Abstract
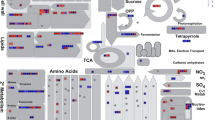
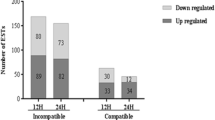
The Asian rice gall midge [Orseolia oryzae (Wood-Mason)] is an important rice pest causing an annual average yield loss of about US $80 million in India. Rice varieties possess several discrete resistance (R) genes conferring resistance against the pest in two distinct ways, i.e., with (HR+ type) or without (HR− type) the expression of hypersensitive reaction (HR). The aim of the present work is to understand the molecular basis of compatible and incompatible (HR− type) rice gall midge interactions between the rice variety Kavya and the two gall midge biotypes: the virulent GMB4M and the avirulent GMB1 using transcriptional microarray gene expression analysis. A large number of differentially expressed genes (602genes in incompatible interaction and 1,330 genes in compatible interaction with at least twofold changes, p value <0.05) was obtained from the microarray analysis that could be grouped into six clusters based on their induction during both or either of the interactions. MapMan software was used for functional characterization of these genes into 13 categories (BINs). Real-time polymerase chain reaction validation of 26 genes selected through the analysis revealed four genes viz. NADPH oxidase, AtrbohF, cinnamoyl-CoA reductase, and von Willebrand factor type A domain containing protein coding genes to be significantly upregulated during the incompatible interaction. But most of the signature genes related to HR+ type resistance like salicylic acid pathway-related genes and disease resistance protein coding genes were downregulated. On the other hand, during the compatible interaction, genes related to primary metabolism and nutrient transport were upregulated and genes for defense and signaling were downregulated. We propose a hypothesis that HR− type of resistance in the rice variety Kavya against gall midge could be due to the constitutive expression of an R gene and a case of extreme resistance which is devoid of cell death. Compatible interaction, however, modulated a large number of differentially expressed transcripts to reprogram cell organization, cell remodeling, and relocation of nutrients through transport to support insect growth.







Similar content being viewed by others
References
Alexander D, Goodman RM, Gut-Rella M, Glascock C, Weymann K, Friedrich L, Maddox D, Ahl-Goy P, Luntz T, Ward E (1993) Increased tolerance to 2 oomycete pathogens in transgenic tobacco expressing pathogenesis related protein-1α. Proc Natl Acad Sci USA 90:7327–7331
Alvarez ME, Pennell RI, Meijer PJ, Ishikawa A, Dixon RA, Lamb C (1998) Reactive oxygen intermediates mediate a systemic signal network in the establishment of plant immunity. Cell 92:773–784
Anderson KG, Harris MO (2006) Does R gene resistance allow wheat to prevent plant growth effects associated with Hessian fly (Diptera: Cecidomyiidae) attack? J Econ Entomol 99:1842–1853
Atkinson MM, Mildland SL, Sims JJ, Keen NT (1996) Syringolide triggers Ca2+ influx, K+ efflux, and extracellular alkalization in soybean cells carrying the disease-resistance gene rpG4. Plant Physiol 112:297–302
Baulcombe DC, Gilbert JE, Goulden M, Köhm BA, Santa Cruz S (1994) Molecular biology of resistance to potato virus X in potato. Biochem Soc Symp 60:207–218
Bendahmane A, Kanyuka K, Baulcombe DC (1999) The Rx gene from potato controls separate virus resistance and cell death responses. Plant Cell 11:781–791
Bentur JS, Kalode MB (1996) Hypersensitive reaction and induced resistance in rice against Asian rice gall midge, Orseolia oryzae. Entomol Exp Appl 78:77–81
Bentur JS, Pasalu IC, Sarma NP, Prasada Rao U, Mishra B (2003) Gall midge resistance in rice. Directorate of Rice Research, Hyderabad
Bentur JS, Cheralu C, Rao PRM (2008) Monitoring virulence in Asian rice gall midge populations in India. Entomol Exp Appl 129:96–106
Biradar SK, Sundaram RM, Thirumurugan T, Bentur JS, Amudhan S, Shenoy VV, Mishra B, Bennet J, Sarma NP (2004) Identification of flanking SSR markers for a major rice gall midge resistance gene Gm1 and their validation. Theor Appl Genet 10:1468–1472
Bolton MD, Kolmer JA, Xu WW, Garvin DF (2008) Lr-34 mediated leaf rust resistance in wheat transcript profiling reveals a high energetic supported by transient recruitment of multiple metabolic pathways. Mol Plant Microbe Interact 21:1515–1527
