Abstract
Background
Gastric lymphoepithelioma-like carcinoma (LELC) is a rare entity that is closely associated with Epstein–Barr virus (EBV). However, the EBV latency pattern and genome polymorphisms in gastric LELC have not been systematically explored.
Methods
The clinicopathological features, EBV latency pattern and genome polymorphisms of EBV-positive gastric LELC in Guangzhou, southern China were investigated and compared with those of ordinary EBV-associated gastric carcinoma (EBVaGC) in the same area.
Results
Ten (1.42 %) of 702 gastric carcinoma cases were identified as gastric LELC, in which eight (80 %) cases were EBV-positive. The clinicopathological characteristics and EBV latency pattern of EBV-positive gastric LELC were similar to those of ordinary EBVaGC. In EBV genotype analysis, type A strain, type F, I, mut-W1/I, XhoI− and del-LMP1 variants were predominant among EBV-positive gastric LELCs, accounting for eight (100 %), six (75 %), eight (100 %), seven (87.5 %), five (62.5 %) and six (75 %) cases, respectively, which are similar to those in ordinary EBVaGC. For EBNA1 polymorphisms, the V-leu and P-ala subtypes were predominant in EBV-positive gastric LELC, which is different from the predominant V-val subtype in ordinary EBVaGC. EBV-positive gastric LELC has a favorable prognosis when compared to ordinary EBVaGC (median survival time 43.0 vs. 18.0 months).
Conclusions
Gastric LELC is strongly associated with EBV and EBV-positive gastric LELC should be regarded as a special subtype of EBVaGC. This, to our best knowledge, is the first time in the world that the EBV latency pattern and genome polymorphisms of EBV-positive gastric LELC are systematically revealed.
Similar content being viewed by others
Introduction
Lymphoepithelioma-like carcinomas (LELCs) are defined as tumors with histologic features similar to those of undifferentiated nasopharyngeal carcinoma (NPC) [1–3]. They are rare and have been reported in different anatomic sites, including the salivary gland, thymus, larynx, lung, esophagus, uterine cervix, urinary bladder and skin [4]. Like undifferentiated NPC, LELC show intense lymphocytic infiltration, and invariably, infiltrating lymphocytes are more than tumor cells [1–3]. Two histologic patterns are described. The first one, the so-called Regaud type, displays well-defined epithelial nests separated by broad areas of lymphocytic reaction. Tumor cells growing in a diffuse manner mimicking malignant lymphoma characterize the second pattern, designated the Schmincke type [5, 6].
LELC of the stomach is a rare form of gastric carcinoma that was first reported by Watanabe et al. in 1976 [7]. It constitutes about 1.1–4.6 % of all gastric carcinomas, and demonstrates a male predominance, predisposition to the proximal stomach [8, 9]. Besides, gastric LELC has been reported to have a favorable prognosis [10–13]. Gastric LELC consists of two subsets, Epstein–Barr virus (EBV)-positive and microsatellite instability (MSI)-high carcinomas [13]. It is demonstrated that EBV is present in more than 80 % of gastric LELCs, suggesting that gastric LELC is closely associated with EBV infection [7, 8, 10, 14]. The prevalence of MSI-high carcinoma in gastric LELC ranges from 7 to 39 %, with apparent geographic variability [13]. The hypermethylation of MLH1 promoter, strongly correlating with the loss of MLH1 expression, is the main inactivation mechanism of the MLH1 gene responsible for MSI-high in gastric carcinoma [15]. It is reported that EBV positivity and MSI-high status are mutually exclusive in gastric carcinomas [13].
EBV is a lymphotropic virus that belongs to the Herpesviridae family and infects over 90 % of adults worldwide. It is closely associated with a wide range of human lymphoid and epithelial malignancies, including Burkitt’s lymphoma (BL), Hodgkin lymphoma (HL), NPC and a subset of gastric carcinoma that is defined as EBV-associated gastric carcinoma (EBVaGC) [16–18]. Besides, a close relationship between EBV and LELC, especially those arising from the embryologic foregut derived organs, such as stomach [13, 19, 20], lung [5, 21], salivary gland [22, 23], and thymus [24], has been reported.
EBVaGC, comprising about 10 % of all gastric carcinoma cases worldwide, represents a distinct type of gastric carcinoma as it has special clinicopathological characteristics such as male predominance and predisposition to the proximal stomach [25, 26]. Histopathologically, EBVaGC has two subtypes, namely LELC and ordinary adenocarcinoma (ordinary EBVaGC) [27].
After infection, EBV persists in the host in a latency cycle and constitutively expresses a limited set of viral gene products, the so-called latent products, which comprise six EBV nuclear antigens (EBNAs 1, 2, 3A, 3B, 3C and -LP), three latent membrane proteins (LMPs 1, 2A and 2B), two EBV-encoded small non-coding RNAs (EBERs 1 and 2) and the BamHI A rightward transcripts (BARTs) [16]. Three latency types have been described, depending on which of these latent products are expressed. Latency I is limited to only EBERs, BARTs and EBNA1 expression; latency II includes LMP1 and 2 in addition; and latency III is defined by expression of EBERs, BARTs, all six EBNA proteins and two LMP proteins. Among them, EBNA1 is the only viral protein consistently expressed in all EBV-associated malignancies [28].
