSMβ1 and PSMβ2) [178]. They are expressed to a higher extent in CA-MRSA [such as USA300 (ST8), USA400 (ST1), ST30, and ST59], than in HA-MRSA [such as New York/Japan (ST5)] [178, 179]. In addition, HA-MRSA carries additional cytolytic peptide genes on SCCmecII and SCCmecIII [180]. The PSM-mec gene is not present in SCCmecIV, which is common in CA-MRSA.
Superantigens
Superantigens bind to major histocompatibility complex class II (MHC II) molecules and the variable part of the β-chain of certain T-cell receptors (Vβ-TCRs), resulting in unusual activation of T-cells [181–185]. Staphylococcal superantigens are also called SE, and include TSST-1 (previously SEF), SEA, SEB, SEC1, SEC2, SEC3, SED, SEE, SEG, SEH, SEI, SEJ, SEK, SEL, SEK, SEL, SEM, SEN, SEO, SEP, SEQ, SER, SES, SET, SEU, and SEV [185, 186]. SEA (and a combination of multiple superantigens) is associated with severe diseases [171, 187]. Moreover, TSST-1 and SEB may suppress the motility of PMNs through the inhibition of exoprotein expression, and allow MRSA to invade and damage tissues [165, 166].
Superantigen genes are located on a pathogenicity island [165, 188], such as SaPI5 carrying seq2 and sek2, ΦSa3mw carrying sek, sea, sek2, and seq, SaPI3 carrying seb, sek, seq, or seb, sel, and sek, SaPI1 carrying tst, and sek, and SaPIm1/n1 carrying tst, sec, and sel. At least SaPI1 and likely SaPI3 are mobile pathogenicity islands, and can be transferred from bacterial cell to cell by the action of a phage in such a way that SaPI is excised from the chromosome, and packed into phage particles to be transferred to other cells [189].
Exfoliative toxins
Exfoliative toxins are proteinase from S. aureus, which digest desmoglein 1, resulting in exfoliation of the epidermis cells [190]. Exfoliative toxin involves ETA, ETB, and ETD. Of those, ETA and ETB are responsible for staphylococcal scalded skin syndrome (SSSS) in infants and bullous impetigo in children, while ETD is responsible for not only bullous impetigo but also deep pyoderma in adults [191]. ETA is expressed in ST88 CA-MRSA, and ETB in ST89 or ST91 CA-MRSA. ST89 CA-MRSA is rarely positive for both ETA and ETB [141]. ETD is found in ST80 CA-MRSA [35, 71]. The ETA gene is carried by a phage [192], while the ETB gene is located on a plasmid [193]. The ETD gene is located on a pathogenicity island on the chromosome [194].
Structure, activity, and clinical importance of PVL
PVL is considered as an immunoresistant factor due to its cytotoxity against human PMNs and monocytes [195]. Possible association of PVL with necrotizing lesion development has been described. PVL, a member of staphylococcal two-component pore-forming toxins, consists of PVL-S (284 amino acids, 32 kDa) and PVL-F (301 amino acids, 34 kDa) (Fig. 6a) [196–199], which can be purified from bacterial culture supernatants as a monomeric form [173]. Crystal structures of PVL-S and PVL-F water-soluble monomers were reported in 2004 [199] and 1999 [196], respectively. Sequences of those crystal structures were identical to those of the PVL gene core sequence (in ST30 CA-MRSA and others) in Fig. 7 [32, 200, 201]. PVL-S monomer, but not PVL-F monomer, can bind to PMN membrane with high affinity in vitro, indicating that PVL-S mediates receptor binding [202]. PVL-S and PVL-F assemble into the channel complex on cell membrane and induce necrosis of PMNs and monocytes within 30 min (Fig. 6b). Based on studies of PVL and its homologues [196, 203, 204], deduced channel complexes are hexamer-octamer. Amino acid substitutions have been identified by phyrogenetic analysis of the PVL gene (Fig. 7). Some of those are located at the PVL-S surface contacting adjacent PVL-F or membrane and membrane spanning regions of PVL-S (Fig. 6a), and may alter receptor binding of PVL-S or channel formation. Neither PVL-S nor PVL-F alone induces necrosis even at more than ten times higher concentrations (Fig. 6b). PVL also induces apoptosis of PMNs within 4 h (Fig. 6c) through mitochondrial pathway (cytochrome c/caspase-9/3 cascade) [195].
Structures and functions of PVL. a Data from Refs. [173, 196–204]. Protein Data Base (PDB) codes of PVL-F monomer (PDB code, 1PVL) [196] and PVL-S monomer (PDB code, 1T5R) [197] were downloaded from the RCSB Protein Data Bank (http://www.pdb.org/) [198], and their crystal structures were reconstructed using SWISS-MODEL and the Swiss-Pdb Viewer (http://www.expasy.org/spdbv/) [199]. b Schematic representation of primary structures of recombinant histidine (His)-tagged PVL-S (H-PVL-S) and PVL-F (PVL-F-H) and their cell lysis activities are shown. Necrosis of human PMNs and monocytes was determined by morphological change and trypan blue staining. Percents of cell lysis (mean ± SD) are shown (black lines, human PMNs; blue line, human monocytes). In the boxes, light micrographs of PMNs (incubated with the indicated concentrations of PVL) are also shown. c PVL-induced apoptosis of human PMNs was detected by TUNEL staining (green) under a fluorescent microscope. Nuclei were also stained with propidium iodide (PI). d Neutralization of PVL activity with intravaneous immunoglobulin G (IVIG) was examined. PVL proteins were pre-incubated with the different concentrations of IVIG for 15 min at 37°C and then incubated with human PMNs
Schematic representation of the mutation sites in the PVL genes (a), phyrogenetic analysis of the PVL genes (b), and amino acid substitution sites in three-dimensional structures of PVL-S and PVL-F proteins (c). Data, in part, from Refs. [32, 200, 201]. a A schematic representation of the PVL gene core sequence of ST30 CA-MRSA (GenBank, AB186917) is shown. S and Mature represent the sequences encoding signal peptides and mature proteins, respectively. The sites of nucleotide substitutions and amino acid substitutions are indicated below the scheme. b A phylogenetic tree of the PVL gene is shown. Box colors correspond to those of amino acid substitutions in a. Clones indicated as the same color carry PVL genes giving the same amino acid sequence. The PVL gene core sequence [including MRSA ST30 (its single locus variant ST1335) and MSSA ST50] is indicated as a black-lined red box. c Positions of amino acid substitutions (shown in a) are indicated in three-dimensional structures of (PDB code, 1PVL) [196] and PVL-S (PDB code, 1T5R) [197]. PDB codes were downloaded from the RCSB Protein Data Bank (http://www.pdb.org/) [198], and crystal structures were reconstructed using SWISS-MODEL and the Swiss-Pdb Viewer (http://www.expasy.org/spdbv/) [199]. For the crystal structures, β-sandwich domain is yellow in PVL-S and light yellow in PVL-F, rim domain is blue in PVL-S and light blue in PVL-F, and pre-stem domain is green in PVL-S and dark green in PVL-F. Deduced mode of action of PVL was constructed according to the Refs. [32, 173, 196–204]. Amino acid substitutions in PVL-S may alter the spectrum of its receptor recognition
PVL-S has been stained in tissue cells in vivo. Despite its presence in patients’ infected tissue and in vitro cytolytic activity, the role of PVL in necrotizing lesion development is still controversial. Subcutaneous injection of rabbits [205] and nasal injection of murine [163] with purified PVL developed necrotizing lesions, respectively. Nasal injection of murine with PVL-positive S. aureus, but not its PVL-knocked down mutant, also induced necrotizing pneumonia with PMN infiltration, tissue necrosis, and hemorrhage [163]. A synergistic effect of PVL and Spa was proposed. Brown et al. have indicated the importance of PVL in CA-MRSA (USA300)-induced muscle lesions and necrotizing pneumonia in the murine model and protective effects of PVL immunization [206]. These data suggest the role of PVL in lesion development. In contrast, Voyich et al. [207] and Bubeck Wardenburg et al. [208, 209] provided counterevidence. They showed no significant difference in virulence (e.g., abscess volume, lesion size, mortality) between PVL-positive CA-MRSA and its PVL-knocked down mutants in the murine models (skin abscess, dermonecrosis, pneumonia, and bacteremia). Actually, necrotizing fasciitis caused by PVL-negative MRSA has also been reported clinically [210].
Currently, the role of PVL in bacterial survival at the early stage of infection has also been described elsewhere [211]. Other than cytotoxity, the possible role of the PVL-S signal peptide that attaches to the cell wall after cleavage in bacterial adhesion to collagens (components of tissue extracellular matrix) has also been described [159]. Although the exact role of PVL as a cytotoxin is still controversial, possession of PVL may be an advantage for CA-MRSA at the early stage of infection. Since patients with PVL-positive CA-MRSA infection complain of severe pain and much pus, it is also possible that PVL may induce higher cytotoxity in human than that in other animal species.
Subinhibitory concentrations of clindamycin, rifampicin, and linezolid inhibit PVL production, while β-lactam agents enhanced PVL production [212, 213]. Intravenous immunoglobulin G (IVIG), a product of Japan, neutralizes PVL activity against human PMNs at the levels of normal blood IgG (8.7–17 mg/ml) or less (Fig. 6d). This was the case even for human monocytes. Therefore, these agents may have therapeutic potential for infection with PVL-positive CA-MRSA.
PVL gene
The PVL region is a polycistronic operon, which contains the S component gene (luk PV S, 939 bp) upstream and the F component gene (luk PV F, 978 bp) downstream, with a single-base intercistronic region [214, 215].
Gene mutations and amino acid changes of the PVL gene that have been identified are summarized in Fig. 7a [32, 200, 201]. All gene mutations were substitutions: 11 substitutions in the S gene and 5 substitutions in the F gene. In the S gene, 6 of 11 were non-synonymous substitutions, which change amino acids; all of the amino acid changes occurred in the mature S component. In contrast, in the F gene, of five synonymous substitutions (which do not change amino acids), only one caused amino acid change in the mature F component. The facts that amino acid changes accumulate in the S component and the F component is highly conserved strongly suggest that the S component plays a role in receptor bindings in tissues and the F component plays a role as a toxic component. This result is consistent with a previous notion, obtained by purified component analysis, that the S component acts as a receptor binding, while the F component acts as a toxic component.
The PVL gene is found in various MSSA and MRSA clones. Phylogenetic analysis of the PVL gene is summarized in Fig. 7b [32, 200, 201]. The PVL gene has been identified in 12 ST types. Genes encoding the PVL component with different amino acid sequences are distinguished by different colors. Based on phylogenetic tree analysis, the core sequences could be those of the ST30/ST1335 (single locus variant of ST30)/ST50 group—ST30 includes CA-MRSA and MRSA in the 1980s and 1990s, and ST50 includes CA-MSSA associated with SSTIs; the ST30/ST1335/ST50 group could be an origin of various PVL gene sequences. The same PVL amino acid sequences as those of the ST30/ST1335/ST50 group can be found in ST1, ST22, ST25, ST59, ST80, ST121, and ST1010 (although the gene nucleotide sequences were divergent from those of the ST30/ST1337/ST50 group). However, it should be noted that there existed distinct PVL amino acid sequences in ST30, ST1, and ST8. Distinct PVL amino acid sequences are also found in five other minor cases (Fig. 7b). Based on the previous data of PVL and its homologues [32, 173], and amino acid changes in PVL crystal structures[196–204] and deduced mode of action of PVL are shown in Fig. 7C. In PVL-S, a change at His68 is located at the bottom surface of rim domain (blue, Fig. 7c), suggesting the effect on receptor binding. Of four amino acid changes located in β-sandwich domain (yellow, Fig. 7c), Val27, His148, and Asn233 are located on the surface contacting adjacent PVL-F. These three, as well as one more change at Phe129 located in pre-stem (membrane spanning) domain (green, Fig. 7c), may alter channel formation. In contrast to PVL-S, Val 11, only one amino acid change in PVL-F, is located at the loop-out region of β-sandwich domain (Fig. 7c). Therefore, PVL-F is functionally conserved, suggesting that PVL-F may play a critical role in PVL activity in addition to channel formation.
Genome and SCCmec
Genome
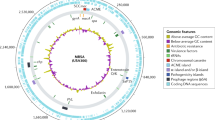
The entire genome structure of USA300 [USA300-0114 (strain FPR3757)] [72] is shown in Fig. 8a. The PVL genes are located in the prophage called ΦSA2USA. At the opposite position on genome USA300, SCCmec(IVa) is present. USA300 SCCmecIVa is linked to ACME. The connection of SCCmec to ACME is most probably a recombination process at invert repeat (IR) sequences (Fig. 8b). Moreover, SCCmecIVa and ACME could be excised by the ccr recombinase function at the repeat sequences, and the circular DNA generated could be transferred to other bacterial cells [12, 43].
Structures of the USA300 genome (a) and ACME-SCCmecIVa linkage structure (b). Data from Ref. [72]. a Genomic structures of the USA300 chromosome (accession no. NC 007793) and plasmids (accession nos. NC 007790, NC 007791, and NC 007792) are shown. Positions of the ACME-SCCmecIVa linkage, spa, coagulase gene (coa), seven housekeeping genes for MLST, drug resistance genes, and virulence genes including SaPI structures are also indicated. b The ACME-SCCmecIVa linkage structure (orfX–orfY region [137]) is shown. Positions of the mec complex, ccr complex (in SCCmecIVa), acr cluster, opp-3 cluster (in ACME), orfX, and orfY are indicated. Sequences at the 5′- and 3′- ends of SCCmecIVa and ACME are also shown below the scheme. The positions of direct and inverted repeats are indicated by green arrows
USA300 (USA300-0114) is resistant to fluoroquinolones (e.g., ciprofloxacin) macrolides (erythromycin), lincosamides (clindamycin), and tetracycline. Of those resistances, fluoroquinolone resistance is due to the gyraseA (gyrA) mutation on the chromosome, while the other resistance genes are on the plasmid (Fig. 8a).
