Abstract
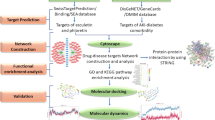
A significant proportion of patients develop left ventricular (LV) dysfunction and heart failure (HF) after acute myocardial infarction (MI). Existing biomarkers of HF provide limited information after MI. To identify new prognostic biomarkers in MI patients, we designed an approach combining protein interaction networks and microarray analysis of blood cells. Blood samples for RNA and protein analysis were taken from 127 acute MI patients. Echocardiography was performed at one month. Assuming that angiogenesis is related to cardiac repair after MI, a protein-protein interaction network of angiogenesis was constructed and analyzed. Among the 556 proteins and 686 interactions of this network, a cluster of 53 proteins highly specialized in regulation of cell growth was identified. Of these 53 proteins, 38 were found differentially expressed by microarrays between low (≤ 40%) and high (>40%) LV ejection fraction (EF) patients (n = 32). Among these 38 genes, prediction analysis identified a set of three genes able to predict significant LV dysfunction (EF ≤ 40%) with an area under the receiver operating characteristic curve (AUC) of 0.82. These three genes—vascular endothelial growth factor B, thrombospondin-1 and placental growth factor—had a stronger predictive value than brain natriuretic peptide and troponin T (AUC of 0.63). Independent validations on protein expression and quantitative PCR datasets confirmed the results. In conclusion, a new strategy is described that allows identifying new potential biomarkers. The three specific biomarkers described here remain to be validated in a larger patient population.






Similar content being viewed by others
References
Al-Shahrour F, Minguez P, Tarraga J, Montaner D, Alloza E, Vaquerizas JM, Conde L, Blaschke C, Vera J, Dopazo J (2006) BABELOMICS: a systems biology perspective in the functional annotation of genome-scale experiments. Nucleic Acids Res 34:W472–W476
Bader GD, Hogue CW (2003) An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 4:2
Bellomo D, Headrick JP, Silins GU, Paterson CA, Thomas PS, Gartside M, Mould A, Cahill MM, Tonks ID, Grimmond SM, Townson S, Wells C, Little M, Cummings MC, Hayward NK, Kay GF (2000) Mice lacking the vascular endothelial growth factor-B gene (Vegfb) have smaller hearts, dysfunctional coronary vasculature, and impaired recovery from cardiac ischemia. Circ Res 86:E29–E35
Braunwald E (2008) Biomarkers in heart failure. N Engl J Med 358:2148–2159
Brindle JT, Antti H, Holmes E, Tranter G, Nicholson JK, Bethell HW, Clarke S, Schofield PM, McKilligin E, Mosedale DE, Grainger DJ (2002) Rapid and noninvasive diagnosis of the presence and severity of coronary heart disease using 1H-NMR-based metabonomics. Nat Med 8:1439–1444
Cappuzzello C, Napolitano M, Arcelli D, Melillo G, Melchionna R, Di Vito L, Carlini D, Silvestri L, Brugaletta S, Liuzzo G, Crea F, Capogrossi MC (2009) Gene expression profiles in peripheral blood mononuclear cells of chronic heart failure patients. Physiol Genomics 38:233–240
Carmeliet P, Baes M (2008) Metabolism and therapeutic angiogenesis. N Engl J Med 358:2511–2512
Chatila K, Ren G, Xia Y, Huebener P, Bujak M, Frangogiannis NG (2007) The role of the thrombospondins in healing myocardial infarcts. Cardiovasc Hematol Agents Med Chem 5:21–27