EBV has two distinct strains, type A and type B (also called types 1 and 2) [28]. In addition, based on alignment to the prototype B95.8 genome, five major variants of EBV have been identified. They are F and ‘‘f’’ variants at BamHI-F region, I and ‘‘i’’ variants, wt-W1/I1 and mut-W1/I1 variants at BamHI W1/I1 boundary region, XhoI+ and XhoI− variants at LMP1 exon 1, as well as wt-LMP1 and del-LMP1 at LMP1 exon 3 [29]. Moreover, based on the signature changes at amino acid (aa) residue 487, EBNA1 has been classified into five distinct subtypes, including P-ala, P-thr, V-leu, V-val and V-pro [30, 31].
Although it has been demonstrated that gastric LELC is closely associated with EBV, the EBV latency pattern and genome polymorphisms in gastric LELC have not been systematically explored. Therefore, in the present study, ten cases of gastric LELC screened from 702 consecutive gastric carcinoma cases from Guangzhou, southern China, where NPC is endemic, were investigated for the presence of EBV. The clinicopathological features, EBV latency pattern and genome polymorphisms of EBV-positive gastric LELC were also examined and compared with those of ordinary EBVaGC in the same area.
Methods
Subjects
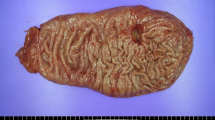
In the present study, gastric LELC was defined as a tumor that is arranged in nests and clusters of poorly or undifferentiated cells showing a syncytial growth pattern surrounded by an intense lymphoplasmacytic cell infiltration. The tumor cells have ill-defined borders with round, oval, or elongated vesicular nuclei with distinct eosinophilic nucleoli. Gastric carcinoma with lymphoid stroma but showing regional glandular differentiation is not considered as gastric LELC [8, 14].
Ten (1.42 %) gastric LELC cases were screened from 702 consecutive specimens of resected gastric carcinoma diagnosed in the Second and the Third Affiliated Hospitals of Sun Yat-sen University, and the Guangzhou First Municipal People’s Hospital, Guangzhou, southern China, from 1 January 2000 to 31 December 2006. Forty-five ordinary EBVaGC cases identified from the remaining 692 gastric carcinoma cases using tumor tissue microarray (TMA) and EBER-1 in situ hybridization (ISH) assay were used as controls.
Clinicopathological data were obtained from the archives of the medical records, including age and sex of the patient as well as the anatomical site, invasion depth and lymphatic and hematogenous metastases of the tumor. Gross features and the TNM staging of the tumor were classified according to the guidelines of the Japanese Research Society for Gastric Cancer [32]. The clinical outcome was followed up from the date of gastric carcinoma resection until the date of death. The follow-up period was 5–71 months (median 24.5 months).
Human tissues used in this study were obtained and managed in accordance with the provisions of the Declaration of Helsinki. Written informed consents were taken from all the patients. This study was approved by the Clinical Research Ethics Committee of the Third Affiliated Hospital, Sun Yat-sen University.
Construction of tumor tissue microarrays
The TMA blocks were constructed as described by Kononen et al. [33]. A 12 × 8 cylinder matrix was defined on a 3.5 × 2.5 cm paraffin wax block, and position-specific blank cylinders were adopted for orientation during microscopy analysis. A total of 15 TMA blocks were constructed, where each TMA block contained up to 94 tissue cores from 47 gastric carcinoma cases and each case was composed of two tissue cores. An adequate case was defined by tumor occupancy of more than 10 % of the core area.
In situ hybridization
ISH assay was performed with a commercially available EBV oligonucleotide probe complementary to EBER-1 (PanPath, Amsterdam, Netherlands), according to the manufacturer’s instructions. Positive signals were recognized as dark brown nuclear staining under light microscope. Sections from a known EBER-1-positive NPC tissue were used as the positive control and a sense probe for EBER-1 was used as the negative control.
Immunohistochemistry
Immunohistochemical analysis was performed in accordance with the manufacturer’s instructions. Monoclonal antibodies used in this study and their retrieval methods as well as dilutions are shown in Table S1 (Online Resource 1). Tumors were considered positive if 10 % or more of the neoplastic cells were stained.
Preparation of DNA
DNA was extracted from formalin-fixed, paraffin-embedded EBER-1-positive gastric LELC and ordinary EBVaGC tissues using the NucleoSpin® Tissue Kit (MACHEREY-NAGEL GmbH & Co. KG, Germany), according to the manufacturer’s protocol. Paraffin blocks without any samples were used as negative controls. DNA extracted from the formalin-fixed, paraffin-embedded EBV-positive cell lines B95.8 and Raji was used as positive controls.
PCR, RFLP and DNA sequencing
Polymerase chain reaction (PCR) was performed with 2 μl of DNA in a total amount of 50 μl reaction mixture comprised of 10 mM Tris–HCl (pH 8.0), 50 mM KCl, 1.5 mM MgCl2, 200 μM dNTP, 0.5 μM of each primer and 1.25U Taq Polymerase (TaKaRa, Dalian, China). Fourteen different regions of EBV genome, including EBNA-3C region, EBNA1 region, BamHI-F region, BamHI-W1/I1 boundary region, exon 1 and exon 3 of LMP1 gene, and exons 1–8 of LMP2A gene, were amplified. The primers and the amplification protocol used in this study are shown in Table S2 (Online Resource 1). The purified PCR products were then subjected to restriction fragment length polymorphism (RFLP) analysis and/or DNA sequencing, respectively.