Dangl JL, Dietrich RA, Richberg MH (1996) Death don’t have no mercy: cell death programs in plant–microbe interactions. Plant Cell 8:1793–1807
Delledonne M, Murgia I, Ederle D, Sbicego PF, Biondani A, Polverari A, Lamb C (2002) Reactive oxygen intermediates modulate nitric oxide signaling in the plant hypersensitive disease response. Plant Physiol Biochem 40:605–610
Freialdenhoven A, Scherag B, Hollricher K, Collinge DB, Thordal-Christensen H, Schulze-Leferd P (1994) Nar-1 and Nar-2, two loci required for Mla 12 specified race specific resistance to powdery mildew in Barley. Plant cell 10:947–956
Fung RWM, Gonzalo M, Fekete C, Kovacs LG, He Y, Marsh E, McIntyre LM, Schachtman DP, Qiu W (2008) Powdery mildew induces defense-oriented reprogramming of the transcriptome in a susceptible but not in a resistant grapevine. Plant Physiol 146:236–249
Garza R, Vera J, Cardona C, Barcenas N, Singh SP (2001) Hypersensitive response of beans to Apion godmani (Coleoptera: Curculionidae). J Econ Entomol 94:958–962
Gatehouse JA (2002) Plant resistance towards insect herbivores: a dynamic interaction. New Phytol 156:145–169
Greenberg JT, Yao N (2004) The role and regulation of programmed cell death in plant–pathogen interactions. Cellular Microbiol 6:201–211
Grover PBJ (1995) Hypersensitive response of wheat to Hessian fly. Entomol Exp Appl 74:283–294
Hammond-Kosack KE, Silverman P, Raskin I, Jones JDG (1996) Race-specific elicitors of Cladosporium fulvum induce changes in cell morphology and the synthesis of ethylene and salicylic acid in tomato cells carrying the corresponding Cf disease resistance gene. Plant Physiol 110:1381–1394
Harris MO, Freeman TP, Rohfritsch O, Anderson KG, Payne SA, Moore JA (2006) Virulent Hessian fly (Diptera: Cecidomyiidae) larvae induce a nutritive tissue during compatible interactions with wheat. Ann Entomol Soc Am 99:305–316
Heath MC (2000) Hypersensitive response-related death. Plant Mol Biol 44:321–334
Herdt RW (1991) Research priorities for rice biotechnology. In: Khush GS, Toenniessen GH (eds) Rice biotechnology. CABI and International Rice Research Institute, Manila (Philippines), pp 19–54
Himabindu K (2009) Identification, tagging and mapping of rice gall midge resistance genes using microsatellite markers. Dissertation, Acharya Nagarjuna University
Himabindu K, Suneetha K, Sama VSAK, Bentur JS (2010) A new rice gall midge resistance gene in the breeding line CR57-MR1523, mapping with flanking markers and development of NILs. Euphytica 174:179–187
Hoglund S, Larsson S, Wingsle G (2005) Both hypersensitive and non-hypersensitive responses are associated with resistance in Salix viminalis against the gall midge Dasineura marginemtorquens. J Exp Bot 56:3215–3222
Hulbert SH, Bai J, Fellers JP, Pacheco MG, Bowden RL (2007) Gene expression patterns in near isogenic lines for wheat rust resistance gene Lr34/Yr18. Phytopathol 97:1083–1093
Kawasaki T, Koita H, Nakatsubo T, Hasegawa K, Wakabayashi K (2006) Cinnamoyl-CoA reductase a key enzyme in lignin biosynthesis, is an effector of small GTPase Rac in defense signaling in rice. Proc Natl Acad Sci USA 103:230–235
Keen NT, Littlefield LJ (1979) The possible association of phytoalexins with resistance gene expression in flax to Melamspora lini. Physiol Plant Pathol 14:265–280
Knogge W (1996) Fungal infection of plants. Plant Cell 8:1711–1722
Kohm BA, Goulden MG, Gilbert JE, Kavanagh TA, Baulcombe DC (1993) A potato virus X resistance gene mediates an induced, nonspecific resistance in protoplasts. Plant Cell 5:913–920
Li C, Bai Y, Jacobsen E, Visser R, Lindhout P, Bonnema G (2006) Tomato defense to the powdery mildew fungus: differences in expression of genes in susceptible, monogenic- and polygenic-resistance responses are mainly in timing. Plant Mol Biol 62:127–140
Liu X, Bai J, Huang L, Zhu L, Liu X et al (2007) Gene expression of different wheat genotypes during attack by virulent and avirulent Hessian fly (Mayetiola destructor) larvae. J Chem Ecol 33:2171–2194
Liu X, Williams CE, Nemacheck JA, Wang H, Subramanyam S, Zheng C, Chen MS (2010) Reactive oxygen species are involved in plant defense against a gall midge. Plant Physiol 152:985–999
