Abstract
Stem blister canker, caused by Botryosphaeria dothidea, is becoming the most serious disease of poplar in China. The molecular basis of the poplar in response to stem blister canker is not well understood. To reveal the global transcriptional changes of poplar to infection by B. dothidea, Solexa paired-end sequencing of complementary DNAs (cDNAs) from control (NB) and pathogen-treated samples (WB) was performed, resulting in a total of 339,283 transcripts and 183,881 unigenes. A total of 206,586 transcripts were differentially expressed in response to pathogen stress (false discovery rate ≤0.05 and an absolute value of log2Ratio (NB/WB) ≥1). In enrichment analysis, energy metabolism and redox reaction-related macromolecules were accumulated significantly in Gene Ontology and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways analyses, indicating components of dynamic defense against the fungus. A total of 852 transcripts (575 upregulated and 277 downregulated transcripts) potentially involved in plant–pathogen interaction were also differentially regulated, including genes encoding proteins linked to signal transduction (putative leucine-rich repeat (LRR) protein kinases and calcium-binding proteins), defense (pathogenesis-related protein 1), and cofactors (jasmonate-ZIM-domain-containing proteins and heat shock proteins). Moreover, transcripts encoding glutathione S-transferase (GST) were accumulated to high levels, revealing key genes and proteins potentially related to pathogen resistance. Poplar RNA sequence data were validated by quantitative real-time PCR (RT-qPCR), which revealed a highly reliability of the transcriptomic profiling data.






Similar content being viewed by others
References
Adhikari BN, Savory EA, Vaillancourt B, Childs KL, Hamilton JP, Day B, Buell CR (2012) Expression profiling of Cucumis sativus in response to infection by Pseudoperonospora cubensis. PLoS One 7(4):e34954
Alexander D, Goodman RM, Gut-Rella M, Glascock C, Weymann K, Friedrich L, Maddox D, Ahl-Goy P, Luntz T, Ward E (1993) Increased tolerance to two oomycete pathogens in transgenic tobacco expressing pathogenesis-related protein 1A. Proc Natl Acad Sci U S A 90(15):7327–7331
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Asai T, Tena G, Plotnikova J, Willmann MR, Chiu W-L, Gomez-Gomez L, Boller T, Ausubel FM, Sheen J (2002) MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415(6875):977–983
Azaiez A, Boyle B, Levée V, Séguin A (2009) Transcriptome profiling in hybrid poplar following interactions with Melampsora rust fungi. Mol Plant Microbe Interact 22(2):190–200
Babashpour S, Aminzadeh S, Farrokhi N, Karkhane A, Haghbeen K (2012) Characterization of a chitinase (Chit62) from Serratia marcescens B4A and its efficacy as a bioshield against plant fungal pathogens. Biochem Genet 50(9–10):722–735
Bai S, Dong C, Li B, Dai H (2012) A PR-4 gene identified from Malusdomestica is involved in the defense responses against Botryosphaeria dothidea. Plant Physiol Biochem 62:23–32
Brown-Rytlewski DE, McManus PS (2000) Virulence of Botryosphaeria dothidea and Botryosphaeria obtusa on apple and management of stem cankers with fungicides. Plant Dis 84(9):1031–1037
Ceserani T, Trofka A, Gandotra N, Nelson T (2009) VH1/BRL2 receptor-like kinase interacts with vascular-specific adaptor proteins VIT and VIK to influence leaf venation. Plant J 57(6):1000–1014
Chain F, Côté-Beaulieu C, Belzile F, Menzies J, Bélanger R (2009) A comprehensive transcriptomic analysis of the effect of silicon on wheat plants under control and pathogen stress conditions. Mol Plant Microbe Interact 22(11):1323–1330
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11(2):113–116
Chen J-H, Jiang H-W, Hsieh E-J, Chen H-Y, Chien C-T, Hsieh H-L, Lin T-P (2012a) Drought and salt stress tolerance of an Arabidopsis glutathione S-transferase U17 knockout mutant are attributed to the combined effect of glutathione and abscisic acid. Plant Physiol 158(1):340–351
Chen L, Ren Y, Zhang Y, Xu J, Zhang Z, Wang Y (2012b) Genome-wide profiling of novel and conserved Populus microRNAs involved in pathogen stress response by deep sequencing. Planta 235(5):873–883