The RFLP analysis was performed at BamHI-F region, BamHI-W1/I1 boundary region and XhoI restriction site in LMP1 exon 1 with either BamHI or XhoI restriction enzyme (TaKaRa, Dalian, China), following the manufacturer’s instructions. Resultant products were visualized by electrophoresis in a 1.5 % agarose gel stained with 0.5 μg/ml of ethidium bromide.
DNA sequencing was carried out on all purified PCR products to confirm the specificity of PCR reaction and determine the nucleotide sequence variations. Sequence analysis was performed on the 3730xl DNA Analyzers (PE Applied Biosystems, Foster City, CA, USA). The sequencing results were compared with the sequence of EBV strain B95.8 (GenBank Accession No. V01555) to determine the difference in the nucleotide sequences.
Statistical analysis
Fisher’s exact tests were used to compare the clinicopathological characteristics and EBV genome polymorphisms of EBV-positive gastric LELC and ordinary EBVaGC. Survival rates were calculated by Kaplan–Meier method, and differences between survival curves were examined with Log-rank test. The results were considered to be statistically significant at a p value of less than 0.05.
Results
Clinicopathological features of gastric LELC
Clinicopathological features of gastric LELC are summarized in Table 1. In this study, ten LELC cases including eight (80 %) male cases and two female cases were investigated. The mean age was 61.4 ± 14.9 years (range 38–79 years) for all patients, and 61.0 ± 16.0 years (range 38–79 years) for male patients.
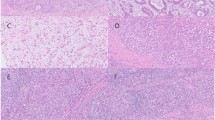
All cases were poorly differentiated gastric carcinomas. Eight (80 %) cases belonged to the Regaud type, showing small nests or confluent sheets of cancer cells, interspersed by wide stromal tissue that contained intense reactive lymphoplasmacytic cells and other inflammatory cells. The tumor cell arranged syncytially with ill-defined cytoplasmic borders (Fig. 1a). The remaining two (20 %) cases (cases #2 and #9) belonged to the Schmincke type, in which the majority of carcinoma cells grow in a diffuse manner, along with intense lymphoplasmacytic cell infiltration (Fig. 1b). In general, tumor cell nuclei were round, oval, or elongated, with mildly irregular nuclear outline, vesicular chromatin, and prominent eosinophilic nucleoli.
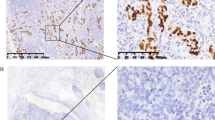
Microscopic features and expression of EBER-1, EBNA1, LMP2A, LMP1, EBNA2, ZEBRA in gastric LELC. a Regaud-type gastric LELC showed large nests of cancer cells with a syncytial arrangement, with ill-defined cytoplasmic borders infiltrated by lymphoplasmacytic cells (H&E, ×200). b Schmincke-type gastric LELC showed diffuse cancer cells with oval nuclei and prominent eosinophilic nucleoli, along with intense lymphoplasmacytic cell infiltration (H&E, ×200). c, d EBER-1-positive signals by in situ hybridization were restricted only to the tumor nuclei, but were not in surrounding infiltrating lymphocytes (c ×100, d ×400). e Immunohistochemistry staining for EBNA1. Positive staining can be seen in almost all tumor nuclei. f Immunohistochemistry staining for LMP2A. Positive staining can be seen in the cytoplasm and membrane of the tumor cells. g–i Immunohistochemistry staining for LMP1, EBNA2 and ZEBRA of the same case, respectively. No positive signals can be seen (×200)
EBER-1 and MLH1 expressed in 80 % gastric LELC
Eight (80 %) out of the ten gastric LELC cases showed both EBER-1 ISH positive signals and MLH1 expression, and were defined as EBV-positive gastric LELCs. The EBER-1 positive signals were restricted only to the nuclei of almost all tumor cells, but were absent in surrounding infiltrating lymphocytes and non-neoplastic gastric mucosa (Fig. 1c, d). The MLH1 positivity was found in almost all tumor nuclei and scattered in some lymphocytes in the germinal center (Figure S1) (Online Resource 1).
The clinicopathological features of EBV-positive gastric LELC were similar to those of ordinary EBVaGC (Table 2).
EBNA1, LMP2A rather than EBNA2, LMP1 and ZEBRA expressed in EBV-positive gastric LELC
The immunohistochemistry assay was performed in eight EBV-positive gastric LELC cases. Seven cases (87.5 %) displayed diffuse positivity for EBNA1 in the nuclei of tumor cells (Fig. 1e). Expression of LMP2A was detected in five cases (62.5 %) (Fig. 1f), while EBNA2, LMP1 and ZEBRA were all negative (Fig. 1g–i). Thus, like ordinary EBVaGC (Table S3) (Online Resource 1), the EBV latency pattern in EBV-positive gastric LELC was type I.