Chin BS, Blann AD, Gibbs CR, Chung NA, Conway DG, Lip GY (2003) Prognostic value of interleukin-6, plasma viscosity, fibrinogen, von Willebrand factor, tissue factor and vascular endothelial growth factor levels in congestive heart failure. Eur J Clin Invest 33:941–948
Cleary JG, Trigg L (1995) K*: an instance-based learner using an entropic distance measure. Proc Machine Learning '95, Lake Tahoe, USA:108–114
Ferrara N, Gerber HP, LeCouter J (2003) The biology of VEGF and its receptors. Nat Med 9:669–676
Frank E, Hall M, Trigg L, Holmes G, Witten IH (2004) Data mining in bioinformatics using Weka. Bioinformatics 20:2479–2481
Good DJ, Polverini PJ, Rastinejad F, Le Beau MM, Lemons RS, Frazier WA, Bouck NP (1990) A tumor suppressor-dependent inhibitor of angiogenesis is immunologically and functionally indistinguishable from a fragment of thrombospondin. Proc Natl Acad Sci U S A 87:6624–6628
Hoeben A, Landuyt B, Highley MS, Wildiers H, Van Oosterom AT, De Bruijn EA (2004) Vascular endothelial growth factor and angiogenesis. Pharmacol Rev 56:549–580
Horwitz PA, Tsai EJ, Putt ME, Gilmore JM, Lepore JJ, Parmacek MS, Kao AC, Desai SS, Goldberg LR, Brozena SC, Jessup ML, Epstein JA, Cappola TP (2004) Detection of cardiac allograft rejection and response to immunosuppressive therapy with peripheral blood gene expression. Circulation 110:3815–3821
King JY, Ferrara R, Tabibiazar R, Spin JM, Chen MM, Kuchinsky A, Vailaya A, Kincaid R, Tsalenko A, Deng DX-F, Connolly A, Zhang P, Yang E, Watt C, Yakhini Z, Ben-Dor A, Adler A, Bruhn L, Tsao P, Quertermous T, Ashley EA (2005) Pathway analysis of coronary atherosclerosis. Physiol Genomics 23:103–118
Kolakowski S, Berry MF, Atluri P, Grand T, Fisher O, Moise MA, Cohen J, Hsu V, Woo YJ (2006) Placental growth factor provides a novel local angiogenic therapy for ischemic cardiomyopathy. J Card Surg 21:559–564
Le Brigand K, Barbry P (2007) Mediante: a web-based microarray data manager. Bioinformatics 23:1304–1306
Le Brigand K, Russell R, Moreilhon C, Rouillard JM, Jost B, Amiot F, Magnone V, Bole-Feysot C, Rostagno P, Virolle V, Defamie V, Dessen P, Williams G, Lyons P, Rios G, Mari B, Gulari E, Kastner P, Gidrol X, Freeman TC, Barbry P (2006) An open-access long oligonucleotide microarray resource for analysis of the human and mouse transcriptomes. Nucleic Acids Res 34:e87
Lee SH, Wolf PL, Escudero R, Deutsch R, Jamieson SW, Thistlethwaite PA (2000) Early expression of angiogenesis factors in acute myocardial ischemia and infarction. N Engl J Med 342:626–633
Michowitz Y, Goldstein E, Wexler D, Sheps D, Keren G, George J (2007) Circulating endothelial progenitor cells and clinical outcome in patients with congestive heart failure. Heart 93:1046–1050
Montaner D, Tarraga J, Huerta-Cepas J, Burguet J, Vaquerizas JM, Conde L, Minguez P, Vera J, Mukherjee S, Valls J, Pujana MA, Alloza E, Herrero J, Al-Shahrour F, Dopazo J (2006) Next station in microarray data analysis: GEPAS. Nucleic Acids Res 34:W486–W491
Nonaka-Sarukawa M, Yamamoto K, Aoki H, Nishimura Y, Tomizawa H, Ichida M, Eizawa T, Muroi K, Ikeda U, Shimada K (2007) Circulating endothelial progenitor cells in congestive heart failure. Int J Cardiol 119:344–348
Novikov E, Barillot E (2007) Software package for automatic microarray image analysis (MAIA). Bioinformatics 23:639–640