SCCmec
SCCmec types were reported previously [25–27]; however, SCCmec typing has recently been reproposed by the SCCmec Working Committee (IWS-SCC) [28]. SCCmec types, including those reproposed by the committee (shown in parenthesis) [25–28, 32, 137], are shown in Fig. 9. Since SCCmec is inserted at the attachment site (att), which is located at the right end of orfX, all SCCmec structures contain the att sequence at both ends. SCCmec contains core structures consisting of a mec complex representing methicillin resistance and a ccr complex. The locations of primers used for previous typing are shown in Fig. 9 (arrows). On the other hand, retyping recommended by the committee is determined by the mec complex and ccr complex, located to the right side of the mec complex. In some cases, SCCmec type III is linked to SCCmercury (HG). SCCmec type IVa (of USA300) is linked to ACME. SCCmec types II and III carry the peptide toxin gene (PSM-mec) [180]. Besides methicillin resistance, SCCmecII confers kanamycin resistance and erythromycin/clindamycin resistance. SCCmercury/type III carries the erythromycin/clindamycin resistance gene, tetracycline resistance gene, and cadmium resistance gene. SCCmec types II and III are seen in pandemic HA-MRSA clones.
Basic structures of SCCmec. Data from Refs. [25–28, 32, 137]. Roman numerals in parentheses represent a new type of SCCmec proposed by the SCCmec Working Committee (IWG-SCC). Arrows indicate the region of detection by PCR method to decide the SCCmec type. ACME is located downstream of SCCmecIVa in USA300
SCCmecVII (currently SCCmec type V) contains the ccrC-carrying unit at the orfX end side and adjacent to IS431 [137]. This core unit is also seen in other SCCmec structures, suggesting that it is used to construct SCCmec mosaic structures through recombination (Fig. 10).
Rapid detection of PVL-positive CA-MRSA
Multiplex PCR is developed for the rapid detection of major PVL-positive CA-MRSA clones. The targets for an assay are a combination of the gene or DNA sequence unique to each clone. Targets include the PVL gene, the MW1409 and MW0756 loci, and the seh gene for ST1 (USA400) [35, 36]; the PVL gene and the arcA gene (on ACME) for ST8 (USA300) [35–37]; the PVL gene and the etd gene for ST80 [35]; and the PVL gene and the cna and bbp genes for ST30 [38]. The mecA gene and the nuc gene (encoding for S. aureus-specific thermostable nuclease) may also be included in an assay to differentiate MRSA from MSSA and S. aureus from CNS, respectively [36].
Drug resistance
Drug resistance genes and their resistance mechanisms in MRSA are summarized in Table 2. Some of them are found in CA-MRSA. Breakpoints for drug resistance have been defined by the CLSI.
β-lactam agents [32, 35, 71, 73, 216–219]
Ampicillin resistance is encoded for by the blaZ gene. Methicillin resistance is encoded by the mecA gene, whose product is methicillin-resistant penicillin binding protein 2 (peptidoglycan-associated transpeptidase, PBP2), known as PBP2a or PBP2′. The expression level of the mecA gene is under the control of chromosomal genes such as fem. Some MRSA exhibits a low oxacillin MIC level (MIC, <4 μg/ml), and thus are susceptible to oxacillin according to the CLSI’s breakpoints; such MRSA is called borderline MRSA. To improve controversial phenomena, and to diagnose borderline MRSA, cefoxitin is used to judge MRSA.
CA-MRSA exhibits low oxacillin and carbapenem (especially imipenem) MIC levels compared to HA-MRSA. For instance, oxacillin MIC90s are 32 μg/ml for CA-MRSA and ≥256 μg/ml for HA-MRSA, and imipenem MIC90s are 1 μg/ml for CA-MRSA and 64 μg/ml for HA-MRSA. This phenomenon is even more remarkably observed: imipenem MIC90s 0.12 μg/ml for PVL-positive CA-MRSA and 2 μg/ml for PVL-negative CA-MRSA. It is possible that HA-MRSA strains that have been selected in more cases over longer periods by β-lactam agents in hospitals have resulted in higher resistance. For example, in Japan, MRSA isolated from hospitals from 1976 to 1989 was susceptible, or only slightly resistant, to imipenem (MIC90, 12.5 μg/ml), while that isolated in 1990 and 1991 was highly resistant to imipenem (MIC90, ≥100 μg/ml). If this is the case, in the future CA-MRSA could also manifest high oxacillin and imipenem resistance, just as HA-MRSA has been selected.
As for distribution in the community in Japan, MRSA is isolated from children at a few percent, no MRSA is isolated from university students, and it is isolated in public transport (trains and subways) at 2.5%.
Vancomycin [220–223]
Vancomycin is a first-line antimicrobial agent for the treatment of MRSA infections. Vancomycin-intermediate S. aureus (VISA, MIC; 4-8 μg/ml) is characterized by cell wall thickness (although the responsible gene has not been identified yet). VISA is MRSA in many cases. Heterogeneous VISA (hVISA) has also been noted. Vancomycin-resistant S. aureus (VRSA, MIC; ≥16 μg/ml), which is always MRSA, carries the vanA gene (on the Tn1546-like structure in a plasmid). The vanA gene is transmissible. The vanA plasmid pLW1043 confers multiple-drug resistance, including ampicillin (blaZ), gentamicin (aacA/aphD), trimethoprim (dfrA/thyA), and quaternary ammonium compound resistance (qacC). VRSA has been isolated not only from the US, but also from India and Iran.
VRSA is highly associated with USA100 (New York/Japan clone) and USA800 (pediatric clone) in the US. VISA has also been found in the most remarkable CA-MRSA clone in the US, USA300.
Linezolid [224, 225]
Linezolid is an alternative to vancomycin for the treatment of MRSA infections. Linezolid is able to penetrate into tissues (such as lung tissues) better than vancomycin. One of the resistance mechanisms is G to U substitution at position 2576 in the peptidyl transferase center of 23S rRNA, exhibiting MIC levels ≥8 μg/ml. The G2576U mutation-mediated resistance levels are affected by the number of copies of mutated 23S rRNA genes on the chromosome (present in five copies per cell). The other mechanism is methylation of adenine at position 2503 in the peptidyl transferase center of 23S rRNA, exhibiting 8 μg/ml. The responsible methyltransferase is encoded by the cfr gene; cfr also confers on host bacterial cells multiple resistance to chloramphenicol, clindamycin, tiamulin, and quinupristin/dalfopristin, in addition to oxazolidinones.
Aminoglycosides [32, 35, 71, 142, 145, 219, 226–233]
Gentamicin has widely been used for the treatment of S. aureus infections in the community, such as bullous impetigo, in Japan. Arbekacin has been used as an anti-MRSA agent in Japan. Resistance to gentamicin is mainly due to gentamicin modification (inactivation) by 6′-aminoglycoside N-acetyltransferase-2′′-aminoglycoside O-phosphotransferase [AAC(6′)-APH(2′′)] (Fig. 11). Higher production of AAC(6′)-APH(2′′) results in resistance to arbekacin as well; a breakpoint for arbekacin resistance is 2 μg/ml (Japanese Society for Chemotherapy).
Chemical structures of aminoglycosides and target residues of enzymatic modification. Chemical structures of gentamicin, kanamycin, and arbekacin are shown. Hydroxyl and amino residues shown in red and blue, respectively, are the targets of the enzymes. The inactivating enzymes are from Table 2
Fluoroquinolones [32, 72, 234, 235]
In S. aureus, fluoroquinolone resistance is caused by a mutation in the DNA gyrase gene (gyr) and/or topoisomerase IV gene (grl). The quinolone resistance-determining region (QRDR) is located in gyrA and grlA. Fluoroquionlone resistance is at a severe level for HA-MRSA. Ciprofloxacin and levofloxacin resistance has been noted. Some CA-MRSA clones have also become fluoroquionlone resistant. For example, the USA300 clone is now multiple-drug resistant, and ciprofloxacin resistance (one of such multiple resistance) is due to a Ser84Leu chromosomal mutation of gyrase A encoded by the gyrA gene.
Rifampicin [236–238]
Rifampicin exhibits extremely high in vitro activity against S. aureus including MRSA. However, single use of rifampicin results in the rapid emergence of resistance. Rifampicin resistance is caused by a mutation in the RFP resistance-determining clusters I and II located in the β-subunit gene (rpoB) of bacterial RNA polymerase.
To prevent rifampicin-resistance, combination therapy with appropriative drugs (such as linezolid) has been continuously investigated. Moreover, rifampicin-resistant MRSA has been isolated in tuberculosis wards.
Macrolide-lincosamide-streptogramin B (MLSB) [32, 35, 71, 72, 111, 145, 219, 239–243]
Clindamycin has been, in part, used for the treatment of S. aureus infections including MRSA infections. However, failure of treatment by clindamycin has already been noted due to the presence of inducible clindamycin resistance. Inducible-clindamycin-resistant strains are always resistant to erythromycin, and inducible clindamycin resistance can be induced by erythromycin. Inducible clindamycin resistance assay is called the D test (Fig. 12b, c). Clindamycin cannot be an effective inducer. The mechanism of clindamycin resistance is considered to be that in the absence of an erythromycin inducer, a ribosome-binding (SD) sequence of clindamycin resistance mRNA is hidden due to the mRNA secondary structure, and thus clindamycin resistance mRNA cannot be translated, while with the presence of the inducer, this secondary structure of mRNA is disrupted resulting in translation of clindamycin resistance mRNA (Fig. 12a).
Model for translational regulation of ermC expression (a) and D test (b, c). Data from Refs. [298, 299]. a Secondary structures upstream of ermC mRNA in the absence and presence of erythromycin are shown. The leader peptide sequence (19 amino acids) and ermC are indicated as gray and dark gray lines, respectively. SD-1 Shine-Dalgarno sequence for leader peptide translation, SD-2 Shine-Dalgarno sequence for ermC translation. In the absence of EM, SD-2 is masked by stem-loop (constructed by sequences 3 and 4). In the presence of EM, a complex of ribosome and EM tightly binds to the leader peptide sequence. This induces disruption of the two stem-loops, resulting in initiation of ermC translation from unmasked SD-2. b, c Results of D test of ermC-positive strain (b) and ermC-negative erythromycin-resistant strain (c) are shown. EM erythromycin, CLDM clindamycin
Trimethoprim–sulfamethoxazole (TMP–SMX) [244]
Mild SSTIs may be treated with TMP-SMX. Occasionally, the dfrA gene is associated with SCCmec-n1.
Mupirocin [72, 111, 245, 246]
Mupirocin has been used in the eradication of naris-colonizing MRSA. As for resistance, ileS mutation is low-level resistance (MICs; 8–256 μg/ml), while the mupA (ileS-2) gene confers high-level resistance (MICs; ≥512 μg/ml). The multiple-drug-resistant USA300 clone (ST8) carries the mupA gene on a plasmid. Also, the multiple-drug-resistant ST80 clone carries the mupA gene.
Fusidic acid [145, 247–249]
High-level fusidic acid-resistance (MICs; >12 μg/ml) involves mutation of the fusA gene. ST80 CA-MRSA, a worldwide clone, carries the fusA gene, and thus the fusA gene has been a marker of ST80 CA-MRSA spread in Europe. Occasionally, the fusB1 gene is found on SCCmec (SCCmec-n1 carrying dfrA).
MRSA in the community in Japan
Young school age children have nasal MRSA at higher levels with a peak at 5–9 years (isolation rate 2.2%), compared with other age groups, e.g., university students (isolation rate <0.25%) [250]. In other reports, isolation rates were 3.8 or 4.3% for children in kindergartens and day-care centers [251, 252]. All MRSA strains are PVL-negative. In their families, MRSA transmits among family members [250]. Examples include ST8/SCCmecIVx MRSA or ST764/SCCmecII MRSA. Moreover, children have ST88/SCCmecIVx MRSA (associated with bullous impetigo), ST857/SCCmecX (IIb) MRSA (new type), or ST380/SCCmecIVc and ST22/SCCmecI MRSA (rare isolates). The HA-MRSA New York/Japan clone (ST5/SCCmecII) is also detected from children; in one study, it was only from pediatric outpatients [250].
MRSA is also distributed in the environment. Although no MRSA was isolated from play settings such as karaoke, pachinko, amusement arcades, and gyms, MRSA was isolated from public transport (trains, subways, or buses) at 2.5% [155]. They were all PVL-negative. Such MRSA falls into three groups: i) bullous impetigo-related CA-MRSA, such as ST8/SCCmecIVx or I MRSA and ST88/SCCmecIVx MRSA, ii) the HA-MRSA New York/Japan clone, and iii) uncategorized MRSA [155]. It is conceivable that such MRSA is transferred from the hands of passengers in trains, subways, or buses.
The New York/Japan clone, a typical HA-MRSA clone in Japan, is occasionally isolated from the community, as described above. It could be due to (e.g.) frequent pediatric visits to hospitals (which could be a high risk factor for pediatric MRSA colonization); for instance, some children visit hospitals for treatment several times a month. More importantly, ST764/SCCmecII MRSA is spreading not only in hospitals but also widely in the community [155, 253]. The ST764 clone is a derivative of the New York/Japan clone and is multiple-drug-resistant just like the New York/Japan clone; however, it seems to persist more in the community. The ST764 MRSA clone was first isolated from bacteremia in an inpatient in Sado (Niigata) in 2005 [253], and then from the hands of a medical student (undergoing clinical practice) in 2007 [155]. ST764 was also isolated from an intrafamilial transmission case in 2008 [250], from necrotizing fasciitis in the community in 2008 (unpublished data), and in enteritis in the community in 2009 (unpublished data), suggesting that the ST764 clone is an emerging hybrid-type clone with a higher ability to adapt to the community. The ST764 clone lacks SaPIm1/n1 (carrying three superantigen genes, tst, sec, and sel) present in the New York/Japan clone, but instead has a superantigen gene seb2 [a variant of seb (seb1)] [138]. The seb1 gene is detected in the major PVL-positive CA-MRSA [ST59/SCCmecVII (or V)] in Taiwan [32]. The seb gene may contribute to adaptation to the community.