Pfitzner UM, Goodman HM (1987) Isolation and characterization of cDNA clones encoding pathogenesis-related proteins from tobacco mosaic virus infected tobacco plants. Nucleic Acids Res 15:4449–4465
Poncianoa G, Yoshikawa M, Lee JL, Ronald PC, Whalena MC (2006) Pathogenesis-related gene expression in rice is correlated with developmentally controlled Xa21-mediated resistance against Xanthomonas oryzae pv. oryzae. Physiol Mol Plant Pathol 69:131–139
Rawat N (2012) Identification and characterization of differentially expressed genes in rice involved in rice gall midge interactions. Dissertation, Osmania University
Rawat N, Sinha DK, Rajendrakumar P, Srivastava P, Neeraja CN, Sundaram RM, Nair S, Bentur JS (2010) Role of pathogenesis related genes in rice gall midge interactions. Curr Sci 99:1361–1368
Regupathy A, Subramanian A (1972) Effect of different doses of fertilizers on the mineral metabolism of IR8 rice in relation to its susceptibility to gall fly Pachydiplosis oryzae, Wood-Manson, and leaf roller, Cnaphalocrosis medinalis Guenee. Oryza 9:81–85
Ryals JA, Neuenschwander UH, Willits MG, Molina A, Steiner HY, Hunt MD (1996) Systemic acquired resistance. Plant Cell 8:1809–1819
Sauge MH, Kervella J, Pascal T (1998) Settling behaviour and reproductive potential of the green peach aphid Myzus persicae on peach varieties and a related wild Prunus. Entomol Exp Appl 89:233–242
Staskawicz BJ, Ausubel FM, Baker BJ, Ellis JG, Jones JDG (1995) Molecular genetics of plant disease resistance. Science 268:661–667
Swarbrick PJ, Huang K, Liu G, Slate J, Press MC, Scholes JD (2008) Global patterns of gene expression in rice cultivars undergoing a susceptible or resistant interaction with the parasitic plant Striga hermonthica. New Phytol 179:515–529
Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Kruger P, Selbig J, Muller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
Torres MA, Dangl JL, Jones JD (2002) Arabidopsis gp91phox homologues AtrbohD and AtrbohF are required for accumulation of reactive oxygen intermediates in the plant defense response. Proc Natl Acad Sci USA 99:517–522
Torres MA, Jones JD, Dangl JL (2005) Pathogen-induced, NADPH oxidase-derived reactive oxygen intermediates suppress spread of cell death in Arabidopsis thaliana. Nat Genet 37:1130–1134
Torres MA, Jones JDG, Dangl JL (2006) Reactive oxygen species signaling in response to pathogens. Plant Physiol 141:373–378
Usadel B, Nagel A, Thimm O, Redestig H, Blaesing OE, Palacios-Rojas N, Selbig J, Hannemann J, Piques MC, Steinhauser D (2005) Extension of the visualization tool MapMan to allow statistical analysis of arrays, display of corresponding genes, and comparison with known responses. Plant Physiol 138:1195–1204
Vijaya Lakshmi P, Amudhan S, Himabindu K, Cheralu C, Bentur JS (2006) A new biotype of the Asian rice gall midge Orseolia oryzae (Diptera: Cecidomyiidae) characterized from the Warangal population in Andhra Pradesh, India. Int J Trop Sci 26:207–211
Walling LL (2000) The myriad plant responses to herbivores. J Plant Growth Regul 19:195–216
Yu IC, Parker J, Bent AF (1998) Gene-for-gene disease resistance without the hypersensitive response in Arabidopsis dnd 1 mutant. Proc Natl Acad Sci USA 95:7819–7824
Acknowledgments
We thank the Project Director, Directorate of Rice Research, Hyderabad and the Director, International Centre for Genetic Engineering and Biotechnology, New Delhi for the facilities and encouragement. This work was supported by a research grant (NFBSFARA/PCN/AP01/2006-07) from the National Fund of the National Agricultural Innovative Project of the Indian Council of Agricultural Research, New Delhi, to JSB.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary materials
Below is the link to the electronic supplementary material.
10142_2012_275_MOESM3_ESM.xls
List of different rice pathways, downregulated in the Kavya–GMB4M interaction. Shaded area in the sheet represents the significant pathways list (p < 0.05; XLS 54 kb)
Rights and permissions
About this article
Cite this article
Rawat, N., Chiruvuri Naga, N., Raman Meenakshi, S. et al. A novel mechanism of gall midge resistance in the rice variety Kavya revealed by microarray analysis. Funct Integr Genomics 12, 249–264 (2012). https://doi.org/10.1007/s10142-012-0275-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-012-0275-2