Peri S, Navarro JD, Amanchy R, Kristiansen TZ, Jonnalagadda CK, Surendranath V, Niranjan V, Muthusamy B, Gandhi TK, Gronborg M, Ibarrola N, Deshpande N, Shanker K, Shivashankar HN, Rashmi BP, Ramya MA, Zhao Z, Chandrika KN, Padma N, Harsha HC, Yatish AJ, Kavitha MP, Menezes M, Choudhury DR, Suresh S, Ghosh N, Saravana R, Chandran S, Krishna S, Joy M, Anand SK, Madavan V, Joseph A, Wong GW, Schiemann WP, Constantinescu SN, Huang L, Khosravi-Far R, Steen H, Tewari M, Ghaffari S, Blobe GC, Dang CV, Garcia JG, Pevsner J, Jensen ON, Roepstorff P, Deshpande KS, Chinnaiyan AM, Hamosh A, Chakravarti A, Pandey A (2003) Development of human protein reference database as an initial platform for approaching systems biology in humans. Genome Res 13:2363–2371
Platt J (1998) Sequential minimal optimization: a fast algorithm for training support vector machines. Microsoft Research Technical Report MSR-TR-98-14. http://research.microsoft.com/en-us/um/people/jplatt/smotr.pdf. Accessed 16 April 2010
Richards AM, Nicholls MG, Espiner EA, Lainchbury JG, Troughton RW, Elliott J, Frampton C, Turner J, Crozier IG, Yandle TG (2003) B-type natriuretic peptides and ejection fraction for prognosis after myocardial infarction. Circulation 107:2786–2792
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504
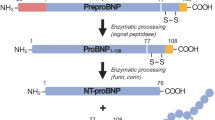
Talwar S, Squire IB, Downie PF, McCullough AM, Campton MC, Davies JE, Barnett DB, Ng LL (2000) Profile of plasma N-terminal proBNP following acute myocardial infarction; correlation with left ventricular systolic dysfunction. Eur Heart J 21:1514–1521
Torabi A, Cleland JG, Khan NK, Loh PH, Clark AL, Alamgir F, Caplin JL, Rigby AS, Goode K (2008) The timing of development and subsequent clinical course of heart failure after a myocardial infarction. Eur Heart J 29:859–870
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci U S A 98:5116–5121
Ware JH (2006) The limitations of risk factors as prognostic tools. N Engl J Med 355:2615–2617
Wingrove JA, Daniels SE, Sehnert AJ, Tingley W, Elashoff M, Rosenberger S, Buellesfeld L, Grube E, Newby LK, Ginsburg GS, Kraus WE (2008) Correlation of peripheral-blood gene expression with the extent of coronary artery stenosis. Circ Cardiovasc Genet 1:31–38
Zentilin L, Puligadda U, Lionetti V, Zacchigna S, Collesi C, Pattarini L, Ruozi G, Camporesi S, Sinagra G, Pepe M, Recchia FA, Giacca M (2009) Cardiomyocyte VEGFR-1 activation by VEGF-B induces compensatory hypertrophy and preserves cardiac function after myocardial infarction. FASEB J DOI. doi:10.1096/fj.09-143180
Zethelius B, Berglund L, Sundstrom J, Ingelsson E, Basu S, Larsson A, Venge P, Arnlov J (2008) Use of multiple biomarkers to improve the prediction of death from cardiovascular causes. N Engl J Med 358:2107–2116
Acknowledgments
The authors thank Céline Jeanty, Malou Gloesener and Loredana Jacobs for expert technical assistance. We also thank Dr. Georges Gilson, Laboratory of Biochemistry, Centre Hospitalier, Luxembourg, for NT-pro-BNP assessment. This work was supported by grants from the Society for Research on Cardiovascular Diseases, the National Funds for Research, and the Ministry of Culture, Higher Education and Research of Luxembourg.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Devaux, Y., Azuaje, F., Vausort, M. et al. Integrated protein network and microarray analysis to identify potential biomarkers after myocardial infarction. Funct Integr Genomics 10, 329–337 (2010). https://doi.org/10.1007/s10142-010-0169-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-010-0169-0