It has also turned out recently that ST8/SCCmecIVx is a major CA-MRSA clone in Japan. It exhibits mainly spa606 (with some minor spa types) and is PVL- and ACME-negative and distinct from USA300. It has been associated with a wide range of community infections, including bullous impetigo [141], atopic dermatitis (unpublished data), slight wound infection in a sumo-wrestling team [254], bacteremia and subsequent epidural abscesses [255], and fatal pneumonia [256]. A very similar MRSA was also isolated from cutaneous abscesses (in the community) of Japanese family members who had moved to Hong Kong [137].
Bullous impetigo is a common disease among children, and CA-MRSA is frequently isolated from these infected children (at 19–51%) [141, 194]. Such bullous impetigo-associated CA-MRSA (biCA-MRSA) belongs to ST8, ST88, ST89, and ST91 [141]. Those isolates are positive for the cna gene in many cases (at 86.4%) [257]. We speculate that biCA-MRSA infects skin soft tissues through binding to collagens, exposed by scratching, to cause bullous impetigo. biCA-MRSA is highly resistant to gentamicin (at 80–90%) [141, 258], reflecting frequent use of gentamicin ointment for the treatment of bullous impetigo.
Recently, a variety of PVL-positive MRSA clones have been accumulating in Japan (albeit still at a low frequency), including ST30/SCCmecIVa [92], ST30/SCCmecIVc [75, 141], and their ST single locus variants, ST765/SCCmecIVx (or I) [71] and ST1335/SCCmecIVc (unpublished data), ST59/SCCmecVII (or V) [138], ST8/SCCmecIVa (US300 clone) [125], and ST22/SCCmecIVa [144]. Of those, ST59 may originate in Taiwan (although the patient and his family members had never been to Taiwan). As for USA300, the first isolate was from a visitor (Indian girl) from the US in 2007 in Tokyo. This was followed by a Japanese child infection case in Saitama in 2008 (this patient and her family members had never been abroad), indicating the spread of USA300 in Japan, although the two isolates in 2007 and 2008 are divergent [126].
CA-MRSA infections in other areas of Asia
Taiwan
In Taiwan, the ST59/SCCmecVII (or alternatively assigned as V) clone has been noted as a major CA-MRSA clone [32, 259, 260]. They are mostly PVL-positive [32, 259]. On the other hand, ST59/SCCmecIV is mostly PVL-negative [32, 259, 260]. Nasal colonization of healthy adults (age of >18 years) with MRSA has also been reported (isolation rate 3.8%) [260]. Of those, PVL-positive ST59 and PVL-negative ST59 accounted for approximately 38 and 46%, respectively. The PVL-positive and -negative ST59/SCCmecIV/V clones were relatively multidrug-resistant [(e.g.) tetracycline, kanamycin, MLSB, chloramphenicol] [32]. Although the most frequent clone in the hospital in Taiwan is ST239/SCCmecIII (Hungarian clone), PVL-positive and -negative ST59 clones are spreading not only in the community but also in the hospital [32, 259]. Currently, a fatal case of a 12-year-old boy who was infected by USA300 was reported [128].
China
Recent studies have shown the appearance of a worldwide clone, ST30 CA-MRSA, in China [106, 261, 262]. Yu et al. [261] investigated community- and hospital- acquired S. aureus isolates from the patients of a teaching hospital in Wenzhou and found two CA-MRSA isolates. One was a worldwide clone, PVL-positive ST30/SCCmecIV, and mostly susceptible to non-β-lactam antimicrobial agents; the other was PVL-positive ST398/SCCmecIV, resistant to erythromycin and gentamicin. In the same study, ST398/SCCmecIV was also found as HA-MRSA.
More information about the molecular characteristics of CA-MRSA is available in Hong Kong [136, 262, 263]. Ho and his colleagues collected CA-MRSA strains from patients and their household members from 2004 to 2007 [136, 263]. Most of the CA-MRSA isolates (78% of all CA-MRSA isolates) were associated with SSTI. Among the CA-MRSA isolates, 72% were positive for PVL. Sixty-seven percent of the PVL-positive CA-MRSA isolates showed ST30/SCCmecIV, typically characteristic of worldwide clones, and were mostly susceptible to non-β lactam antimicrobial agents. The other set of PVL-positive CA-MRSA isolates in Hong Kong was ST59. ST59 clones showed multidrug resistance (MLSB, tetracycline, or chloramphenicol), indicating similar characteristics to PVL-positive ST59 seen in Taiwan [32]. PVL-negative CA-MRSA isolates were also isolated and showed ST8/SCCmecIVx [262], which has typical characteristics of PVL-negative CA-MRSA in Japan [141]. Recently, after PVL-negative ST88/SCCmecIV CA-MRSA was reported in Japan [250], a similar PVL-negative ST88/SCCmecIVA CA-MRSA has also been isolated in Guangzhou, a city adjacent to Hong Kong [264]. A PVL-negative ST59/SCCmecIVA CA-MRSA was isolated in this study as well.
In hospitals in Hong Kong, ST239/SCCmecIII/IIIA is a major clone, and ST5/SCCmecII and ST45/SCCmecIV (and its variant) are found as well [58]. Currently, the ST45/SCCmecIV clone is increasing.
These data indicate that CA-MRSA strains in China have divergent genotypes closely related to clones isolated worldwide and in areas around mainland China including Taiwan and Japan.
Russia
There are many distinct MRSA clones in the territory of the Russian Federation. Among them, the ST239 Hungarian clone could be a major clone in Russian hospitals [265]. In addition, another unique ST239 MRSA clone (so-called Russian clone) was found in the far eastern part of Russia [57]. It can transfer multidrug resistance to other S. aureus or MRSA strains. In addition, one strain of PVL-positive worldwide CA-MRSA was isolated. Its genotype was ST30/spa19/SCCmecIVc [139]. A multidrug-resistant ST239 clone, described above, has been isolated from outpatients, in addition to inpatients, in Russia, indicating that the multidrug-resistant ST239 clone is spreading even in the community. This may be, in part, because the patients in Russia can obtain antibiotics without doctor’s prescription in the community. Therefore, selection pressure by antibiotics in the community is slightly higher than that in other countries like Japan or the US, suggesting that different criteria for CA-MRSA should be considered. However, further study is needed.
Korea
In Korea, MRSA is spreading in hospitals and in the community like in other countries [266]. But the major ST type is very unique. ST72, which is SCCmecIV and PVL negative, is the major CA-MRSA clone in Korea, but does not spread in other areas. The ST5/SCCmecII and ST239/SCCmecIIIa clone are spreading both in hospitals and in the community [266]. Only one case of a USA300-like clone has been reported [127].
MRSA in animals
MRSA has also been isolated from companion and livestock animals. Those MRSA strains have currently been noted as a source of infection in humans, and transmission cases are summarized in Table 3 [267–278]. For example, in case of companion animals, the EMRSA-15 clone (ST22/SCCmecIV/PVL-negative) has been isolated from dogs and cats in Europe, including the UK, Ireland, and Germany [279–284]; the Canadian epidemic MRSA-2 clone (ST5/SCCmecII/PVL-negative) from dogs and cats [275]; and the Canadian epidemic MRSA-5 clone (ST8/SCCmecIV/PVL-negative) from horses in North America [285]. In addition, human PVL-positive CA-MRSA clones have been isolated from companion animals [274, 276]. Sing et al. [274] have reported a recurrent infection case of PVL-positive ST80 CA-MRSA through intrafamilial transmission among households and cat. To prevent recurrent infections, the eradication of MRSA from companion animals, as well as patients and their households, is important. Other than the ST80 clone, transmission of the USA300 strain from a cat has also been reported [276].
On the other hand, MRSA strains have also been isolated from livestock (e.g., pigs, cattle, and chickens) in (e.g.) the Netherlands, Belgium, Germany, Hungary, the US, Australia, and Korea [267, 269, 286–290]. van Rijen et al. [291] concluded in their report that the infection risk for pig and cow farmers (32%) is three times higher than for people who do not have direct contact with those animals. ST398 clones with SCCmecIV or V, as animal-associated CA-MRSA clones, have been widely spreading among livestock [269, 286–290, 292] and companion animals (e.g., horses) [293]. ST398 clones were originally known as non-typeable MRSA in PFGE due to methylation of C-residues in the SmaI recognition site [294]. Currently, transmission of those animal clones to farmers and their households has been noted (Table 3). Furthermore, isolation of the ST398 clone from a patient, family member (carrier), and nurse (carrier) who was taking care of that patient has also been observed, indicating possible transmission from human to human [268]. Those ST398 clones were PVL-negative. However, PVL-positive ST398 CA-MRSA has also been isolated from a human patient with abscesses in China although there had been no description of contact with animals [261].
Other than ST398, possible transmission of ST9 clones with SCCmecIII or V among pigs and farmers has also been reported in Asia. In those cases, no ST398 clones were observed in contrast to Europe, North America, and Australia, suggesting a spread of different clones in Asia [295, 296].
In Japan, MRSA with SCCmecIV has been isolated from chicken meat, although MLST was unknown [297]. On the other hand, we have examined the carriage of MRSA among healthy companion animals (mainly dogs and cats) in Japan. However, no MRSA has been isolated from healthy companion animals, although other methicillin-resistant Staphylococcus spp., such as methicillin-resistant S. intermedius, have been isolated.
Companion and livestock animals, which are carriers of MRSA, are potential risk factors for MRSA infection in humans. In cases of recurrent MRSA infection among family members, tests should be conducted on MRSA carriage in animals in the house or farm. If positive, MRSA eradication should be carefully considered.
Closing remarks
Historically, several waves of MRSA infections have attacked humans, including those due to penicillin G-resistant S. aureus in the mid 1940s, multidrug-resistant S. aureus in the late 1950s, and MRSA since 1961 (but with worldwide outbreaks in the 1970s–1980s). They all seem to have been driven by high levels of antimicrobial agent use in hospitals. The concept of MRSA infection has drastically changed since the late 1990s, when CA-MRSA was noted worldwide. CA-MRSA is well adapted to the community and triggers even unexpected cases of diseases for clinicians. Apparently, new countermeasures against the emerging threat (such as active surveillance of CA-MRSA in high risk groups, such as children and their families, and its eradication) must be considered and constructed.
References
Tenover F, Gorwitz RJ. The epidemiology of Staphylococcus infections In: Fischetti VA, Novick RP, Ferretti JJ, Portnoy DA, Rood JI, editors. Gram-positive pathogens. 2nd ed. Washington, DC: American Society for Microbiology; 2006. p. 526-34.
Lowy FD. Staphylococcus aureus infections. N Engl J Med. 1998;339:520–32.
Safdar N, Bradley EA. The risk of infection after nasal colonization with Staphylococcus aureus. Am J Med. 2008;121:310–5.
Durupt F, Mayor L, Bes M, Reverdy ME, Vandenesch F, Thomas L, et al. Prevalence of Staphylococcus aureus toxins and nasal carriage in furuncles and impetigo. Br J Dermatol. 2007;157:1161–7.
Bryskier A. Penicillin. In: Andre B, editor. Antimicrobial agents: antibacterials and antifungals. Washington, DC: American Society for Microbiology; 2005. p. 113–62.
Projan SJ, Ruzin A. Antibiotic resistance in the staphylococci. In: Fischetti VA, Novick RP, Ferretti JJ, Portnoy DA, Rood JI, editors. Gram-positive pathogens. 2nd ed. Washington: American Society for Microbiology; 2006. p. 587–97.
Lyon BR, Skurray R. Antimicrobial resistance of Staphylococcus aureus: genetic basis. Microbiol Rev. 1987;51:88–134.
From the Center for Disease Control and Prevention. Four pediatric deaths from community-acquired methicillin-resistant Staphylococcus aureus—Minnesota and North Dakota, 1997–1999. MMWR Morb Mortal Wkly Rep 1999;48:707–10.
Vandenesch F, Naimi T, Enright MC, Lina G, Nimmo GR, Heffernan H, et al. Community-acquired methicillin-resistant Staphylococcus aureus carrying Panton-Valentine leukocidin genes: worldwide emergence. Emerg Infect Dis. 2003;9:978–84.
Zetola N, Francis JS, Nuermberger EL, Bishai WR. Community-acquired methicillin-resistant Staphylococcus aureus: an emerging threat. Lancet Infect Dis. 2005;5:275–86.
Deurenberg RH, Stobberingh EE. The evolution of Staphylococcus aureus. Infect Genet Evol. 2008;8:747–63.
Diep BA, Stone GG, Basuino L, Graber CJ, Miller A, des Etages SA, et al. The arginine catabolic mobile element and staphylococcal chromosomal cassette mec linkage: convergence of virulence and resistance in the USA300 clone of methicillin-resistant Staphylococcus aureus. J Infect Dis. 2008;197:1523–30.
Levin BR, Lipsitch M, Perrot V, Schrag S, Antia R, Simonsen L, et al. The population genetics of antibiotic resistance. Clin Infect Dis. 1997;24(Suppl 1):S9–16.
Levin BR. Minimizing potential resistance: a population dynamics view. Clin Infect Dis. 2001;33(Suppl 3):S161–9.
Ender M, McCallum N, Adhikari R, Berger-Bachi B. Fitness cost of SCCmec and methicillin resistance levels in Staphylococcus aureus. Antimicrob Agents Chemother. 2004;48:2295–7.
Lee SM, Ender M, Adhikari R, Smith JM, Berger-Bachi B, Cook GM. Fitness cost of staphylococcal cassette chromosome mec in methicillin-resistant Staphylococcus aureus by way of continuous culture. Antimicrob Agents Chemother. 2007;51:1497–9.
Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol. 2000;38:1008–15.
Shopsin B, Gomez M, Montgomery SO, Smith DH, Waddington M, Dodge DE, et al. Evaluation of protein A gene polymorphic region DNA sequencing for typing of Staphylococcus aureus strains. J Clin Microbiol. 1999;37:3556–63.
Harmsen D, Claus H, Witte W, Rothganger J, Turnwald D, Vogel U. Typing of methicillin-resistant Staphylococcus aureus in a university hospital setting by using novel software for spa repeat determination and database management. J Clin Microbiol. 2003;41:5442–8.