Type A strain, types F, I, mut-W1/I1, XhoI− and del-LMP1 variants predominated in EBV-positive gastric LELC
Amplification of the EBNA-3C region, BamHI-F region, BamHI-W1/I1 boundary region, exon 1 and exon 3 of the LMP1 gene was performed successfully in eight EBV-positive gastric LELC cases. The genotypes of EBV were determined by PCR–RFLP and/or DNA sequencing. The subtype A strain, prototype F, type I, mut-W1/I1, XhoI− and del-LMP1 variants were predominant among EBV-positive gastric LELCs, accounting for eight (100 %), six (75 %), eight (100 %), seven (87.5 %), five (62.5 %) and six (75 %) cases, respectively. EBV genotypes in EBV-positive gastric LELC were illustrated in Table 3, Figure S2 and Table S4 (Online Resource 1).
V-leu and P-ala subtypes were predominant in EBV-positive gastric LELC
EBNA1 fragment was also successfully amplified and sequenced in seven (87.5 %) of eight EBV-positive gastric LELC cases. All nucleotide variations were identified by comparing with the B95.8 prototype sequence. The sequencing results showed the presence of a single EBNA1 sequence in three (42.9 %) of the seven cases, whereas the remaining four (57.1 %) samples displayed dual EBNA1 sequences. The V-leu subtype was the most predominant (71.4 %, 5/7), followed by P-ala (57.1 %, 4/7). The distribution of EBNA1 subtypes in EBV-positive gastric LELC is illustrated in Table 3, Figure S2 and Table S4 (Online Resource 1). All the tumor samples that harbored the V-leu subtype showed the same clustered point mutations. They included 12 nucleotide mutations, leading to 10 aa substitutions (Table S5) (Online Resource 1).
Sequence variations of LMP2A gene in EBV-positive gastric LELC
Exons 1–8 of LMP2A gene were successfully amplified in eight EBV-positive gastric LELC cases. All sequences were compared with the B95.8 prototype sequence. In total, 31 nucleotide mutations were found, which cause 15 aa changes. Among them, eight loci of aa mutation (codons 23, 153, 169, 208, 370, 391, 426 and 444) were detected in ≥ 50 % isolates. The nucleotide variations of LMP2A exons 1–8 in gastric LELC are shown in Table S6 (Online Resource 1).
As we know, the amino-terminal cytoplasmic domain of LMP2A is encoded by exon 1, which contains several functionally important motifs such as immunoreceptor tyrosine-based activation motif (ITAM) and phosphotyrosine (PY) motifs. The ITAM, consisting of paired pivotal tyrosines and leucines (YxxL) at Y74 and Y85, was conserved in all isolates. The two PY motifs (PPPPY Y60 and Y101), were not affected by mutations in any isolates, either.
In the transmembrane region (exons 2–8) of LMP2A, 17 EBV-specific cytotoxic T lymphocytes (CTLs) target epitopes are identified. In the present study, seven of the 17 identified CTL epitopes were affected by at least one point mutation. The amino acid variations of CTL epitopes in gastric LELC were summarized in Table S7 (Online Resource 1). Among them, the L208I at HLA-B60-restricted CTL epitope (codons 200 to 208) and the S444T at HLA-A25-restricted CTL epitope (codons 442 to 451) were detected in 75 % (6/8) and 62.5 % (5/8) cases, respectively.
EBV-positive gastric LELC has a relative excellent prognosis
Thirty-six gastric carcinoma cases including eight EBV-positive gastric LELC cases and 32 ordinary EBVaGC cases were performed for survival analysis. The overall survival time to event was defined as time of surgery until death, irrespective of cause of death. The estimated median survival time was 43.0 months (27.2–58.8 months) and 18.0 months (3.9–32.1 months) for EBV-positive gastric LELC cases and ordinary EBVaGC cases, respectively. EBV-positive gastric LELC exhibited a longer overall survival (p < 0.05, Fig. 2).
Overall survival curves for EBV-positive gastric LELC and ordinary EBVaGC patients. Thirty-six gastric carcinoma cases, including eight EBV-positive gastric LELC cases and 32 ordinary EBVaGC cases, were performed for survival analysis using Kaplan–Meier method. The EBV-positive gastric LELC group has a favorable overall survival (p = 0.035, Log-rank test)
Discussion
Gastric LELC represents a distinct clinicopathological entity whose morphology is similar to NPC, but it occurs outside of the nasopharynx [1–3]. In the present study, ten LELC cases were screened among 702 consecutive gastric carcinoma cases from Guangzhou. The frequency was 1.42 %, which was consistent with those reported in the literature (1.1–4.6 %) [8, 10, 14, 26]. Eight (80 %) out of the ten gastric LELC cases were EBV-positive. They showed a male preponderance (7:1), predisposition to the proximal part of the stomach and gastric stump and a relative favorable prognosis. They expressed EBNA1 (7/8, 87.5 %) and LMP2A (5/8, 62.5 %) but not EBNA2, LMP1 and ZEBRA. In EBV genotype analysis, type A strain, type F, I, mut-W1/I, XhoI− and del-LMP1 variants were predominant among EBV-positive gastric LELCs, accounting for eight (100 %), six (75 %), eight (100 %), seven (87.5 %), five (62.5 %) and six (75 %) cases, respectively, which are similar to those in ordinary EBVaGC in Guangzhou. For EBNA1 polymorphisms, the V-leu and P-ala subtypes were predominant in EBV-positive gastric LELC, accounting for 71.4 % (5/7) and 57.1 % (4/7), respectively, which is different from the predominant V-val subtype in ordinary EBVaGC in Guangzhou.