Dang Z-h, L-l Z, Wang J, Gao Z, S-b W, Qi Z, Y-c W (2013) Transcriptomic profiling of the salt-stress response in the wild recretohalophyte Reaumuria trigyna. BMC Genomics 14(1):29
Dittrich H, Kutchan TM (1991) Molecular cloning, expression, and induction of berberine bridge enzyme, an enzyme essential to the formation of benzophenanthridine alkaloids in the response of plants to pathogenic attack. Proc Natl Acad Sci U S A 88(22):9969–9973
Dixon DP, Edwards R (2009) Selective binding of glutathione conjugates of fatty acid derivatives by plant glutathione transferases. J Biol Chem 284(32):21249–21256
Dixon DP, Hawkins T, Hussey PJ, Edwards R (2009) Enzyme activities and subcellular localization of members of the Arabidopsis glutathione transferase superfamily. J Exp Bot 60(4):1207–1218
Duplessis S, Major I, Martin F, Séguin A (2009) Poplar and pathogen interactions: insights from Populus genome-wide analyses of resistance and defense gene families and gene expression profiling. Crit Rev Plant Sci 28(5):309–334
Eulgem T (2006) Dissecting the WRKY web of plant defense regulators. Plos Pathog 2(11):e126
Fode B, Siemsen T, Thurow C, Weigel R, Gatz C (2008) The Arabidopsis GRAS protein SCL14 interacts with class II TGA transcription factors and is essential for the activation of stress-inducible promoters. Plant Cell 20(11):3122–3135
Fu B, He S (2012) Transcriptome analysis of silver carp (Hypophthalmichthys molitrix) by paired-end RNA sequencing. DNA Res 19(2):131–142
Garg R, Patel RK, Tyagi AK, Jain M (2011) De novo assembly of chickpea transcriptome using short reads for gene discovery and marker identification. DNA Res 18(1):53–63
Giraud E, Ivanova A, Gordon CS, Whelan J, Considine MJ (2012) Sulphur dioxide evokes a large scale reprogramming of the grape berry transcriptome associated with oxidative signalling and biotic defence responses. Plant Cell Environ 35(2):405–417
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36(10):3420–3435
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29(7):644–652
Harding SA, Roberts DM (1998) Incompatible pathogen infection results in enhanced reactive oxygen and cell death responses in transgenic tobacco expressing a hyperactive mutant calmodulin. Planta 206(2):253–258
Harding SA, Oh S-H, Roberts DM (1997) Transgenic tobacco expressing a foreign calmodulin gene shows an enhanced production of active oxygen species. EMBO J 16(6):1137–1144
He P, Shan L, Sheen J (2007) Elicitation and suppression of microbe-associated molecular pattern‐triggered immunity in plant–microbe interactions. Cell Microbiol 9(6):1385–1396
Jain M, Ghanashyam C, Bhattacharjee A (2010) Comprehensive expression analysis suggests overlapping and specific roles of rice glutathione S-transferase genes during development and stress responses. BMC Genomics 11(1):73
Jung HW, Kim W, Hwang BK (2003) Three pathogen-inducible genes encoding lipid transfer protein from pepper are differentially activated by pathogens, abiotic, and environmental stresses. Plant Cell Environ 26(6):915–928
Kaku H, Nishizawa Y, Ishii-Minami N, Akimoto-Tomiyama C, Dohmae N, Takio K, Minami E, Shibuya N (2006) Plant cells recognize chitin fragments for defense signaling through a plasma membrane receptor. Proc Natl Acad Sci U S A 103(29):11086–11091
Knoth C, Ringler J, Dangl JL, Eulgem T (2007) Arabidopsis WRKY70 is required for full RPP4-mediated disease resistance and basal defense against Hyaloperonospora parasitica. Mol Plant Microbe Interact 20(2):120–128
Lan T, Wang X-R, Zeng Q-Y (2013) Structural and functional evolution of positively selected sites in pine glutathione S-transferase enzyme family. J Biol Chem 288(34):24441–24451
Latinović J, Mazzaglia A, Latinović N, Ivanović M, Gleason ML (2013) Resistance of olive cultivars to Botryosphaeria dothidea, causal agent of olive fruit rot in Montenegro. Crop Prot 48:35–40
Liang H, Maynard CA, Allen RD, Powell WA (2001) Increased Septoria musiva resistance in transgenic hybrid poplar leaves expressing a wheat oxalate oxidase gene. Plant Mol Biol 45(6):619–629
Lu R, Malcuit I, Moffett P, Ruiz MT, Peart J, Wu A-J, Rathjen JP, Bendahmane A, Day L, Baulcombe DC (2003) High throughput virus-induced gene silencing implicates heat shock protein 90 in plant disease resistance. EMBO J 22(21):5690–5699