Koreen L, Ramaswamy SV, Graviss EA, Naidich S, Musser JM, Kreiswirth BN. spa typing method for discriminating among Staphylococcus aureus isolates: implications for use of a single marker to detect genetic micro- and macrovariation. J Clin Microbiol. 2004;42:792–9.
Strommenger B, Cuny C, Werner G, Witte W. Obvious lack of association between dynamics of epidemic methicillin-resistant Staphylococcus aureus in central Europe and agr specificity groups. Eur J Clin Microbiol Infect Dis. 2004;23:15–9.
Gilot P, Lina G, Cochard T, Poutrel B. Analysis of the genetic variability of genes encoding the RNA III-activating components Agr and TRAP in a population of Staphylococcus aureus strains isolated from cows with mastitis. J Clin Microbiol. 2002;40:4060–7.
Kinoshita M, Kobayashi N, Nagashima S, Ishino M, Otokozawa S, Mise K, et al. Diversity of staphylocoagulase and identification of novel variants of staphylocoagulase gene in Staphylococcus aureus. Microbiol Immunol. 2008;52:334–48.
Sakai F, Takemoto A, Watanabe S, Aoyama K, Ohkubo T, Yanahira S, et al. Multiplex PCRs for assignment of Staphylocoagulase types and subtypes of type VI Staphylocoagulase. J Microbiol Methods. 2008;75:312–7.
Oliveira DC, de Lencastre H. Multiplex PCR strategy for rapid identification of structural types and variants of the mec element in methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother. 2002;46:2155–61.
Zhang K, McClure JA, Elsayed S, Louie T, Conly JM. Novel multiplex PCR assay for characterization and concomitant subtyping of staphylococcal cassette chromosome mec types I to V in methicillin-resistant Staphylococcus aureus. J Clin Microbiol. 2005;43:5026–33.
Ito T, Ma XX, Takeuchi F, Okuma K, Yuzawa H, Hiramatsu K. Novel type V staphylococcal cassette chromosome mec driven by a novel cassette chromosome recombinase, ccrC. Antimicrob Agents Chemother. 2004;48:2637–51.
International Working Group on the Classification of Staphylococcal Cassette Cromosome Elements (IWG-SCC). Classification of staphylococcal cassette chromosome mec (SCCmec): guidelines for reporting novel SCCmec elements. Antimicrob Agents Chemother. 2009;53:4961–7.
Enright MC, Robinson DA, Randle G, Feil EJ, Grundmann H, Spratt BG. The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA). Proc Natl Acad Sci USA. 2002;99:7687–92.
Robinson DA, Enright MC. Evolutionary models of the emergence of methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother. 2003;47:3926–34.
Boyle-Vavra S, Ereshefsky B, Wang CC, Daum RS. Successful multiresistant community-associated methicillin-resistant Staphylococcus aureus lineage from Taipei, Taiwan, that carries either the novel Staphylococcal chromosome cassette mec (SCCmec) type V T or SCCmec type IV. J Clin Microbiol. 2005;43:4719–30.
Takano T, Higuchi W, Otsuka T, Baranovich T, Enany S, Saito K, et al. Novel characteristics of community-acquired methicillin-resistant Staphylococcus aureus strains belonging to multilocus sequence type 59 in Taiwan. Antimicrob Agents Chemother. 2008;52:837–45.
Tenover FC, McDougal LK, Goering RV, Killgore G, Projan SJ, Patel JB, et al. Characterization of a strain of community-associated methicillin-resistant Staphylococcus aureus widely disseminated in the United States. J Clin Microbiol. 2006;44:108–18.
Tenover FC, Goering RV. Methicillin-resistant Staphylococcus aureus strain USA300: origin and epidemiology. J Antimicrob Chemother. 2009;64:441–6.
Strommenger B, Kettlitz C, Werner G, Witte W. Multiplex PCR assay for simultaneous detection of nine clinically relevant antibiotic resistance genes in Staphylococcus aureus. J Clin Microbiol. 2003;41:4089–94.
Zhang K, McClure JA, Elsayed S, Louie T, Conly JM. Novel multiplex PCR assay for simultaneous identification of community-associated methicillin-resistant Staphylococcus aureus strains USA300 and USA400 and detection of mecA and Panton-Valentine leukocidin genes, with discrimination of Staphylococcus aureus from coagulase-negative staphylococci. J Clin Microbiol. 2008;46:1118–22.
Bonnstetter KK, Wolter DJ, Tenover FC, McDougal LK, Goering RV. Rapid multiplex PCR assay for identification of USA300 community-associated methicillin-resistant Staphylococcus aureus isolates. J Clin Microbiol. 2007;45:141–6.
Reva I, Higuchi W, Takano T, Singur O, Ozaki K, Isobe H, et al. A rapid screening method for Panton-Valentine leucocidin-positive community-acquired methicillin-resistant Staphylococcus aureus belonging to multilocus sequence type 30 and its related clone using a combination of multiplex PCR and pulsed-field gel electrophoresis. J Infect Chemother. 2009;15:75–83.
Brumfitt W, Hamilton-Miller J. Methicillin-resistant Staphylococcus aureus. N Engl J Med. 1989;320:1188–96.
Naimi TS, LeDell KH, Como-Sabetti K, Borchardt SM, Boxrud DJ, Etienne J, et al. Comparison of community- and health care-associated methicillin-resistant Staphylococcus aureus infection. JAMA. 2003;290:2976–84.
Tacconelli E, Tumbarello M, Cauda R. Staphylococcus aureus infections. N Engl J Med. 1998;339:2026–7.
Aires de Sousa M, Conceicao T, Simas C, de Lencastre H. Comparison of genetic backgrounds of methicillin-resistant and -susceptible Staphylococcus aureus isolates from Portuguese hospitals and the community. J Clin Microbiol. 2005;43:5150–7.
Noto MJ, Kreiswirth BN, Monk AB, Archer GL. Gene acquisition at the insertion site for SCCmec, the genomic island conferring methicillin resistance in Staphylococcus aureus. J Bacteriol. 2008;190:1276–83.
Oliveira DC, Tomasz A, de Lencastre H. The evolution of pandemic clones of methicillin-resistant Staphylococcus aureus: identification of two ancestral genetic backgrounds and the associated mec elements. Microb Drug Resist. 2001;7:349–61.
EMRSA-16: a new epidemic strain of Staphylococcus aureus. Commun Dis Rep CDR Wkly 1993;3:25.
Aucken HM, Ganner M, Murchan S, Cookson BD, Johnson AP. A new UK strain of epidemic methicillin-resistant Staphylococcus aureus (EMRSA-17) resistant to multiple antibiotics. J Antimicrob Chemother. 2002;50:171–5.
Conceicao T, Aires-de-Sousa M, Fuzi M, Toth A, Paszti J, Ungvari E, et al. Replacement of methicillin-resistant Staphylococcus aureus clones in Hungary over time: a 10-year surveillance study. Clin Microbiol Infect. 2007;13:971–9.
Gordon RJ, Lowy FD. Pathogenesis of methicillin-resistant Staphylococcus aureus infection. Clin Infect Dis. 2008;46(Suppl 5):S350–9.
Hsu LY, Koh TH, Tan TY, Ito T, Ma XX, Lin RT, et al. Emergence of community-associated methicillin-resistant Staphylococcus aureus in Singapore: a further six cases. Singapore Med J. 2006;47:20–6.
Chongtrakool P, Ito T, Ma XX, Kondo Y, Trakulsomboon S, Tiensasitorn C, et al. Staphylococcal cassette chromosome mec (SCCmec) typing of methicillin-resistant Staphylococcus aureus strains isolated in 11 Asian countries: a proposal for a new nomenclature for SCCmec elements. Antimicrob Agents Chemother. 2006;50:1001–12.
Deurenberg RH, Vink C, Kalenic S, Friedrich AW, Bruggeman CA, Stobberingh EE. The molecular evolution of methicillin-resistant Staphylococcus aureus. Clin Microbiol Infect. 2007;13:222–35.
Krishna BV, Patil A, Chandrasekhar MR. Methicillin resistant Staphylococcus aureus infections-implications in hospital infection control. Indian J Public Health. 2007;51:43–6.
Boucher HW, Corey GR. Epidemiology of methicillin-resistant Staphylococcus aureus. Clin Infect Dis. 2008;46(Suppl 5):S344–9.
Stratchounski LS, Dekhnich AV, Kretchikov VA, Edelstain IA, Narezkina AD, Afinogenov GE, et al. Antimicrobial resistance of nosocomial strains of Staphylococcus aureus in Russia: results of a prospective study. J Chemother. 2005;17:54–60.
van Trijp MJ, Melles DC, Hendriks WD, Parlevliet GA, Gommans M, Ott A. Successful control of widespread methicillin-resistant Staphylococcus aureus colonization and infection in a large teaching hospital in the Netherlands. Infect Control Hosp Epidemiol. 2007;28:970–5.
Simor AE, Loeb M. The management of infection and colonization due to methicillin-resistant Staphylococcus aureus: a CIDS/CAMM position paper. Can J Infect Dis. 2004;15:39–48.
Higuchi W, Singur O, Iwao Y, Hung WC, Shi D, Takano T, et al. A novel multiple drug-resistant ST239 methicillin-resistant Staphylococcus aureus clone which is capable of donating drug resistance from Russia (abstract no. C2-119). In: 49th interscience conference on antimicrobial agents and chemotherapy. San Francisco, CA: American Scoiety for Microbiology; 2009.
Ho PL, Chow KH, Lo PY, Lee KF, Lai EL. Changes in the epidemiology of methicillin-resistant Staphylococcus aureus associated with spread of the ST45 lineage in Hong Kong (DMID-08–518 revised). Diagn Microbiol Infect Dis. 2009;64:131–7.
Holden MT, Lindsay JA, Corton C, Quail MA, Cockfield JD, Pathak S, et al. Genome sequence of a recently emerged highly-transmissible, multi-antibiotic and antiseptic resistant, variant of methicillin-resistant Staphylococcus aureus (MRSA) sequence-type 239 (TW). J Bacteriol. 2009. doi:10.1128/JB.01255-09.
Edgeworth JD, Yadegarfar G, Pathak S, Batra R, Cockfield JD, Wyncoll D, et al. An outbreak in an intensive care unit of a strain of methicillin-resistant Staphylococcus aureus sequence type 239 associated with an increased rate of vascular access device-related bacteremia. Clin Infect Dis. 2007;44:493–501.
Ko KS, Lee JY, Suh JY, Oh WS, Peck KR, Lee NY, et al. Distribution of major genotypes among methicillin-resistant Staphylococcus aureus clones in Asian countries. J Clin Microbiol. 2005;43:421–6.
Xu BL, Zhang G, Ye HF, Feil EJ, Chen GR, Zhou XM, et al. Predominance of the Hungarian clone (ST 239-III) among hospital-acquired methicillin-resistant Staphylococcus aureus isolates recovered throughout mainland China. J Hosp Infect. 2009;71:245–55.
Amaral MM, Coelho LR, Flores RP, Souza RR, Silva-Carvalho MC, Teixeira LA, et al. The predominant variant of the Brazilian epidemic clonal complex of methicillin-resistant Staphylococcus aureus has an enhanced ability to produce biofilm and to adhere to and invade airway epithelial cells. J Infect Dis. 2005;192:801–10.
Vivoni AM, Diep BA, de Gouveia Magalhaes AC, Santos KR, Riley LW, Sensabaugh GF, et al. Clonal composition of Staphylococcus aureus isolates at a Brazilian university hospital: identification of international circulating lineages. J Clin Microbiol. 2006;44:1686–91.
Bartels MD, Nanuashvili A, Boye K, Rohde SM, Jashiashvili N, Faria NA, et al. Methicillin-resistant Staphylococcus aureus in hospitals in Tbilisi, the Republic of Georgia, are variants of the Brazilian clone. Eur J Clin Microbiol Infect Dis. 2008;27:757–60.
Szczepanik A, Koziol-Montewka M, Al-Doori Z, Morrison D, Kaczor D. Spread of a single multiresistant methicillin-resistant Staphylococcus aureus clone carrying a variant of staphylococcal cassette chromosome mec type III isolated in a university hospital. Eur J Clin Microbiol Infect Dis. 2007;26:29–35.
Klevens RM, Morrison MA, Nadle J, Petit S, Gershman K, Ray S, et al. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA. 2007;298:1763–71.
Zeller JL, Burke AE, Glass RM. JAMA patient page. MRSA infections. JAMA. 2007;298:1826.
From the Centers for Disease Control and Prevention. MRSA in Healthcare Settings. 2006. http://www.cdc.gov/ncidod/dhqp/ar_MRSA_spotlight_2006.html.
Siegel JD, Rhinehart E, Jackson M, Chiarello L. Management of multidrug-resistant organisms in healthcare settings, 2006. Atlanta: Centers for Disease Control and Prevention; 2006.
Yamamoto T, Dohmae S, Saito K, Otsuka T, Takano T, Chiba M, et al. Molecular characteristics and in vitro susceptibility to antimicrobial agents, including the des-fluoro(6) quinolone DX-619, of Panton-Valentine leucocidin-positive methicillin-resistant Staphylococcus aureus isolates from the community and hospitals. Antimicrob Agents Chemother. 2006;50:4077–86.
Diep BA, Gill SR, Chang RF, Phan TH, Chen JH, Davidson MG, et al. Complete genome sequence of USA300, an epidemic clone of community-acquired methicillin-resistant Staphylococcus aureus. Lancet. 2006;367:731–9.
Takano T, Higuchi W, Yamamoto T. Superior in vitro activity of carbapenems over anti-methicillin-resistant Staphylococcus aureus (MRSA) and some related antimicrobial agents for community-acquired MRSA but not for hospital-acquired MRSA. J Infect Chemother. 2009;15:54–7.
Redziniak DE, Diduch DR, Turman K, Hart J, Grindstaff TL, MacKnight JM, et al. Methicillin-resistant Staphylococcus aureus (MRSA) in the Athlete. Int J Sports Med. 2009;30:557–62.
Okubo T, Yabe S, Otsuka T, Takizawa Y, Takano T, Dohmae S, et al. Multifocal pelvic abscesses and osteomyelitis from community-acquired methicillin-resistant Staphylococcus aureus in a 17-year-old basketball player. Diagn Microbiol Infect Dis. 2008;60:313–8.