As the reliable and highly sensitive method for detection of latent EBV in paraffin sections, EBER-1 ISH assay was performed in LELCs and ordinary adenocarcinomas of the stomach. Eight of ten LELC cases and 45 of 692 ordinary adenocarcinoma cases had positive signals in the nuclei of almost all tumor cells, which indicated that these cases are EBV-associated. It is reported that gastric LELC consists of two subsets, EBV-positive and MSI-high ones [13]. In the present study, all eight EBV-positive gastric LELCs showed positivity of MLH1, indicating that they are not MSI-high gastric carcinomas; while the other two EBV-negative gastric LELCs showed loss expression of MLH1, suggesting that they are MSI-high gastric carcinomas. This is consistent with previous studies showing that EBV positivity and MSI-high status are mutually exclusive in gastric carcinomas [13]. The prevalence of EBV in gastric LELC is apparently higher than that in ordinary gastric adenocarcinoma (80 vs. 6.5 %). This is consistent with the previous studies [34–36], which proves a strong association between EBV and gastric LELC. However, like ordinary EBVaGCs, EBV-positive gastric LELCs show a male preponderance (7:1), and predisposition to the proximal part of the stomach and gastric stump. Besides, the prognosis of EBV-positive gastric LELC is relatively good. In our series, the median survival time of EBV-positive gastric LELC was 43.0 months, which is longer than that (18.0 months) of ordinary EBVaGC. It is known that gastric LELC always shows abundant lymphocytic infiltration. Thus, the relative favorable survival may be due to the heavy lymphoid infiltrate in gastric LELC. Further studies are needed to elucidate the relationship among EBV infection, abundant lymphocytic infiltration and relatively good survival in gastric LELC.
In the present study, both EBV-positive gastric LELCs and ordinary EBVaGCs expressed EBNA1 rather than LMP1, EBNA2 and ZEBRA, suggesting that the EBV latency pattern in EBV-positive gastric LELC and ordinary EBVaGC was type I, which is similar to that in BL [37], but different from that in NPC [38, 39]. However, LMP2A was detected in 62.5 % (5/8) EBV-positive gastric LELC cases and 53.3 % (24/45) ordinary EBVaGC cases by immunohistochemistry. Zur et al. [40], Sugiura et al. [41] and Luo et al. [42] also showed that LMP2A mRNA was detected in seven of nine (77.8 %), three of seven (42.9 %) and four of 11 (36.4 %) EBVaGC cases, respectively. These findings indicate that LMP2A is expressed in about half of the EBVaGC cases, which is different from that in BL [37].
Genotyping of EBV displayed that the subtype A strain, prototype F, type I, wt-W1/I1, XhoI− and del-LMP1 variants were predominant among the isolates from EBV-positive gastric LELCs. The findings were similar to those in ordinary EBVaGC. The predominant prototype F variant in EBV-positive gastric LELC and ordinary EBVaGC was significantly different from that found in NPC (predominant ‘‘f’’ variant) in the same area [43, 44]. Additionally, the mut-W1/I1 EBV variants that harbored a newly identified mutation in the BamHI-W1/I1 boundary region (148,972 T → C) were also predominant among EBV-positive gastric LELCs and ordinary EBVaGCs. However, the mutation could not be found in the NPC-derived strain GD1 [45]. These findings suggested that some EBV sequence variations may be disease restricted and the pathogenic mechanisms of EBV in EBVaGC and NPC may be distinct. In our previous study, we found that the predominant variants harbored in throat washings (TWs) of healthy EBV carriers in this area were prototype F and type wt-W1/I1, which were different from that in NPC (predominant “f” variant) and that in EBVaGC (predominant mut-W1/I1 variant) [46]. These findings seem to suggest that the same virus infects the population, then different mutants develop within the patients and result in different types of tumors. This, to some extent, might be another piece of evidence supporting that the pathogenic mechanisms of EBV in EBVaGC and NPC may be distinct. On the other hand, the combination of I/XhoI− was found in five (62.5 %) of eight EBV-positive gastric LELC cases and 34 (75.6 %) of 45 ordinary EBVaGC cases. These findings were similar to those in EBVaGC in eastern countries [47, 48], but different from those in EBVaGC in Latin American countries [25], suggesting that some EBV sequence variations may be geographically distributed.
For EBNA1 polymorphisms, the V-leu and P-ala subtypes were predominant in EBV-positive gastric LELC, accounting for 71.4 % (5/7) and 57.1 % (4/7), respectively. This is different from that in ordinary EBVaGC, in which V-val subtype was predominant (13/25, 52 %). As we known, the most important distinction feature of EBV-positive gastric LELC from ordinary EBVaGC is that EBV-positive gastric LELC always shows dense lymphocytic infiltration in the tumor [13, 19, 20]. Whether the predominant V-leu and P-ala subtypes in EBV-positive gastric LELC are associated with the dense infiltration of lymphocytes in the tumor needs to be further investigated.