Martin F, Gianinazzi‐Pearson V, Hijri M, Lammers P, Requena N, Sanders I, Shachar-Hill Y, Shapiro H, Tuskan G, Young J (2008) The long hard road to a completed Glomus intraradices genome. New Phytol 180(4):747–750
Martinelli F, Reagan RL, Uratsu SL, Phu ML, Albrecht U, Zhao W, Davis CE, Bowman KD, Dandekar AM (2013) Gene regulatory networks elucidating Huanglongbing disease mechanisms. PLoS One 8(9):e74256
Mizrachi E, Hefer CA, Ranik M, Joubert F, Myburg AA (2010) De novo assembled expressed gene catalog of a fast-growing Eucalyptus tree produced by Illumina mRNA-Seq. BMC Genomics 11(1):681
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621–628
Niderman T, Genetet I, Bruyere T, Gees R, Stintzi A, Legrand M, Fritig B, Mosinger E (1995) Pathogenesis-related PR-1 proteins are antifungal (isolation and characterization of three 14-kilodalton proteins of tomato and of a basic PR-1 of tobacco with inhibitory activity against Phytophthora infestans). Plant Physiol 108(1):17–27
Nie Q, Fang M, Jia X, Zhang W, Zhou X, He X, Zhang X (2011) Analysis of muscle and ovary transcriptome of Sus scrofa: assembly, annotation and marker discovery. DNA Res 18(5):343–351
Passos MA, de Cruz VO, Emediato FL, de Teixeira CC, Azevedo VCR, Brasileiro AC, Amorim EP, Ferreira CF, Martins NF, Togawa RC (2013) Analysis of the leaf transcriptome of Musa acuminata during interaction with Mycosphaerella musicola: gene assembly, annotation and marker development. BMC Genomics 14(1):78
Petre B, Morin E, Tisserant E, Hacquard S, Da Silva C, Poulain J, Delaruelle C, Martin F, Rouhier N, Kohler A (2012) RNA-Seq of Early-Infected Poplar Leaves by the Rust Pathogen Melampsora larici-populina Uncovers PtSultr3; 5, a Fungal-Induced Host Sulfate Transporter. Plos One 7(8):e44408
Poovaiah B, Du L, Wang H, Yang T (2013) Recent advances in calcium/calmodulin-mediated signaling with an emphasis on plant-microbe interactions. Plant Physiol 163(2):531–542
Rietz S, Bernsdorff FE, Cai D (2012) Members of the germin-like protein family in Brassica napus are candidates for the initiation of an oxidative burst that impedes pathogenesis of Sclerotinia sclerotiorum. J Exp Bot 63(15):5507–5519
Roberts DM, Harmon AC (1992) Calcium-modulated proteins: targets of intracellular calcium signals in higher plants. Annu Rev Plant Biol 43(1):375–414
Saijo Y, Tintor N, Lu X, Rauf P, Pajerowska-Mukhtar K, Häweker H, Dong X, Robatzek S, Schulze-Lefert P (2009) Receptor quality control in the endoplasmic reticulum for plant innate immunity. EMBO J 28(21):3439–3449
Salem M, Rexroad C, Wang J, Thorgaard G, Yao J (2010) Characterization of the rainbow trout transcriptome using Sanger and 454-pyrosequencing approaches. BMC Genomics 11(1):564
Santamaria M, Thomson CJ, Read ND, Loake GJ (2001) The promoter of a basic PR1-like gene, AtPRB1, from Arabidopsis establishes an organ-specific expression pattern and responsiveness to ethylene and methyl jasmonate. Plant Mol Biol 47(5):641–652
Schaller A, Oecking C (1999) Modulation of plasma membrane H+-ATPase activity differentially activates wound and pathogen defense responses in tomato plants. Plant Cell 11(2):263–272
Schlumbaum A, Mauch F, Vögeli U, Boller T (1986) Plant chitinases are potent inhibitors of fungal growth. Nature 324:365–367
Shimaoka T, Ohnishi M, Sazuka T, Mitsuhashi N, Hara-Nishimura I, Shimazaki K-I, Maeshima M, Yokota A, Tomizawa K-I, Mimura T (2004) Isolation of intact vacuoles and proteomic analysis of tonoplast from suspension-cultured cells of Arabidopsis thaliana. Plant Cell Physiol 45(6):672–683
Srivastava A, Rogers WL, Breton CM, Cai L, Malmberg RL (2011) Transcriptome analysis of Sarracenia, an insectivorous plant. DNA Res 18(4):253–261
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313(5793):1596–1604
Varshney RK, Nayak SN, May GD, Jackson SA (2009) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotechnol 27(9):522–530
Vijayan P, Shockey J, Lévesque CA, Cook RJ (1998) A role for jasmonate in pathogen defense of Arabidopsis. Proc Natl Acad Sci U S A 95(12):7209–7214
Wan J, Pentecost G (2013) Potential Application of Chitin Signaling in Engineering Broad-Spectrum Disease Resistance to Fungal and Bacterial Pathogens in Plants. Adv Crop Sci Tech 1:e103
