Abstract
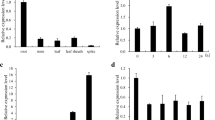
The expression of the six rice ASR genes is differentially regulated in a tissue-dependent manner according to environmental conditions and reproductive stages. OsASR1 and OsASR3 are the most abundant and are found in most tissues; they are enriched in the leaves and roots, respectively. Coexpression analysis of OsASR1 and OsASR3 and a comparison of the cis-acting elements upstream of OsASR1 and OsASR3 suggested that their expression is regulated in common by abiotic stresses but differently regulated by hormone and sugar signals. The results of quantitative real-time PCR analyses of OsASR1 and OsASR3 expression under various conditions further support this model. The expression of both OsASR1 and OsASR3 was induced by drought stress, which is a major regulator of the expression of all ASR genes in rice. In contrast, ABA is not a common regulator of the expression of these genes. OsASR1 transcription was highly induced by ABA, whereas OsASR3 transcription was strongly induced by GA. In addition, OsASR1 and OsASR3 expression was significantly induced by sucrose and sucrose/glucose treatments, respectively. The induction of gene expression in response to these specific hormone and sugar signals was primarily observed in the major target tissues of these genes (i.e., OsASR1 in leaves and OsASR3 in roots). Our data also showed that the overexpression of either OsASR1 or OsASR3 in transgenic rice plants increased their tolerance to drought and cold stress. Taken together, our results revealed that the transcriptional control of different rice ASR genes exhibit different tissue-dependent sugar and hormone-sensitivities.
Similar content being viewed by others
References
Abe, H., Urao, T., Ito, T., Seki, M., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2003). Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15, 63–78.
Amitai-Zeigerson, H., Scolnik, P.A., and Bar-Zvi, D. (1994). Genomic nucleotide sequence of tomato Asr2, a second member of the stress/ripening-induced Asr1 gene family. Plant Physiol. 106, 1699–1700.
Arenhart, R.A., Lima, J.C., Pedron, M., Carvalho, F.E., Silveira, J.A., Rosa, S.B., Caverzan, A., Andrade, C.M., Schunemann, M., Margis, R., et al. (2013). Involvement of ASR genes in aluminium tolerance mechanisms in rice. Plant Cell Environ. 36, 52–67.
Artus, N.N., Uemura, M., Steponkus, P.L., Gilmour, S.J., Lin, C., and Thomashow, M.F. (1996). Constitutive expression of the cold-regulated Arabidopsis thaliana COR15a gene affects both chloroplast and protoplast freezing tolerance. Proc. Natl. Acad. Sci. USA 93, 13404–13409.
Blackwell, T.K., Bowerman, B., Priess, J.R., and Weintraub, H. (1994). Formation of a monomeric DNA binding domain by Skn-1 bZIP and homeodomain elements. Science 266, 621–628.
Boyer, J.S. (1982). Plant productivity and environment. Science 218, 443–448.
Cakir, B., Agasse, A., Gaillard, C., Saumonneau, A., Delrot, S., and Atanassova, R. (2003). A grape ASR protein involved in sugar and abscisic acid signaling. Plant Cell 15, 2165–2180.
Canel, C., Bailey-Serres, J.N., and Roose, M.L. (1995). Pummelo fruit transcript homologous to ripening-induced genes. Plant Physiol. 108, 1323–1324.
Chen, W., Provart, N.J., Glazebrook, J., Katagiri, F., Chang, H.S., Eulgem, T., Mauch, F., Luan, S., Zou, G., Whitham, S.A., et al. (2002). Expression profile matrix of Arabidopsis transcription factor genes suggests their putative functions in response to environmental stresses. Plant Cell 14, 559–574.
Chen, P.W., Chiang, C.M., Tseng, T.H., and Yu, S.M. (2006). Interaction between rice MYBGA and the gibberellin response element controls tissue-specific sugar sensitivity of alpha-amylase genes. Plant Cell 18, 2326–2340.
Chen, J.Y., Liu, D.J., Jiang, Y.M., Zhao, M.L., Shan, W., Kuang, J.F., and Lu, W.J. (2011). Molecular characterization of a strawberry FaASR gene in relation to fruit ripening. PLoS One 6, e24649.
Dai, J.R., Liu, B., Feng, D.R., Liu, H.Y., He, Y.M., Qi, K.B., Wang, H.B., and Wang, J.F. (2011). MpAsr encodes an intrinsically unstructured protein and enhances osmotic tolerance in transgenic Arabidopsis. Plant Cell Rep. 30, 1219–1230.
Dominguez, P.G., Frankel, N., Mazuch, J., Balbo, I., Iusem, N.D., Fernie, A.R., and Carrari, F. (2013). Asr1 mediates glucose-hormone crosstalk by affecting sugar trafficking in tobacco plants. Plant Physiol. 161, 1486–1500.