It is interesting to find that some tumor samples in the present study displayed dual EBNA1 sequences, suggesting co-existence of two subtypes of EBNA1 within one tumor. Although we could not fully exclude the possibility that the one subtype is derived from the cancer, and the other from the infiltrating lymphocytes, this finding may also suggest that EBV subtypes in EBVaGC are generated in vivo from a single infecting virus subtype by mutations in the EBNA1 gene. In nasal NK/T-cell lymphomas, Gutiérrez et al. also found that only six of the 39 nasal NK/T-cell lymphomas studied carried a single EBNA1 sequence. The remaining 33 cases carried dual EBNA1 sequences [49]. In their report, they found that although nasal NK/T-cell lymphomas carried a mixture of EBNA1 subtypes, the EBV in the tumors was still monoclonal. Moreover, they showed evidence that EBV subtype present in most nasal lymphomas undergoes and accumulates multiple mutations consistent with the generation of variant species of EBNA1 in vivo. This might also be true for EBV in EBVaGC. However, of course, this needs further studies and will be the topic of a future research project.
As for LMP2A, 43 nucleotide mutations were found in the sequences of exons 1–8 in total. The distribution of nucleotide variations of LMP2A in EBV-positive gastric LELC and ordinary EBVaGC did not differ significantly (all p > 0.05, Table S6). These nucleotide mutations were mainly distributed in the transmembrane region (exons 2–8) of LMP2A, which possesses 17 target epitopes of cytotoxic T lymphocytes (CTLs) [50–54]. In the present study, eight out of the 17 identified CTL epitopes were affected by at least one point mutation. As we know, CTLs are very important in controlling the long-term persistence of EBV infection, and mutation of the targets would be expected to affect the CTL response to cells expressing LMP2A and result in a growth advantage of the cells in the host [55]. Therefore, the presence of CTL epitopes mutation may reduce or abrogate CTL responses, and confer an advantage on EBV for immune escape and virus persistence in tumor cells, thus playing a role in tumorigenesis and tumor maintenance [55]. Besides, the CTL epitopes mutations in LMP2A may also implicate that the effect of LMP2A polymorphisms should be considered when LMP2A-targeted immunotherapy is conducted.
In conclusion, the present study not only described the frequency and clinicopathological characteristics of gastric LELC, but also for the first time in the world systematically revealed the EBV latency pattern and genome polymorphisms of EBV-positive gastric LELC. The clinicopathological characteristics, EBV latency pattern and genome polymorphisms of EBV-positive gastric LELC were mostly similar to those of ordinary EBVaGC. However, the predominant EBNA1 subtype(s) in EBV-positive gastric LELC was different from that in ordinary EBVaGC. In addition, EBV-positive gastric LELC always shows dense lymphocytic infiltration and generally has a favorable prognosis. Therefore, EBV-positive gastric LELC should be regarded as a special subtype of EBVaGC.
References
Iezzoni JC, Gaffey MJ, Weiss LM. The role of Epstein–Barr virus in lymphoepithelioma-like carcinomas. Am J Clin Pathol. 1995;103:308–15.
Anagnostopoulos I, Hummel M. Epstein–Barr virus in tumours. Histopathology. 1996;29:297–315.
Weiss LM, Gaffey MJ, Shibata D. Lymphoepithelioma-like carcinoma and its relationship to Epstein–Barr virus. Am J Clin Pathol. 1991;96:156–8.
Herrmann K, Niedobitek G. Epstein–Barr virus-associated carcinomas: facts and fiction. J Pathol. 2003;199:140–5.
Han AJ, Xiong M, Gu YY, Lin SX, Xiong M. Lymphoepithelioma-like carcinoma of the lung with a better prognosis. A clinicopathologic study of 32 cases. Am J Clin Pathol. 2001;115:841–50.
Chetty R. Gastrointestinal cancers accompanied by a dense lymphoid component: an overview with special reference to gastric and colonic medullary and lymphoepithelioma-like carcinomas. J Clin Pathol. 2012;65:1062–5.
Watanabe H, Enjoji M, Imai T. Gastric carcinoma with lymphoid stroma. Its morphologic characteristics and prognostic correlations. Cancer. 1976;38:232–43.
Shibata D, Tokunaga M, Uemura Y, Sato E, Tanaka S, Weiss LM. Association of Epstein–Barr virus with undifferentiated gastric carcinomas with intense lymphoid infiltration. Lymphoepithelioma-like carcinoma. Am J Pathol. 1991;139:469–74.
Shibata D, Weiss LM. Epstein–Barr virus-associated gastric adenocarcinoma. Am J Pathol. 1992;140:769–74.
Nakamura S, Ueki T, Yao T, Ueyama T, Tsuneyoshi M. Epstein–Barr virus in gastric carcinoma with lymphoid stroma. Special reference to its detection by the polymerase chain reaction and in situ hybridization in 99 tumors, including a morphologic analysis. Cancer. 1994;73:2239–49.
Kijima Y, Ishigami S, Hokita S, Koriyama C, Akiba S, Eizuru Y, et al. The comparison of the prognosis between Epstein–Barr virus (EBV)-positive gastric carcinomas and EBV-negative ones. Cancer Lett. 2003;200:33–40.
Adachi Y, Yoh R, Konishi J, Iso Y, Matsumata T, Kasai T, et al. Epstein–Barr virus-associated gastric carcinoma. J Clin Gastroenterol. 1996;23:207–10.
Grogg KL, Lohse CM, Pankratz VS, Halling KC, Smyrk TC. Lymphocyte-rich gastric cancer: associations with Epstein–Barr virus, microsatellite instability, histology, and survival. Mod Pathol. 2003;16:641–51.
Oda K, Tamaru J, Takenouchi T, Mikata A, Nunomura M, Saitoh N, et al. Association of Epstein–Barr virus with gastric carcinoma with lymphoid stroma. Am J Pathol. 1993;143:1063–71.