Wijaya L, Hsu LY, Kurup A. Community-associated methicillin-resistant Staphylococcus aureus: overview and local situation. Ann Acad Med Singapore. 2006;35:479–86.
Workshop 47-11. Origin, Epidemiology, Diagnosis and Management of CA-MRSA. In: 47th interscience conference on antimicrobial agents and chemotherapy. Chicago, IL: American Scoiety for Microbiology; 2007.
Johannes RS. Epidemiology of early-onset bloodstream infection and implications for treatment. Am J Infect Control. 2008;36:S171.e13–7.
von Eiff C, Becker K, Machka K, Stammer H, Peters G. Nasal carriage as a source of Staphylococcus aureus bacteremia. Study Group. N Engl J Med. 2001;344:11–6.
Demling RH, Waterhouse B. The increasing problem of wound bacterial burden and infection in acute and chronic soft-tissue wounds caused by methicillin-resistant Staphylococcus aureus. J Burns Wounds. 2007;7:e8.
Wertheim HF, Vos MC, Ott A, van Belkum A, Voss A, Kluytmans JA, et al. Risk and outcome of nosocomial Staphylococcus aureus bacteraemia in nasal carriers versus non-carriers. Lancet. 2004;364:703–5.
Baba-Moussa L, Anani L, Scheftel JM, Couturier M, Riegel P, Haikou N, et al. Virulence factors produced by strains of Staphylococcus aureus isolated from urinary tract infections. J Hosp Infect. 2008;68:32–8.
Blomquist PH. Methicillin-resistant Staphylococcus aureus infections of the eye and orbit (an American Ophthalmological Society thesis). Trans Am Ophthalmol Soc. 2006;104:322–45.
Bruns AS, Sood N. Community-acquired methicillin-resistant Staphylococcus aureus epidural abscess with bacteremia and multiple lung abscesses: case report. Am J Crit Care. 2009;18(88):6–7.
Munckhof WJ, Krishnan A, Kruger P, Looke D. Cavernous sinus thrombosis and meningitis from community-acquired methicillin-resistant Staphylococcus aureus infection. Intern Med J. 2008;38:283–7.
Kefala-Agoropoulou K, Protonotariou E, Vitti D, Sarafidou S, Anastasiou A, Kollios K, et al. Life-threatening infection due to community-acquired methicillin-resistant Staphylococcus aureus: case report and review. Eur J Pediatr. 2010;169:47–53.
Miller LG, Perdreau-Remington F, Rieg G, Mehdi S, Perlroth J, Bayer AS, et al. Necrotizing fasciitis caused by community-associated methicillin-resistant Staphylococcus aureus in Los Angeles. N Engl J Med. 2005;352:1445–53.
Dohin B, Gillet Y, Kohler R, Lina G, Vandenesch F, Vanhems P, et al. Pediatric bone and joint infections caused by Panton-Valentine leukocidin-positive Staphylococcus aureus. Pediatr Infect Dis J. 2007;26:1042–8.
Enany S, Higuchi W, Okubo T, Takano T, Enany M, Yamamoto T. Brain abscess caused by Panton-Valentine leukocidin-positive community-acquired methicillin-resistant Staphylococcus aureus in Egypt, April 2007. Euro Surveill 2007;12:E070927.2.
Fowler A, Mackay A. Community-acquired methicillin-resistant Staphylococcus aureus pyomyositis in an intravenous drug user. J Med Microbiol. 2006;55:123–5.
Fortes CQ, Espanha CA, Bustorff FP, Zappa BC, Ferreira AL, Moreira RB, et al. First reported case of infective endocarditis caused by community-acquired methicillin-resistant Staphylococcus aureus not associated with healthcare contact in Brazil. Braz J Infect Dis. 2008;12:541–3.
Ito T, Iijima M, Fukushima T, Nonoyama M, Ishii M, Baranovich T, et al. Pediatric pneumonia death caused by community-acquired methicillin-resistant Staphylococcus aureus, Japan. Emerg Infect Dis. 2008;14:1312–4.
Adem PV, Montgomery CP, Husain AN, Koogler TK, Arangelovich V, Humilier M, et al. Staphylococcus aureus sepsis and the Waterhouse-Friderichsen syndrome in children. N Engl J Med. 2005;353:1245–51.
Hageman JC, Uyeki TM, Francis JS, Jernigan DB, Wheeler JG, Bridges CB, et al. Severe community-acquired pneumonia due to Staphylococcus aureus, 2003–04 influenza season. Emerg Infect Dis. 2006;12:894–9.
Mandell LA, Wunderink RG, Anzueto A, Bartlett JG, Campbell GD, Dean NC, et al. Infectious Diseases Society of America/American Thoracic Society consensus guidelines on the management of community-acquired pneumonia in adults. Clin Infect Dis. 2007;44(Suppl 2):S27–72.
Hota B, Ellenbogen C, Hayden MK, Aroutcheva A, Rice TW, Weinstein RA. Community-associated methicillin-resistant Staphylococcus aureus skin and soft tissue infections at a public hospital: do public housing and incarceration amplify transmission? Arch Intern Med. 2007;167:1026–33.
From the Centers for Disease Control and Prevention. Methicillin-resistant Staphylococcus aureus infections in correctional facilities-Georgia, California, and Texas, 2001–2003. MMWR Morb Mortal Wkly Rep. 2003;52:992–6.
Ellis MW, Griffith ME, Jorgensen JH, Hospenthal DR, Mende K, Patterson JE. Presence and molecular epidemiology of virulence factors in methicillin-resistant Staphylococcus aureus strains colonizing and infecting soldiers. J Clin Microbiol. 2009;47:940–5.
Olsen RJ, Burns KM, Chen L, Kreiswirth BN, Musser JM. Severe necrotizing fasciitis in a human immunodeficiency virus-positive patient caused by methicillin-resistant Staphylococcus aureus. J Clin Microbiol. 2008;46:1144–7.
Seybold U, Kourbatova EV, Johnson JG, Halvosa SJ, Wang YF, King MD, et al. Emergence of community-associated methicillin-resistant Staphylococcus aureus USA300 genotype as a major cause of health care-associated blood stream infections. Clin Infect Dis. 2006;42:647–56.
Jenkins TC, McCollister BD, Sharma R, McFann KK, Madinger NE, Barron M, et al. Epidemiology of healthcare-associated bloodstream infection caused by USA300 strains of methicillin-resistant Staphylococcus aureus in 3 affiliated hospitals. Infect Control Hosp Epidemiol. 2009;30:233–41.
Malani PN, Rana MM, Banerjee M, Bradley SF. Staphylococcus aureus bloodstream infections: the association between age and mortality and functional status. J Am Geriatr Soc. 2008;56:1485–9.
Cooke FJ, Gkrania-Klotsas E, Howard JC, Stone M, Kearns AM, Ganner M, et al. Clinical, molecular and epidemiological description of a cluster of community-associated methicillin-resistant Staphylococcus aureus isolates from injecting drug users with bacteraemia. Clin Microbiol Infect. 2009. doi:10.1111/j.1469-0691.2009.02969.x.
Carrillo-Marquez MA, Hulten KG, Hammerman W, Mason EO, Kaplan SL. USA300 is the predominant genotype causing Staphylococcus aureus septic arthritis in children. Pediatr Infect Dis J. 2009;28:1076–80.
Hidron AI, Low CE, Honig EG, Blumberg HM. Emergence of community-acquired methicillin-resistant Staphylococcus aureus strain USA300 as a cause of necrotising community-onset pneumonia. Lancet Infect Dis. 2009;9:384–92.
Cheng VC, Lau YK, Lee KL, Yiu KH, Chan KH, Ho PL, et al. Fatal co-infection with swine origin influenza virus A/H1N1 and community-acquired methicillin-resistant Staphylococcus aureus. J Infect. 2009;59:366–70.
Christianson S, Golding GR, Campbell J, Mulvey MR. Comparative genomics of Canadian epidemic lineages of methicillin-resistant Staphylococcus aureus. J Clin Microbiol. 2007;45:1904–11.
Gilbert M, MacDonald J, Gregson D, Siushansian J, Zhang K, Elsayed S, et al. Outbreak in Alberta of community-acquired (USA300) methicillin-resistant Staphylococcus aureus in people with a history of drug use, homelessness or incarceration. CMAJ. 2006;175:149–54.
Li M, Diep BA, Villaruz AE, Braughton KR, Jiang X, DeLeo FR, et al. Evolution of virulence in epidemic community-associated methicillin-resistant Staphylococcus aureus. Proc Natl Acad Sci USA. 2009;106:5883–8.
Highlander SK, Hulten KG, Qin X, Jiang H, Yerrapragada S, Mason EO, Jr, et al. Subtle genetic changes enhance virulence of methicillin resistant and sensitive Staphylococcus aureus. BMC Microbiol. 2007;7:99.
Diep BA, Chambers HF, Graber CJ, Szumowski JD, Miller LG, Han LL, et al. Emergence of multidrug-resistant, community-associated, methicillin-resistant Staphylococcus aureus clone USA300 in men who have sex with men. Ann Intern Med. 2008;148:249–57.
Han LL, McDougal LK, Gorwitz RJ, Mayer KH, Patel JB, Sennott JM, et al. High frequencies of clindamycin and tetracycline resistance in methicillin-resistant Staphylococcus aureus pulsed-field type USA300 isolates collected at a Boston ambulatory health center. J Clin Microbiol. 2007;45:1350–2.
Graber CJ, Wong MK, Carleton HA, Perdreau-Remington F, Haller BL, Chambers HF. Intermediate vancomycin susceptibility in a community-associated MRSA clone. Emerg Infect Dis. 2007;13:491–3.
Hageman JC, Patel J, Franklin P, Miscavish K, McDougal L, Lonsway D, et al. Occurrence of a USA300 vancomycin-intermediate Staphylococcus aureus. Diagn Microbiol Infect Dis. 2008;62:440–2.
Mwangi MM, Wu SW, Zhou Y, Sieradzki K, de Lencastre H, Richardson P, et al. Tracking the in vivo evolution of multidrug resistance in Staphylococcus aureus by whole-genome sequencing. Proc Natl Acad Sci USA. 2007;104:9451–6.
Kennedy AD, Otto M, Braughton KR, Whitney AR, Chen L, Mathema B, et al. Epidemic community-associated methicillin-resistant Staphylococcus aureus: recent clonal expansion and diversification. Proc Natl Acad Sci USA. 2008;105:1327–32.
Goering RV, Morrison D, Al-Doori Z, Edwards GF, Gemmell CG. Usefulness of mec-associated direct repeat unit (dru) typing in the epidemiological analysis of highly clonal methicillin-resistant Staphylococcus aureus in Scotland. Clin Microbiol Infect. 2008;14:964–9.
Ellington MJ, Yearwood L, Ganner M, East C, Kearns AM. Distribution of the ACME-arcA gene among methicillin-resistant Staphylococcus aureus from England and Wales. J Antimicrob Chemother. 2008;61:73–7.
Witte W, Strommenger B, Cuny C, Heuck D, Nuebel U. Methicillin-resistant Staphylococcus aureus containing the Panton-Valentine leucocidin gene in Germany in 2005 and 2006. J Antimicrob Chemother. 2007;60:1258–63.
Witte W, Braulke C, Strommenger B. Community-associated methicillin-resistant Staphylococcus aureus ST8 (“USA300”) in an HIV-positive patient in Cologne, Germany, February 2008. Euro Surveill 2008;13:pii:8080.
Larsen AR, Goering R, Stegger M, Lindsay JA, Gould KA, Hinds J, et al. Two distinct clones of methicillin-resistant Staphylococcus aureus (MRSA) with the same USA300 pulsed-field gel electrophoresis profile: a potential pitfall for identification of USA300 community-associated MRSA. J Clin Microbiol. 2009;47:3765–8.
Fenner L, Widmer AF, Dangel M, Frei R. Distribution of spa types among methicillin-resistant Staphylococcus aureus isolates during a 6 year period at a low-prevalence University Hospital. J Med Microbiol. 2008;57:612–6.
Ruppitsch W, Stoger A, Schmid D, Fretz R, Indra A, Allerberger F, et al. Occurrence of the USA300 community-acquired Staphylococcus aureus clone in Austria. Euro Surveill. 2007;12:E071025.1.
Marchese A, Gualco L, Maioli E, Debbia E. Molecular analysis and susceptibility patterns of methicillin-resistant Staphylococcus aureus (MRSA) strains circulating in the community in the Ligurian area, a northern region of Italy: emergence of USA300 and EMRSA-15 clones. Int J Antimicrob Agents. 2009;34:424–8.
Shibuya Y, Hara M, Higuchi W, Takano T, Iwao Y, Yamamoto T. Emergence of the community-acquired methicillin-resistant Staphylococcus aureus USA300 clone in Japan. J Infect Chemother. 2008;14:439–41.
Higuchi W, Mimura S, Kurosawa Y, Takano T, Iwao Y, Yabe S, et al. Emergence of the community-acquired methicillin-resistant Staphylococcus aureus USA300 clone in Japanese child, demonstrating multiple divergent strains in Japan. J Infect Chemother. doi:10.1007/s10156-010-0051-y.
Park C, Lee DG, Kim SW, Choi SM, Park SH, Chun HS, et al. Predominance of community-associated methicillin-resistant Staphylococcus aureus strains carrying staphylococcal chromosome cassette mec type IVA in South Korea. J Clin Microbiol. 2007;45:4021–6.
Wu CT, Lin JJ, Hsia SH. Cutaneous pustular manifestations associated with disseminated septic embolism due to a Panton-Valentine leukocidin-producing strain of community-acquired methicillin-resistant Staphylococcus aureus. Int J Dermatol. 2008;47:942–3.
Monecke S, Ehricht R, Slickers P, Tan HL, Coombs G. The molecular epidemiology and evolution of the Panton-Valentine leukocidin-positive, methicillin-resistant Staphylococcus aureus strain USA300 in Western Australia. Clin Microbiol Infect. 2009;15:770–6.