Dubouzet, J.G., Sakuma, Y., Ito, Y., Kasuga, M., Dubouzet, E.G., Miura, S., Seki, M., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2003). OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and coldresponsive gene expression. Plant J. 33, 751–763.
Eulgem, T., Rushton, P.J., Robatzek, S., and Somssich, I.E. (2000). The WRKY superfamily of plant transcription factors. Trends Plant Sci. 5, 199–206.
Ezcurra, I., Ellerstrom, M., Wycliffe, P., Stalberg, K., and Rask, L. (1999). Interaction between composite elements in the napA promoter: both the B-box ABA-responsive complex and the RY/G complex are necessary for seed-specific expression. Plant Mol. Biol. 40, 699–709.
Fillion, L., Ageorges, A., Picaud, S., Coutos-Thevenot, P., Lemoine, R., Romieu, C., and Delrot, S. (1999). Cloning and expression of a hexose transporter gene expressed during the ripening of grape berry. Plant Physiol. 120, 1083–1094.
Frankel, N., Carrari, F., Hasson, E., and Iusem, N.D. (2006). Evolutionary history of the Asr gene family. Gene 378, 74–83.
Frankel, N., Nunes-Nesi, A., Balbo, I., Mazuch, J., Centeno, D., Iusem, N.D., Fernie, A.R., and Carrari, F. (2007). ci21A/Asr1 expression influences glucose accumulation in potato tubers. Plant Mol. Biol. 63, 719–730.
Gilad, A., Amitai-Zeigerson, H., Scolnik, P.A., and Bar-Zvi, D. (1997). Asr1, a tomato water-stress regulated gene: genomic organization, developmental regulation and DNA-binding activity. Acta Hort. 447, 447–453.
Goldgur, Y., Rom, S., Ghirlando, R., Shkolnik, D., Shadrin, N., Konrad, Z., and Bar-Zvi, D. (2007). Desiccation and zinc binding induce transition of tomato abscisic acid stress ripening 1, a water stress- and salt stress-regulated plant-specific protein, from unfolded to folded state. Plant Physiol. 143, 617–628.
Grierson, C., Du, J.S., de Torres Zabala, M., Beggs, K., Smith, C., Holdsworth, M., and Bevan, M. (1994). Separate cis sequences and trans factors direct metabolic and developmental regulation of a potato tuber storage protein gene. Plant J. 5, 815–826.
Gubler, F., and Jacobsen, J.V. (1992). Gibberellin-responsive elements in the promoter of a barley high-pI alpha-amylase gene. Plant Cell 4, 1435–1441.
Gubler, F., Raventos, D., Keys, M., Watts, R., Mundy, J., and Jacobsen, J.V. (1999). Target genes and regulatory domains of the GAMYB transcriptional activator in cereal aleurone. Plant J. 17, 1–9.
Hirasawa, T. (1999). Physiological characterization of rice plant for tolerance of water deficit. In Genetic Improvement of Rice for Water-Limited Environments, O. Ito, J.C., O’Toole, and B. Hardy, eds. (Los Baños, Philippines: International Rice Research Institute), pp. 89–98.
Hong, S.H., Kim, I.J., Yang, D.C., and Chung, W.I. (2002). Characterization of an abscisic acid responsive gene homologue from Cucumis melo. J. Exp. Bot. 53, 2271–2272.
Hsu, Y.F., Yu, S.C., Yang, C.Y., and Wang, C.S. (2011). Lily ASR protein-conferred cold and freezing resistance in Arabidopsis. Plant Physiol. Biochem. 49, 937–945.
Hu, W., Huang, C., Deng, X., Zhou, S., Chen, L., Li, Y., Wang, C., Ma, Z., Yuan, Q., Wang, Y., et al. (2013). TaASR1, a transcription factor gene in wheat, confers drought stress tolerance in transgenic tobacco. Plant Cell Environ. doi: 10.1111/pce.12074. [Epub ahead of print].
Huang, J.C., Lin, S.M., and Wang, C.S. (2000). A pollen-specific and desiccation-associated transcript in Lilium longiflorum during development and stress. Plant Cell Physiol. 41, 477–485.
Hwang, Y.S., Karrer, E.E., Thomas, B.R., Chen, L., and Rodriguez, R.L. (1998). Three cis-elements required for rice alpha-amylase Amy3D expression during sugar starvation. Plant Mol. Biol. 36, 331–341.
Ingram, J., and Bartels, D. (1996). The molecular basis of dehydration tolerance in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 47, 377–403.
Itoh, J., Nonomura, K., Ikeda, K., Yamaki, S., Inukai, Y., Yamagishi, H., Kitano, H., and Nagato, Y. (2005). Rice plant development: from zygote to spikelet. Plant Cell Physiol. 46, 23–47.