Lee JH, Park SJ, Abraham SC, Seo JS, Nam JH, Choi C, et al. Frequent CpG island methylation in precursor lesions and early gastric adenocarcinomas. Oncogene. 2004;23:4646–54.
Thompson MP, Kurzrock R. Epstein–Barr virus and cancer. Clin Cancer Res. 2004;10:803–21.
Young LS, Rickinson AB. Epstein–Barr virus: 40 years on. Nat Rev Cancer. 2004;4:757–68.
International Agency for Research on Cancer, World Health Organization. Epstein–Barr virus: summary of data reported and evaluation. In: IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. Vol 70. Lyon: IARC Press; 1997. pp. 255–62.
Matsunou H, Konishi F, Hori H, Ikeda T, Sasaki K, Hirose Y, et al. Characteristics of Epstein–Barr virus-associated gastric carcinoma with lymphoid stroma in Japan. Cancer. 1996;77:1998–2004.
Minamoto T, Mai M, Watanabe K, Ooi A, Kitamura T, Takahashi Y, et al. Medullary carcinoma with lymphocytic infiltration of the stomach. Clinicopathologic study of 27 cases and immunohistochemical analysis of the subpopulations of infiltrating lymphocytes in the tumor. Cancer. 1990;66:945–52.
Chen FF, Yan JJ, Lai WW, Jin YT, Su IJ. Epstein–Barr virus-associated nonsmall cell lung carcinoma: undifferentiated “lymphoepithelioma-like” carcinoma as a distinct entity with better prognosis. Cancer. 1998;82:2334–42.
Leung SY, Chung LP, Yuen ST, Ho CM, Wong MP, Chan SY. Lymphoepithelial carcinoma of the salivary gland: in situ detection of Epstein–Barr virus. J Clin Pathol. 1995;48:1022–7.
Kuo T, Hsueh C. Lymphoepithelioma-like salivary gland carcinoma in Taiwan: a clinicopathological study of nine cases demonstrating a strong association with Epstein–Barr virus. Histopathology. 1997;31:75–82.
Dimery IW, Lee JS, Blick M, Pearson G, Spitzer G, Hong WK. Association of the Epstein–Barr virus with lymphoepithelioma of the thymus. Cancer. 1988;61:2475–80.
Corvalan A, Ding S, Koriyama C, Carrascal E, Carrasquilla G, Backhouse C, et al. Association of a distinctive strain of Epstein–Barr virus with gastric cancer. Int J Cancer. 2006;118:1736–42.
Corvalan A, Koriyama C, Akiba S, Eizuru Y, Backhouse C, Palma M, et al. Epstein–Barr virus in gastric carcinoma is associated with location in the cardia and with a diffuse histology: a study in one area of Chile. Int J Cancer. 2001;94:527–30.
Fukayama M, Ushiku T. Epstein–Barr virus-associated gastric carcinoma. Pathol Res Pract. 2011;207:529–37.
Young LS, Murray PG. Epstein–Barr virus and oncogenesis: from latent genes to tumours. Oncogene. 2003;22:5108–21.
Chen JN, He D, Tang F, Shao CK. Epstein–Barr virus-associated gastric carcinoma: a newly defined entity. J Clin Gastroenterol. 2012;46:262–71.
Bhatia K, Raj A, Guitierrez MI, Judde JG, Spangler G, Venkatesh H, et al. Variation in the sequence of Epstein Barr virus nuclear antigen 1 in normal peripheral blood lymphocytes and in Burkitt’s lymphomas. Oncogene. 1996;13:177–81.
Gutierrez MI, Raj A, Spangler G, Sharma A, Hussain A, Judde JG, et al. Sequence variations in EBNA-1 may dictate restriction of tissue distribution of Epstein–Barr virus in normal and tumour cells. J Gen Virol. 1997;78(Pt 7):1663–70.
Japanese Gastric Cancer Association. Japanese classification of gastric carcinoma—2nd English edition. Gastric Cancer. 1998;1:10–24.
Kononen J, Bubendorf L, Kallioniemi A, Barlund M, Schraml P, Leighton S, et al. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med. 1998;4:844–7.
Herath CH, Chetty R. Epstein–Barr virus-associated lymphoepithelioma-like gastric carcinoma. Arch Pathol Lab Med. 2008;132:706–9.
Fukayama M, Hino R, Uozaki H. Epstein–Barr virus and gastric carcinoma: virus-host interactions leading to carcinoma. Cancer Sci. 2008;99:1726–33.
Ott G, Kirchner T, Muller-Hermelink HK. Monoclonal Epstein–Barr virus genomes but lack of EBV-related protein expression in different types of gastric carcinoma. Histopathology. 1994;25:323–9.
Rowe M, Rowe DT, Gregory CD, Young LS, Farrell PJ, Rupani H, et al. Differences in B cell growth phenotype reflect novel patterns of Epstein–Barr virus latent gene expression in Burkitt’s lymphoma cells. EMBO J. 1987;6:2743–51.
Fahraeus R, Fu HL, Ernberg I, Finke J, Rowe M, Klein G, et al. Expression of Epstein–Barr virus-encoded proteins in nasopharyngeal carcinoma. Int J Cancer. 1988;42:329–38.