Arias CA, Rincon S, Chowdhury S, Martinez E, Coronell W, Reyes J, et al. MRSA USA300 clone and VREF—a U.S.-Colombian connection? N Engl J Med. 2008;359:2177–9.
Bekkhoucha SN, Cady A, Gautier P, Itim F, Donnio PY. A portrait of Staphylococcus aureus from the other side of the Mediterranean Sea: molecular characteristics of isolates from Western Algeria. Eur J Clin Microbiol Infect Dis. 2009;28:553–5.
Udo EE, O’Brien FG, Al-Sweih N, Noronha B, Matthew B, Grubb WB. Genetic lineages of community-associated methicillin-resistant Staphylococcus aureus in Kuwait hospitals. J Clin Microbiol. 2008;46:3514–6.
O’Neill AJ, Larsen AR, Henriksen AS, Chopra I. A fusidic acid-resistant epidemic strain of Staphylococcus aureus carries the fusB determinant, whereas fusA mutations are prevalent in other resistant isolates. Antimicrob Agents Chemother. 2004;48:3594–7.
Lannergard J, Norstrom T, Hughes D. Genetic determinants of resistance to fusidic acid among clinical bacteremia isolates of Staphylococcus aureus. Antimicrob Agents Chemother. 2009;53:2059–65.
Udo EE, Sarkhoo E. Genetic Analysis of High-level mupirocin resistance in ST80 clone of community-associated methicillin-resistant Staphylococcus aureus. J Med Microbiol (2009, in press).
Ho PL, Cheung C, Mak GC, Tse CW, Ng TK, Cheung CH, et al. Molecular epidemiology and household transmission of community-associated methicillin-resistant Staphylococcus aureus in Hong Kong. Diagn Microbiol Infect Dis. 2007;57:145–51.
Higuchi W, Takano T, Teng LJ, Yamamoto T. Structure and specific detection of staphylococcal cassette chromosome mec type VII. Biochem Biophys Res Commun. 2008;377:752–6.
Higuchi W, Hung WC, Takano T, Iwao Y, Ozaki K, Isobe H, et al. Molecular characteristics of the Taiwanese multiple drug-resistant ST59 clone of Panton-Valentine leucocidin-positive community-acquired methicillin-resistant Staphylococcus aureus from pediatric cellulitis. J Infect Chemother. doi:10.1007/s10156-010-0029-9.
Baranovich T, Potapov V, Yamamoto T. The first isolation of Panton-Valentine leukocidin (PVL) positive community-acquired methicillin-resistant Staphylococcus aureus (CA-MRSA) in Russia. Euro Surveill. 2007;12:E070315 4.
Otsuka T, Saito K, Dohmae S, Takano T, Higuchi W, Takizawa Y, et al. Key adhesin gene in community-acquired methicillin-resistant Staphylococcus aureus. Biochem Biophys Res Commun. 2006;346:1234–44.
Takizawa Y, Taneike I, Nakagawa S, Oishi T, Nitahara Y, Iwakura N, et al. A Panton-Valentine leucocidin (PVL)-positive community-acquired methicillin-resistant Staphylococcus aureus (MRSA) strain, another such strain carrying a multiple-drug resistance plasmid, and other more-typical PVL-negative MRSA strains found in Japan. J Clin Microbiol. 2005;43:3356–63.
Taneike I, Otsuka T, Dohmae S, Saito K, Ozaki K, Takano M, et al. Molecular nature of methicillin-resistant Staphylococcus aureus derived from explosive nosocomial outbreaks of the 1980 s in Japan. FEBS Lett. 2006;580:2323–34.
Robinson DA, Kearns AM, Holmes A, Morrison D, Grundmann H, Edwards G, et al. Re-emergence of early pandemic Staphylococcus aureus as a community-acquired methicillin-resistant clone. Lancet. 2005;365:1256–8.
Yamamoto T, Yabe S, Shi D, Higuchi W, Iwao Y, Isobe H, et al. Unique adherence characteristics of Panton-Valentine leucocidin-positive ST22 methicillin-resistant Staphylococcus aureus from recurrent and/or multifocal cutaneous abscesses (abstract no. L1-1672). 49th Interscience Conference on Antimicrobial Agents and Chemotherapy. San Francisco, CA: American Scoiety for Microbiology; 2009.
Tristan A, Bes M, Meugnier H, Lina G, Bozdogan B, Courvalin P, et al. Global distribution of Panton-Valentine leukocidin-positive methicillin-resistant Staphylococcus aureus, 2006. Emerg Infect Dis. 2007;13:594–600.
Nimmo GR, Coombs GW. Community-associated methicillin-resistant Staphylococcus aureus (MRSA) in Australia. Int J Antimicrob Agents. 2008;31:401–10.
Degnan BA, Palmer JM, Robson T, Jones CE, Fischer M, Glanville M, et al. Inhibition of human peripheral blood mononuclear cell proliferation by Streptococcus pyogenes cell extract is associated with arginine deiminase activity. Infect Immun. 1998;66:3050–8.
Degnan BA, Fontaine MC, Doebereiner AH, Lee JJ, Mastroeni P, Dougan G, et al. Characterization of an isogenic mutant of Streptococcus pyogenes Manfredo lacking the ability to make streptococcal acid glycoprotein. Infect Immun. 2000;68:2441–8.
Podbielski A, Pohl B, Woischnik M, Korner C, Schmidt KH, Rozdzinski E, et al. Molecular characterization of group A streptococcal (GAS) oligopeptide permease (opp) and its effect on cysteine protease production. Mol Microbiol. 1996;21:1087–99.
Gonzalez BE, Hulten KG, Dishop MK, Lamberth LB, Hammerman WA, Mason EO Jr, et al. Pulmonary manifestations in children with invasive community-acquired Staphylococcus aureus infection. Clin Infect Dis. 2005;41:583–90.
de Bentzmann S, Tristan A, Etienne J, Brousse N, Vandenesch F, Lina G. Staphylococcus aureus isolates associated with necrotizing pneumonia bind to basement membrane type I and IV collagens and laminin. J Infect Dis. 2004;190:1506–15.
Ryden C, Tung HS, Nikolaev V, Engstrom A, Oldberg A. Staphylococcus aureus causing osteomyelitis binds to a nonapeptide sequence in bone sialoprotein. Biochem J. 1997;327:825–9.
Hultenby K, Reinholt FP, Norgard M, Oldberg A, Wendel M, Heinegard D. Distribution and synthesis of bone sialoprotein in metaphyseal bone of young rats show a distinctly different pattern from that of osteopontin. Eur J Cell Biol. 1994;63:230–9.
Tung H, Guss B, Hellman U, Persson L, Rubin K, Ryden C. A bone sialoprotein-binding protein from Staphylococcus aureus: a member of the staphylococcal Sdr family. Biochem J. 2000;345:611–9.
Yabe S, Higuchi W, Iwao Y, Shi D, Takano T, Nishiyama A, et al. Current status of methicillin-resistant Staphylococcus aureus and coagulase-negative staphylococci carriage in medical students and the transmission backgrounds in Japan (abstract no. K-1615). 49th Interscience Conference on Antimicrobial Agents and Chemotherapy. San Francisco, CA: American Scoiety for Microbiology; 2009.
Clarke SR, Foster SJ. Surface adhesins of Staphylococcus aureus. Adv Microb Physiol. 2006;51:187–224.
Rivas JM, Speziale P, Patti JM, Hook M. MSCRAMM—targeted vaccines and immunotherapy for staphylococcal infection. Curr Opin Drug Discov Devel. 2004;7:223–7.
Otto M. Staphylococcal biofilms. Curr Top Microbiol Immunol. 2008;322:207–28.
Tristan A, Benito Y, Montserret R, Boisset S, Dusserre E, Penin F, et al. The signal peptide of Staphylococcus aureus panton valentine leukocidin LukS component mediates increased adhesion to heparan sulfates. PLoS One. 2009;4:e5042.
Grundmeier M, Hussain M, Becker P, Heilmann C, Peters G, Sinha B. Truncation of fibronectin-binding proteins in Staphylococcus aureus strain Newman leads to deficient adherence and host cell invasion due to loss of the cell wall anchor function. Infect Immun. 2004;72:7155–63.
Matic G, Bosch T, Ramlow W. Background and indications for protein A-based extracorporeal immunoadsorption. Ther Apher. 2001;5:394–403.
Kahl BC, Peters G. Microbiology. Mayhem in the lung. Science. 2007;315:1082–3.
Labandeira-Rey M, Couzon F, Boisset S, Brown EL, Bes M, Benito Y, et al. Staphylococcus aureus Panton-Valentine leukocidin causes necrotizing pneumonia. Science. 2007;315:1130–3.
Gomez MI, O’Seaghdha M, Magargee M, Foster TJ, Prince AS. Staphylococcus aureus protein A activates TNFR1 signaling through conserved IgG binding domains. J Biol Chem. 2006;281:20190–6.
Novick RP. Mobile genetic elements and bacterial toxinoses: the superantigen-encoding pathogenicity islands of Staphylococcus aureus. Plasmid. 2003;49:93–105.
Vojtov N, Ross HF, Novick RP. Global repression of exotoxin synthesis by staphylococcal superantigens. Proc Natl Acad Sci USA. 2002;99:10102–7.
Jarraud S, Peyrat MA, Lim A, Tristan A, Bes M, Mougel C, et al. egc, a highly prevalent operon of enterotoxin gene, forms a putative nursery of superantigens in Staphylococcus aureus. J Immunol. 2001;166:669–77.
Sundstedt A, Grundstrom S, Dohlsten M. T cell- and perforin-dependent depletion of B cells in vivo by staphylococcal enterotoxin A. Immunology. 1998;95:76–82.
Lussow AR, MacDonald HR. Differential effects of superantigen-induced “anergy” on priming and effector stages of a T cell-dependent antibody response. Eur J Immunol. 1994;24:445–9.
Hu HL, Cornwell WD, Rogers TJ, Lin YS. In vivo analysis of a superantigen-induced T cell suppressor factor. Cell Immunol. 1996;167:285–92.
Ferry T, Thomas D, Genestier AL, Bes M, Lina G, Vandenesch F, et al. Comparative prevalence of superantigen genes in Staphylococcus aureus isolates causing sepsis with and without septic shock. Clin Infect Dis. 2005;41:771–7.
van Belkum A, Melles DC, Snijders SV, van Leeuwen WB, Wertheim HF, Nouwen JL, et al. Clonal distribution and differential occurrence of the enterotoxin gene cluster, egc, in carriage- versus bacteremia-associated isolates of Staphylococcus aureus. J Clin Microbiol. 2006;44:1555–7.
Kaneko J, Kamio Y. Bacterial two-component and hetero-heptameric pore-forming cytolytic toxins: structures, pore-forming mechanism, and organization of the genes. Biosci Biotechnol Biochem. 2004;68:981–1003.
Dinges MM, Orwin PM, Schlievert PM. Exotoxins of Staphylococcus aureus. Clin Microbiol Rev. 2000;13:16–34.
Bohach GA. Staphylococcus aureus exotoxins. In: Fischetti VA, Novick RP, Ferretti JJ, Portnoy DA, Rood JI, editors. Gram-positive pathogens. 2nd ed. Washington, DC: American Society for Microbiology; 2006. p. 464–77.
Verdon J, Girardin N, Lacombe C, Berjeaud JM, Hechard Y. delta-hemolysin, an update on a membrane-interacting peptide. Peptides. 2009;30:817–23.
Barrio MB, Rainard P, Prevost G. LukM/LukF’-PV is the most active Staphylococcus aureus leukotoxin on bovine neutrophils. Microbes Infect. 2006;8:2068–74.
Wang R, Braughton KR, Kretschmer D, Bach TH, Queck SY, Li M, et al. Identification of novel cytolytic peptides as key virulence determinants for community-associated MRSA. Nat Med. 2007;13:1510–4.
Nygaard TK, DeLeo FR, Voyich JM. Community-associated methicillin-resistant Staphylococcus aureus skin infections: advances toward identifying the key virulence factors. Curr Opin Infect Dis. 2008;21:147–52.
Queck SY, Khan BA, Wang R, Bach TH, Kretschmer D, Chen L, et al. Mobile genetic element-encoded cytolysin connects virulence to methicillin resistance in MRSA. PLoS Pathog. 2009;5:e1000533.
Kalyan S, Chow AW. Human peripheral gammadelta T cells potentiate the early proinflammatory cytokine response to staphylococcal toxic shock syndrome toxin-1. J Infect Dis. 2004;189:1892–6.
Kappler J, Kotzin B, Herron L, Gelfand EW, Bigler RD, Boylston A, et al. V beta-specific stimulation of human T cells by staphylococcal toxins. Science. 1989;244:811–3.
Krakauer T. Immune response to staphylococcal superantigens. Immunol Res. 1999;20:163–73.
Li H, Llera A, Mariuzza RA. Structure-function studies of T-cell receptor-superantigen interactions. Immunol Rev. 1998;163:177–86.
Fraser JD, Proft T. The bacterial superantigen and superantigen-like proteins. Immunol Rev. 2008;225:226–43.
Ono HK, Omoe K, Imanishi K, Iwakabe Y, Hu DL, Kato H, et al. Identification and characterization of two novel staphylococcal enterotoxins, types S and T. Infect Immun. 2008;76:4999–5005.
Takizawa Y, Shimizu T, Iwaya A, Hatakeyama K, Taneike I, Nakagawa S, et al. A multiple superantigenic toxin pattern of methicillin-resistant Staphylococcus aureus (MRSA) as a risk factor in the development of toxic shock syndrome (TSS). Acta Med Biol. 2003;51:141–7.
Yarwood JM, McCormick JK, Paustian ML, Orwin PM, Kapur V, Schlievert PM. Characterization and expression analysis of Staphylococcus aureus pathogenicity island 3. Implications for the evolution of staphylococcal pathogenicity islands. J Biol Chem. 2002;277:13138–47.
Chen J, Novick RP. Phage-mediated intergeneric transfer of toxin genes. Science. 2009;323:139–41.