Wang L, Song X, Gu L, Li X, Cao S, Chu C, Cui X, Chen X, Cao X (2013a) NOT2 proteins promote polymerase II–dependent transcription and interact with multiple microRNA biogenesis factors in arabidopsis. Plant Cell 25(2):715–727
Wang X-C, Zhao Q-Y, Ma C-L, Zhang Z-H, Cao H-L, Kong Y-M, Yue C, Hao X-Y, Chen L, Ma J-Q (2013b) Global transcriptome profiles of Camellia sinensis during cold acclimation. BMC Genomics 14(1):415
Weaver D (1974) A gummosis disease of peach trees caused by Botryosphaeria dothidea. Phytopathology 64(12):1429–1432
Winkler A, Hartner F, Kutchan TM, Glieder A, Macheroux P (2006) Biochemical evidence that berberine bridge enzyme belongs to a novel family of flavoproteins containing a bi-covalently attached FAD cofactor. J Biol Chem 281(30):21276–21285
Xiang Y, Hua X, Zhao J (1979) The identification on the causal organism of the blister-type canker of poplar. Acta Microbiol Sin 19:57–63
Zhao J-P, Jiang X-L, Zhang B-Y, Su X-H (2012) Involvement of microRNA-mediated gene expression regulation in the pathological development of stem canker disease in Populus trichocarpa. Plos One 7(9):e44968
Zhou Y, Liu H, Zhao J, Tan M, Sui P, Wang J, Zhou L (2008) Poplar stem blister canker and its control strategies by plant extracts. World J Microbiol Biotechnol 24(8):1579–1584
Acknowledgments
This work was supported by National Science & Technology Pillar Programe (2012BAD01B0302), National High Technology Research and Development Program (2013AA102703), and the Changjiang Scholars and Innovative Research Team Program (IRT13047) of China. We sincerely thank Professor Wei He at the Beijing Forestry University for providing Botryosphaeria dothidea strain.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
List of DETs highly expressed following infection. Putative functions were determined by BLASTing against the Nr (non-redundant) and Swiss-prot databases using an E-value of 1.00E-5. (XLS 189 kb)
Table S2
Summary and functional annotation of DETs identified related to plant–pathogen interaction. DETs listed in this table were identified by their functional annotations. (XLS 536 kb)
Table S3
Annotation and categories of transcription factors. Transcripts were identified as transcription factors by BLASTing against the TAIR database (http://www.arabidopsis.org/index.jsp), which contains a list of 5597 TFs, classified into 50 TF families, their names, AGI accession numbers, and putative functions. TFs highly expressed following infection are listed with their fold-changes, AGI accession numbers, and putative functions. (XLS 659 kb)
Table S4
Sequences of validated genes and primers used for RT-qPCR verification. (XLS 68 kb)
Figure S1
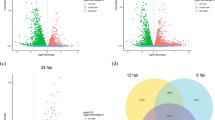
Identification of DETs between the NB and WB libraries. (A) Venn-diagram of expressed poplar transcripts. The upper circle represents for WB library (325,873 transcripts), and the lower circle represents for NB library (276,266 transcripts). The hatched area represents all DETs (206,586 transcripts). 130,159 DETs and 132,697 unchanged transcripts were expressed in both libraries (section c and section b, respectively). 63,017 DETs were expressed only in the WB library (section a), and 13,410 DETs were expressed only in the NB library (section d). (B) Ln-transformed expression levels of poplar transcripts. DETs were determined using a threshold of FDR ≤ 0.05 and an absolute value of log2Ratio (NB/WB) ≥ 1. DETs are colored green (downregulated), red (upregulated), or blue (not DETs). The x-axis represents the ratio of log10 (NB RPKM), and the y-axis represents the ratio of log10 (WB RPKM). (GIF 45 kb)
Figure S2
Validation of RNA-Seq results by RT-qPCR. The x- and y-axis indicate the ratio of log2 fold change in RT-qPCR and RNA-Seq. The figure shows 18 differentially expressed genes, each of which is represented by two treatments (uninfected and infected with B. dothidea). RT-qPCR data represent the mean values of three independent replicates. (GIF 2 kb)
Rights and permissions
About this article
Cite this article
Liao, W., Ji, L., Wang, J. et al. Identification of glutathione S-transferase genes responding to pathogen infestation in Populus tomentosa . Funct Integr Genomics 14, 517–529 (2014). https://doi.org/10.1007/s10142-014-0379-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-014-0379-y