Young LS, Dawson CW, Clark D, Rupani H, Busson P, Tursz T, et al. Epstein–Barr virus gene expression in nasopharyngeal carcinoma. J Gen Virol. 1988;69(Pt 5):1051–65.
Zur HA, Brink AA, Craanen ME, Middeldorp JM, Meijer CJ, van den Brule AJ. Unique transcription pattern of Epstein–Barr virus (EBV) in EBV-carrying gastric adenocarcinomas: expression of the transforming BARF1 gene. Cancer Res. 2000;60:2745–8.
Sugiura M, Imai S, Tokunaga M, Koizumi S, Uchizawa M, Okamoto K, et al. Transcriptional analysis of Epstein–Barr virus gene expression in EBV-positive gastric carcinoma: unique viral latency in the tumour cells. Br J Cancer. 1996;74:625–31.
Luo B, Wang Y, Wang XF, Liang H, Yan LP, Huang BH, et al. Expression of Epstein–Barr virus genes in EBV-associated gastric carcinomas. World J Gastroenterol. 2005;11:629–33.
Lung ML, Lam WP, Sham J, Choy D, Yong-Sheng Z, Guo HY, et al. Detection and prevalence of the “f” variant of Epstein–Barr virus in southern China. Virology. 1991;185:67–71.
Lung ML, Chang RS, Huang ML, Guo HY, Choy D, Sham J, et al. Epstein–Barr virus genotypes associated with nasopharyngeal carcinoma in southern China. Virology. 1990;177:44–53.
Zeng MS, Li DJ, Liu QL, Song LB, Li MZ, Zhang RH, et al. Genomic sequence analysis of Epstein–Barr virus strain GD1 from a nasopharyngeal carcinoma patient. J Virol. 2005;79:15323–30.
Chen JN, Jiang Y, Li HG, Ding YG, Fan XJ, Xiao L, et al. Epstein–Barr virus genome polymorphisms of Epstein–Barr virus-associated gastric carcinoma in gastric remnant carcinoma in Guangzhou, southern China, an endemic area of nasopharyngeal carcinoma. Virus Res. 2011;160:191–9.
Sidagis J, Ueno K, Tokunaga M, Ohyama M, Eizuru Y. Molecular epidemiology of Epstein–Barr virus (EBV) in EBV-related malignancies. Int J Cancer. 1997;72:72–6.
Fukayama M, Hayashi Y, Iwasaki Y, Chong J, Ooba T, Takizawa T, et al. Epstein–Barr virus-associated gastric carcinoma and Epstein–Barr virus infection of the stomach. Lab Invest. 1994;71:73–81.
Gutierrez MI, Spangler G, Kingma D, Raffeld M, Guerrero I, Misad O, et al. Epstein–Barr virus in nasal lymphomas contains multiple ongoing mutations in the EBNA-1 gene. Blood. 1998;92:600–6.
Lee SP, Tierney RJ, Thomas WA, Brooks JM, Rickinson AB. Conserved CTL epitopes within EBV latent membrane protein 2: a potential target for CTL-based tumor therapy. J Immunol. 1997;158:3325–34.
Redchenko IV, Rickinson AB. Accessing Epstein–Barr virus-specific T-cell memory with peptide-loaded dendritic cells. J Virol. 1999;73:334–42.
Meij P, Leen A, Rickinson AB, Verkoeijen S, Vervoort MB, Bloemena E, et al. Identification and prevalence of CD8(+) T-cell responses directed against Epstein–Barr virus-encoded latent membrane protein 1 and latent membrane protein 2. Int J Cancer. 2002;99:93–9.
Whitney BM, Chan AT, Rickinson AB, Lee SP, Lin CK, Johnson PJ. Frequency of Epstein–Barr virus-specific cytotoxic T lymphocytes in the blood of Southern Chinese blood donors and nasopharyngeal carcinoma patients. J Med Virol. 2002;67:359–63.
Wang B, Yao K, Liu G, Xie F, Zhou F, Chen Y. Computational prediction and identification of Epstein–Barr virus latent membrane protein 2A antigen-specific CD8+ T-cell epitopes. Cell Mol Immunol. 2009;6:97–103.
Tanaka M, Kawaguchi Y, Yokofujita J, Takagi M, Eishi Y, Hirai K. Sequence variations of Epstein–Barr virus LMP2A gene in gastric carcinoma in Japan. Virus Genes. 1999;19:103–11.
Acknowledgments
We appreciate Prof. Mu-sheng Zeng from the Cancer Center of Sun Yat-sen University for his generous gift of the LMP2A monoclonal antibody. This work was supported by the National Natural Science Foundation of China (Nos. 81071893, 81272553 and 81301694), the Guangdong Natural Science Foundation (No. S2012020010898) and the Guangzhou Science and Technology Project (No. 2011J4100106), Guangdong Province, China.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding authors
Additional information
N. Cheng and D. Hui contributed equally to this work.
Electronic supplementary material
Below are links to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cheng, N., Hui, Dy., Liu, Y. et al. Is gastric lymphoepithelioma-like carcinoma a special subtype of EBV-associated gastric carcinoma? New insight based on clinicopathological features and EBV genome polymorphisms. Gastric Cancer 18, 246–255 (2015). https://doi.org/10.1007/s10120-014-0376-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-014-0376-9