Mertz PM, Cardenas TC, Snyder RV, Kinney MA, Davis SC, Plano LR. Staphylococcus aureus virulence factors associated with infected skin lesions: influence on the local immune response. Arch Dermatol. 2007;143:1259–63.
Nishifuji K, Sugai M, Amagai M. Staphylococcal exfoliative toxins: “molecular scissors” of bacteria that attack the cutaneous defense barrier in mammals. J Dermatol Sci. 2008;49:21–31.
Yamaguchi T, Hayashi T, Takami H, Nakasone K, Ohnishi M, Nakayama K, et al. Phage conversion of exfoliative toxin A production in Staphylococcus aureus. Mol Microbiol. 2000;38:694–705.
Yamaguchi T, Hayashi T, Takami H, Ohnishi M, Murata T, Nakayama K, et al. Complete nucleotide sequence of a Staphylococcus aureus exfoliative toxin B plasmid and identification of a novel ADP-ribosyltransferase, EDIN-C. Infect Immun. 2001;69:7760–71.
Yamaguchi T, Yokota Y, Terajima J, Hayashi T, Aepfelbacher M, Ohara M, et al. Clonal association of Staphylococcus aureus causing bullous impetigo and the emergence of new methicillin-resistant clonal groups in Kansai district in Japan. J Infect Dis. 2002;185:1511–6.
Genestier AL, Michallet MC, Prevost G, Bellot G, Chalabreysse L, Peyrol S, et al. Staphylococcus aureus Panton-Valentine leukocidin directly targets mitochondria and induces Bax-independent apoptosis of human neutrophils. J Clin Invest. 2005;115:3117–27.
Pedelacq JD, Maveyraud L, Prevost G, Baba-Moussa L, Gonzalez A, Courcelle E, et al. The structure of a Staphylococcus aureus leucocidin component (LukF-PV) reveals the fold of the water-soluble species of a family of transmembrane pore-forming toxins. Structure. 1999;7:277–87.
Guillet V, Roblin P, Werner S, Coraiola M, Menestrina G, Monteil H, et al. Crystal structure of leucotoxin S component: new insight into the staphylococcal beta-barrel pore-forming toxins. J Biol Chem. 2004;279:41028–37.
Berman H, Henrick K, Nakamura H. Announcing the worldwide Protein Data Bank. Nat Struct Biol. 2003;10:980.
Guex N, Peitsch MC. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis. 1997;18:2714–23.
O’Hara FP, Guex N, Word JM, Miller LA, Becker JA, Walsh SL, et al. A geographic variant of the Staphylococcus aureus Panton-Valentine leukocidin toxin and the origin of community-associated methicillin-resistant S. aureus USA300. J Infect Dis. 2008;197:187–94.
Wolter DJ, Tenover FC, Goering RV. Allelic variation in genes encoding Panton-Valentine leukocidin from community-associated Staphylococcus aureus. Clin Microbiol Infect. 2007;13:827–30.
Noda M, Kato I, Matsuda F, Hirayama T. Mode of action of staphylococcal leukocidin: relationship between binding of 125I-labeled S and F components of leukocidin to rabbit polymorphonuclear leukocytes and leukocidin activity. Infect Immun. 1981;34:362–7.
Sugawara-Tomita N, Tomita T, Kamio Y. Stochastic assembly of two-component staphylococcal gamma-hemolysin into heteroheptameric transmembrane pores with alternate subunit arrangements in ratios of 3:4 and 4:3. J Bacteriol. 2002;184:4747–56.
Miles G, Movileanu L, Bayley H. Subunit composition of a bicomponent toxin: staphylococcal leukocidin forms an octameric transmembrane pore. Protein Sci. 2002;11:894–902.
Ward PD, Turner WH. Identification of staphylococcal Panton-Valentine leukocidin as a potent dermonecrotic toxin. Infect Immun. 1980;28:393–7.
Brown EL, Dumitrescu O, Thomas D, Badiou C, Koers EM, Choudhury P, et al. The Panton-Valentine leukocidin vaccine protects mice against lung and skin infections caused by Staphylococcus aureus USA300. Clin Microbiol Infect. 2009;15:156–64.
Voyich JM, Otto M, Mathema B, Braughton KR, Whitney AR, Welty D, et al. Is Panton-Valentine leukocidin the major virulence determinant in community-associated methicillin-resistant Staphylococcus aureus disease? J Infect Dis. 2006;194:1761–70.
Bubeck Wardenburg J, Bae T, Otto M, Deleo FR, Schneewind O. Poring over pores: alpha-hemolysin and Panton-Valentine leukocidin in Staphylococcus aureus pneumonia. Nat Med. 2007;13:1405–6.
Bubeck Wardenburg J, Palazzolo-Ballance AM, Otto M, Schneewind O, DeLeo FR. Panton-Valentine leukocidin is not a virulence determinant in murine models of community-associated methicillin-resistant Staphylococcus aureus disease. J Infect Dis. 2008;198:1166–70.
Orii KO, Iwao Y, Higuchi W, Takano T, Yamamoto T. Molecular characterization of methicillin-resistant Staphylococcus aureus from a fatal case of necrotizing fasciitis in an extremely low-birth-weight infant. Clin Microbiol Infect. 2010;16:289–92.
Diep BA, Palazzolo-Ballance AM, Tattevin P, Basuino L, Braughton KR, Whitney AR, et al. Contribution of Panton-Valentine leukocidin in community-associated methicillin-resistant Staphylococcus aureus pathogenesis. PLoS One. 2008;3:e3198.
Dumitrescu O, Badiou C, Bes M, Reverdy ME, Vandenesch F, Etienne J, et al. Effect of antibiotics, alone and in combination, on Panton-Valentine leukocidin production by a Staphylococcus aureus reference strain. Clin Microbiol Infect. 2008;14:384–8.
Stevens DL, Ma Y, Salmi DB, McIndoo E, Wallace RJ, Bryant AE. Impact of antibiotics on expression of virulence-associated exotoxin genes in methicillin-sensitive and methicillin-resistant Staphylococcus aureus. J Infect Dis. 2007;195:202–11.
Nakagawa S, Taneike I, Mimura D, Iwakura N, Nakayama T, Emura T, et al. Gene sequences and specific detection for Panton-Valentine leukocidin. Biochem Biophys Res Commun. 2005;328:995–1002.
Prevost G, Cribier B, Couppie P, Petiau P, Supersac G, Finck-Barbancon V, et al. Panton-Valentine leucocidin and gamma-hemolysin from Staphylococcus aureus ATCC 49775 are encoded by distinct genetic loci and have different biological activities. Infect Immun. 1995;63:4121–9.
Hartman BJ, Tomasz A. Low-affinity penicillin-binding protein associated with beta-lactam resistance in Staphylococcus aureus. J Bacteriol. 1984;158:513–6.
Utsui Y, Yokota T. Role of an altered penicillin-binding protein in methicillin- and cephem-resistant Staphylococcus aureus. Antimicrob Agents Chemother. 1985;28:397–403.
Hussain M, Pastor FI, Lampen JO. Cloning sequencing of the blaZ gene encoding beta-lactamase III, a lipoprotein of Bacillus cereus 569/H. J Bacteriol. 1987;169:579–86.
Monecke S, Ehricht R. Rapid genotyping of methicillin-resistant Staphylococcus aureus (MRSA) isolates using miniaturised oligonucleotide arrays. Clin Microbiol Infect. 2005;11:825–33.
Bugg TD, Wright GD, Dutka-Malen S, Arthur M, Courvalin P, Walsh CT. Molecular basis for vancomycin resistance in Enterococcus faecium BM4147: biosynthesis of a depsipeptide peptidoglycan precursor by vancomycin resistance proteins VanH and VanA. Biochemistry. 1991;30:10408–15.
Bugg TD, Dutka-Malen S, Arthur M, Courvalin P, Walsh CT. Identification of vancomycin resistance protein VanA as a d-alanine:d-alanine ligase of altered substrate specificity. Biochemistry. 1991;30:2017–21.
Dutka-Malen S, Molinas C, Arthur M, Courvalin P. The VANA glycopeptide resistance protein is related to d-alanyl-d-alanine ligase cell wall biosynthesis enzymes. Mol Gen Genet. 1990;224:364–72.
From the Centers for Disease Control and Prevention. Staphylococcus aureus resistant to vancomycin—United States, 2002. 2002;51:565–7.
Shaw KJ, Poppe S, Schaadt R, Brown-Driver V, Finn J, Pillar CM, et al. In vitro activity of TR-700, the antibacterial moiety of the prodrug TR-701, against linezolid-resistant strains. Antimicrob Agents Chemother. 2008;52:4442–7.
Besier S, Ludwig A, Zander J, Brade V, Wichelhaus TA. Linezolid resistance in Staphylococcus aureus: gene dosage effect, stability, fitness costs, and cross-resistances. Antimicrob Agents Chemother. 2008;52:1570–2.
Ardic N, Sareyyupoglu B, Ozyurt M, Haznedaroglu T, Ilga U. Investigation of aminoglycoside modifying enzyme genes in methicillin-resistant staphylococci. Microbiol Res. 2006;161:49–54.
Ida T, Okamoto R, Shimauchi C, Okubo T, Kuga A, Inoue M. Identification of aminoglycoside-modifying enzymes by susceptibility testing: epidemiology of methicillin-resistant Staphylococcus aureus in Japan. J Clin Microbiol. 2001;39:3115–21.
Ubukata K, Yamashita N, Gotoh A, Konno M. Purification and characterization of aminoglycoside-modifying enzymes from Staphylococcus aureus and Staphylococcus epidermidis. Antimicrob Agents Chemother. 1984;25:754–9.
Byrne ME, Gillespie MT, Skurray RA. 4’, 4’’ adenyltransferase activity on conjugative plasmids isolated from Staphylococcus aureus is encoded on an integrated copy of pUB110. Plasmid. 1991;25:70–5.
Matsumura M, Katakura Y, Imanaka T, Aiba S. Enzymatic and nucleotide sequence studies of a kanamycin-inactivating enzyme encoded by a plasmid from thermophilic bacilli in comparison with that encoded by plasmid pUB110. J Bacteriol. 1984;160:413–20.
Pinto-Alphandary H, Mabilat C, Courvalin P. Emergence of aminoglycoside resistance genes aadA and aadE in the genus Campylobacter. Antimicrob Agents Chemother. 1990;34:1294–6.
Gray GS, Fitch WM. Evolution of antibiotic resistance genes: the DNA sequence of a kanamycin resistance gene from Staphylococcus aureus. Mol Biol Evol. 1983;1:57–66.
Clark NC, Olsvik O, Swenson JM, Spiegel CA, Tenover FC. Detection of a streptomycin/spectinomycin adenylyltransferase gene (aadA) in Enterococcus faecalis. Antimicrob Agents Chemother. 1999;43:157–60.
Ferrero L, Cameron B, Manse B, Lagneaux D, Crouzet J, Famechon A, et al. Cloning and primary structure of Staphylococcus aureus DNA topoisomerase IV: a primary target of fluoroquinolones. Mol Microbiol. 1994;13:641–53.
Hopewell R, Oram M, Briesewitz R, Fisher LM. DNA cloning and organization of the Staphylococcus aureus gyrA and gyrB genes: close homology among gyrase proteins and implications for 4-quinolone action and resistance. J Bacteriol. 1990;172:3481–4.
Jin DJ, Gross CA. Mapping and sequencing of mutations in the Escherichia coli rpoB gene that lead to rifampicin resistance. J Mol Biol. 1988;202:45–58.
Ovchinnikov YA, Monastyrskaya GS, Guriev SO, Kalinina NF, Sverdlov ED, Gragerov AI, et al. RNA polymerase rifampicin resistance mutations in Escherichia coli: sequence changes and dominance. Mol Gen Genet. 1983;190:344–8.
O’Neill AJ, Huovinen T, Fishwick CW, Chopra I. Molecular genetic and structural modeling studies of Staphylococcus aureus RNA polymerase and the fitness of rifampin resistance genotypes in relation to clinical prevalence. Antimicrob Agents Chemother. 2006;50:298–309.
Murphy E. Nucleotide sequence of ermA, a macrolide-lincosamide-streptogramin B determinant in Staphylococcus aureus. J Bacteriol. 1985;162:633–40.
Otsuka T, Zaraket H, Takano T, Saito K, Dohmae S, Higuchi W, et al. Macrolide-lincosamide-streptogramin B resistance phenotypes and genotypes among Staphylococcus aureus clinical isolates in Japan. Clin Microbiol Infect. 2007;13:325–7.
Schmitz FJ, Sadurski R, Kray A, Boos M, Geisel R, Kohrer K, et al. Prevalence of macrolide-resistance genes in Staphylococcus aureus and Enterococcus faecium isolates from 24 European university hospitals. J Antimicrob Chemother. 2000;45:891–4.
Horinouchi S, Weisblum B. Posttranscriptional modification of mRNA conformation: mechanism that regulates erythromycin-induced resistance. Proc Natl Acad Sci USA. 1980;77:7079–83.
Horinouchi S, Byeon WH, Weisblum B. A complex attenuator regulates inducible resistance to macrolides, lincosamides, and streptogramin type B antibiotics in Streptococcus sanguis. J Bacteriol. 1983;154:1252–62.
Qi W, Ender M, O’Brien F, Imhof A, Ruef C, McCallum N, et al. Molecular epidemiology of methicillin-resistant Staphylococcus aureus in Zurich, Switzerland (2003): prevalence of type IV SCCmec and a new SCCmec element associated with isolates from intravenous drug users. J Clin Microbiol. 2005;43:5164–70.
Farmer TH, Gilbart J, Elson SW. Biochemical basis of mupirocin resistance in strains of Staphylococcus aureus. J Antimicrob Chemother. 1992;30:587–96.
Simor AE, Stuart TL, Louie L, Watt C, Ofner-Agostini M, Gravel D, et al. Mupirocin-resistant, methicillin-resistant Staphylococcus aureus strains in Canadian hospitals. Antimicrob Agents Chemother. 2007;51:3880–6.
O’Brien FG, Price C, Grubb WB, Gustafson JE. Genetic characterization of the fusidic acid and cadmium resistance determinants of Staphylococcus aureus plasmid pUB101. J Antimicrob Chemother. 2002;50:313–21.
O’Neill AJ, Chopra I. Molecular basis of fusB-mediated resistance to fusidic acid in Staphylococcus aureus. Mol Microbiol. 2006;59:664–76.
Witte W, Braulke C, Cuny C, Strommenger B, Werner G, Heuck D, et al. Emergence of methicillin-resistant Staphylococcus aureus with Panton-Valentine leukocidin genes in central Europe. Eur J Clin Microbiol Infect Dis. 2005;24:1–5.
Ozaki K, Takano M, Higuchi W, Takano T, Yabe S, Nitahara Y, et al. Genotypes, intrafamilial transmission, and virulence potential of nasal methicillin-resistant Staphylococcus aureus from children in the community. J Infect Chemother. 2009;15:84–91.
Jamaluddin TZ, Kuwahara-Arai K, Hisata K, Terasawa M, Cui L, Baba T, et al. Extreme genetic diversity of methicillin-resistant Staphylococcus epidermidis strains disseminated among healthy Japanese children. J Clin Microbiol. 2008;46:3778–83.
Hisata K, Kuwahara-Arai K, Yamanoto M, Ito T, Nakatomi Y, Cui L, et al. Dissemination of methicillin-resistant staphylococci among healthy Japanese children. J Clin Microbiol. 2005;43:3364–72.
Zaraket H, Otsuka T, Saito K, Dohmae S, Takano T, Higuchi W, et al. Molecular characterization of methicillin-resistant Staphylococcus aureus in hospitals in Niigata, Japan: divergence and transmission. Microbiol Immunol. 2007;51:171–6.
Matsumoto T, Yamamoto T, Ishikawa H, Koshihara K, Matsunaga N, Ishii Y, et al. An outbreak of community-acquired Methicillin-resistant Staphylococcus aureus Infection in a sumo wrestling team (abstract no. L-1499). In: 48th Interscience conference on antimicrobial agents and chemotherapy. Washington, DC. 2008.
Yano M, Ishii R, Yoshida M, Chiyonobu T, Nukina S, Otsuki Y. A case of extradural abscess due to community-acquired MRSA (in Japanese). In: 40th annual meeting of the Japanese society for pediatric infectious diseases; 2008. p. D-5.
Tomita Y, Kawano O, Ichiyasu H, Fukushima T, Fukuda K, Sugimoto M, et al. Two cases of severe necrotizing pneumonia caused by community-acquired methicillin-resistant Staphylococcus aureus (in Japanese). Nihon Kokyuki Gakkai Zasshi. 2008;46:395–403.
Shi D, Higuchi W, Saito K, Yabe S, Takano T, Nishiyama A, et al. Two types of methicillin-resistant Staphylococcus aureus isolated alone or in combination with methicillin-susceptible S. aureus from bullous impetigo in Japan (abstract no. C2-121). In: 49th interscience conference on antimicrobial agents and chemotherapy. San Francisco, CA: American Society for Microbiology; 2009.
Nakaminami H, Noguchi N, Ikeda M, Hasui M, Sato M, Yamamoto S, et al. Molecular epidemiology and antimicrobial susceptibilities of 273 exfoliative toxin-encoding-gene-positive Staphylococcus aureus isolates from patients with impetigo in Japan. J Med Microbiol. 2008;57:1251–8.
Huang YC, Ho CF, Chen CJ, Su LH, Lin TY. Comparative molecular analysis of community-associated and healthcare-associated methicillin-resistant Staphylococcus aureus isolates from children in northern Taiwan. Clin Microbiol Infect. 2008;14:1167–72.
Wang JT, Liao CH, Fang CT, Chie WC, Lai MS, Lauderdale TL, et al. Prevalence of and risk factors for colonization by methicillin-resistant Staphylococcus aureus among adults in community settings in Taiwan. J Clin Microbiol. 2009;47:2957–63.
Yu F, Chen Z, Liu C, Zhang X, Lin X, Chi S, et al. Prevalence of Staphylococcus aureus carrying Panton-Valentine leukocidin genes among isolates from hospitalised patients in China. Clin Microbiol Infect. 2008;14:381–4.
Ho PL, Tse CW, Mak GC, Chow KH, Ng TK. Community-acquired methicillin-resistant Staphylococcus aureus arrives in Hong Kong. J Antimicrob Chemother. 2004;54:845–6.
Ho PL, Chuang SK, Choi YF, Lee RA, Lit AC, Ng TK, et al. Community-associated methicillin-resistant and methicillin-sensitive Staphylococcus aureus: skin and soft tissue infections in Hong Kong. Diagn Microbiol Infect Dis. 2008;61:245–50.
Liu Y, Wang H, Du N, Shen E, Chen H, Niu J, et al. Molecular evidence for spread of two major methicillin-resistant Staphylococcus aureus clones with a unique geographic distribution in Chinese hospitals. Antimicrob Agents Chemother. 2009;53:512–8.
Vorobieva V, Bazhukova T, Hanssen AM, Caugant DA, Semenova N, Haldorsen BC, et al. Clinical isolates of Staphylococcus aureus from the Arkhangelsk region, Russia: antimicrobial susceptibility, molecular epidemiology, and distribution of Panton-Valentine leukocidin genes. APMIS. 2008;116:877–87.
Park SH, Park C, Yoo JH, Choi SM, Choi JH, Shin HH, et al. Emergence of community-associated methicillin-resistant Staphylococcus aureus strains as a cause of healthcare-associated bloodstream infections in Korea. Infect Control Hosp Epidemiol. 2009;30:146–55.
Juhasz-Kaszanyitzky E, Janosi S, Somogyi P, Dan A, van der Graaf-van Bloois L, van Duijkeren E, et al. MRSA transmission between cows and humans. Emerg Infect Dis. 2007;13:630–2.
Voss A, Loeffen F, Bakker J, Klaassen C, Wulf M. Methicillin-resistant Staphylococcus aureus in pig farming. Emerg Infect Dis. 2005;11:1965–6.
Huijsdens XW, van Dijke BJ, Spalburg E, van Santen-Verheuvel MG, Heck ME, Pluister GN, et al. Community-acquired MRSA and pig-farming. Ann Clin Microbiol Antimicrob. 2006;5:26.
van Duijkeren E, Jansen MD, Flemming SC, de Neeling H, Wagenaar JA, Schoormans AH, et al. Methicillin-resistant Staphylococcus aureus in pigs with exudative epidermitis. Emerg Infect Dis. 2007;13:1408–10.
van Duijkeren E, Wolfhagen MJ, Heck ME, Wannet WJ. Transmission of a Panton-Valentine leucocidin-positive, methicillin-resistant Staphylococcus aureus strain between humans and a dog. J Clin Microbiol. 2005;43:6209–11.
van Duijkeren E, Wolfhagen MJ, Box AT, Heck ME, Wannet WJ, Fluit AC. Human-to-dog transmission of methicillin-resistant Staphylococcus aureus. Emerg Infect Dis. 2004;10:2235–7.
Nienhoff U, Kadlec K, Chaberny IF, Verspohl J, Gerlach GF, Schwarz S, et al. Transmission of methicillin-resistant Staphylococcus aureus strains between humans and dogs: two case reports. J Antimicrob Chemother. 2009;64:660–2.
Sing A, Tuschak C, Hormansdorfer S. Methicillin-resistant Staphylococcus aureus in a family and its pet cat. N Engl J Med. 2008;358:1200–1.
Weese JS, Dick H, Willey BM, McGeer A, Kreiswirth BN, Innis B, et al. Suspected transmission of methicillin-resistant Staphylococcus aureus between domestic pets and humans in veterinary clinics and in the household. Vet Microbiol. 2006;115:148–55.
Vitale CB, Gross TL, Weese JS. Methicillin-resistant Staphylococcus aureus in cat and owner. Emerg Infect Dis. 2006;12:1998–2000.
Weese JS, Caldwell F, Willey BM, Kreiswirth BN, McGeer A, Rousseau J, et al. An outbreak of methicillin-resistant Staphylococcus aureus skin infections resulting from horse to human transmission in a veterinary hospital. Vet Microbiol. 2006;114:160–4.
Wannet WJ, Spalburg E, Heck ME, Pluister GN, Tiemersma E, Willems RJ, et al. Emergence of virulent methicillin-resistant Staphylococcus aureus strains carrying Panton-Valentine leucocidin genes in The Netherlands. J Clin Microbiol. 2005;43:3341–5.
Rich M, Roberts L. Methicillin-resistant Staphylococcus aureus isolates from companion animals. Vet Rec. 2004;154:310.
Baptiste KE, Williams K, Willams NJ, Wattret A, Clegg PD, Dawson S, et al. Methicillin-resistant staphylococci in companion animals. Emerg Infect Dis. 2005;11:1942–4.
Loeffler A, Boag AK, Sung J, Lindsay JA, Guardabassi L, Dalsgaard A, et al. Prevalence of methicillin-resistant Staphylococcus aureus among staff and pets in a small animal referral hospital in the UK. J Antimicrob Chemother. 2005;56:692–7.
Leonard FC, Abbott Y, Rossney A, Quinn PJ, O’Mahony R, Markey BK. Methicillin-resistant Staphylococcus aureus isolated from a veterinary surgeon and five dogs in one practice. Vet Rec. 2006;158:155–9.
Moodley A, Stegger M, Bagcigil AF, Baptiste KE, Loeffler A, Lloyd DH, et al. spa typing of methicillin-resistant Staphylococcus aureus isolated from domestic animals and veterinary staff in the UK and Ireland. J Antimicrob Chemother. 2006;58:1118–23.
Strommenger B, Kehrenberg C, Kettlitz C, Cuny C, Verspohl J, Witte W, et al. Molecular characterization of methicillin-resistant Staphylococcus aureus strains from pet animals and their relationship to human isolates. J Antimicrob Chemother. 2006;57:461–5.
Hanselman BA, Kruth S, Weese JS. Methicillin-resistant staphylococcal colonization in dogs entering a veterinary teaching hospital. Vet Microbiol. 2008;126:277–81.
Kwon NH, Park KT, Moon JS, Jung WK, Kim SH, Kim JM, et al. Staphylococcal cassette chromosome mec (SCCmec) characterization and molecular analysis for methicillin-resistant Staphylococcus aureus and novel SCCmec subtype IVg isolated from bovine milk in Korea. J Antimicrob Chemother. 2005;56:624–32.
Nemati M, Hermans K, Lipinska U, Denis O, Deplano A, Struelens M, et al. Antimicrobial resistance of old and recent Staphylococcus aureus isolates from poultry: first detection of livestock-associated methicillin-resistant strain ST398. Antimicrob Agents Chemother. 2008;52:3817–9.
Cuny C, Nathaus R, Layer F, Strommenger B, Altmann D, Witte W. Nasal colonization of humans with methicillin-resistant Staphylococcus aureus (MRSA) CC398 with and without exposure to pigs. PLoS One. 2009;4:e6800.
Smith TC, Male MJ, Harper AL, Kroeger JS, Tinkler GP, Moritz ED, et al. Methicillin-resistant Staphylococcus aureus (MRSA) strain ST398 is present in midwestern U.S. swine and swine workers. PLoS One. 2009;4:e4258.
Springer B, Orendi U, Much P, Hoger G, Ruppitsch W, Krziwanek K, et al. Methicillin-resistant Staphylococcus aureus: a new zoonotic agent? Wien Klin Wochenschr. 2009;121:86–90.
van Rijen MM, Van Keulen PH, Kluytmans JA. Increase in a Dutch hospital of methicillin-resistant Staphylococcus aureus related to animal farming. Clin Infect Dis. 2008;46:261–3.
Monecke S, Kuhnert P, Hotzel H, Slickers P, Ehricht R. Microarray based study on virulence-associated genes and resistance determinants of Staphylococcus aureus isolates from cattle. Vet Microbiol. 2007;125:128–40.
Loeffler A, Kearns AM, Ellington MJ, Smith LJ, Unt VE, Lindsay JA, et al. First isolation of MRSA ST398 from UK animals: a new challenge for infection control teams? J Hosp Infect. 2009;72:269–71.
Bens CC, Voss A, Klaassen CH. Presence of a novel DNA methylation enzyme in methicillin-resistant Staphylococcus aureus isolates associated with pig farming leads to uninterpretable results in standard pulsed-field gel electrophoresis analysis. J Clin Microbiol. 2006;44:1875–6.
Cui S, Li J, Hu C, Jin S, Li F, Guo Y, et al. Isolation and characterization of methicillin-resistant Staphylococcus aureus from swine and workers in China. J Antimicrob Chemother. 2009;64:680–3.
Neela V, Mohd Zafrul A, Mariana NS, van Belkum A, Liew YK, Rad EG. Prevalence of ST9 methicillin-resistant Staphylococcus aureus among pigs and pig handlers in Malaysia. J Clin Microbiol. 2009;47:4138–40.
Kitai S, Shimizu A, Kawano J, Sato E, Nakano C, Uji T, et al. Characterization of methicillin-resistant Staphylococcus aureus isolated from retail raw chicken meat in Japan. J Vet Med Sci. 2005;67:107–10.
Vazquez-Laslop N, Thum C, Mankin AS. Molecular mechanism of drug-dependent ribosome stalling. Mol Cell. 2008;30:190–202.
Clinical and Laboratory Standards Institute. Performance standard for antimicrobial susceptibility testing; 18th informational supplement, vol. 28, no. 1, M100-S18. Wayne, PA:. Clinical and Laboratory Standards Institute; 2008.
Acknowledgments
We thank Tsuneyuki Nakamura, Yoshiko Mine, Yasuhisa Iwao, Tatiana Baranovich, and Shima Enany for helpful discussion and assistance. This work was supported, in part, by grants from Interchange Association, Japan; from the Ministry of Education, Science, Sports and Culture, Japan; and from Tsukada Foundation (Niigata), Japan.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Yamamoto, T., Nishiyama, A., Takano, T. et al. Community-acquired methicillin-resistant Staphylococcus aureus: community transmission, pathogenesis, and drug resistance. J Infect Chemother 16, 225–254 (2010). https://doi.org/10.1007/s10156-010-0045-9
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10156-010-0045-9