Iusem, N.D., Bartholomew, D.M., Hitz, W.D., and Scolnik, P.A. (1993). Tomato (Lycopersicon esculentum) transcript induced by water deficit and ripening. Plant Physiol. 102, 1353–1354.
Jain, M., Nijhawan, A., Arora, R., Agarwal, P., Ray, S., Sharma, P., Kapoor, S., Tyagi, A.K., and Khurana, J.P. (2007). F-box proteins in rice. Genome-wide analysis, classification, temporal and spatial gene expression during panicle and seed development, and regulation by light and abiotic stress. Plant Physiol. 143, 1467–1483.
Jang, I.C., Choi, W.B., Lee, K.H., Song, S.I., Nahm, B.H., and Kim, J.K. (2002). High-level and ubiquitous expression of the rice cytochrome c gene OsCc1 and its promoter activity in transgenic plants provides a useful promoter for transgenesis of monocots. Plant Physiol. 129, 1473–1481.
Jeanneau, M., Gerentes, D., Foueillassar, X., Zivy, M., Vidal, J., Toppan, A., and Perez, P. (2002). Improvement of drought tolerance in maize: towards the functional validation of the Zm-Asr1 gene and increase of water use efficiency by overexpressing C4-PEPC. Biochimie 84, 1127–1135.
Kagaya, Y., Ohmiya, K., and Hattori, T. (1999). RAV1, a novel DNA-binding protein, binds to bipartite recognition sequence through two distinct DNA-binding domains uniquely found in higher plants. Nucleic Acids Res. 27, 470–478.
Kalifa, Y., Gilad, A., Konrad, Z., Zaccai, M., Scolnik, P.A., and Bar-Zvi, D. (2004a). The water- and salt-stress-regulated Asr1 (abscisic acid stress ripening) gene encodes a zinc-dependent DNA-binding protein. Biochem. J. 381, 373–378.
Kalifa, Y., Perlson, E., Gilad, A., Konrad, Z., Scolnik, P.A., and Bar-Zvi, D. (2004b). Over-expression of the water and salt stressregulated Asr1 gene confers an increased salt tolerance. Plant Cell Environ. 27, 1459–1468.
Kang, J.Y., Choi, H.I., Im, M.Y., and Kim, S.Y. (2002). Arabidopsis basic leucine zipper proteins that mediate stress-responsive abscisic acid signaling. Plant Cell 14, 343–357.
Kaplan, B., Davydov, O., Knight, H., Galon, Y., Knight, M.R., Fluhr, R., and Fromm, H. (2006). Rapid transcriptome changes induced by cytosolic Ca2+ transients reveal ABRE-related sequences as Ca2+-responsive cis elements in Arabidopsis. Plant Cell 18, 2733–2748.
Kawasaki, S., Borchert, C., Deyholos, M., Wang, H., Brazille, S., Kawai, K., Galbraith, D., and Bohnert, H.J. (2001). Gene expression profiles during the initial phase of salt stress in rice. Plant Cell 13, 889–905.
Kim, H.J., Kim, Y.K., Park, J.Y., and Kim, J. (2002). Light signalling mediated by phytochrome plays an important role in coldinduced gene expression through the C-repeat/dehydration responsive element (C/DRE) in Arabidopsis thaliana. Plant J. 29, 693–704.
Kim, S.J., Lee, S.C., Hong, S.K., An, K., An, G., and Kim, S.R. (2009). Ectopic expression of a cold-responsive OsAsr1 cDNA gives enhanced cold tolerance in transgenic rice plants. Mol. Cells 27, 449–458.
Kim, I.S., Kim, Y.S., and Yoon, H.S. (2012). Rice ASR1 protein with reactive oxygen species scavenging and chaperone-like activities enhances acquired tolerance to abiotic stresses in Saccharomyces cerevisiae. Mol. Cells 33, 285–293.
Konrad, Z., and Bar-Zvi, D. (2008). Synergism between the chaperone-like activity of the stress regulated ASR1 protein and the osmolyte glycine-betaine. Planta 227, 1213–1219.
Lanahan, M.B., Ho, T.H., Rogers, S.W., and Rogers, J.C. (1992). A gibberellin response complex in cereal alpha-amylase gene promoters. Plant Cell 4, 203–211.
Lata, C., and Prasad, M. (2011). Role of DREBs in regulation of abiotic stress responses in plants. J. Exp. Bot. 62, 4731–4748.
Lee, T.H., Kim, Y.K., Pham, T.T., Song, S.I., Kim, J.K., Kang, K.Y., An, G., Jung, K.H., Galbraith, D.W., Kim, M., et al. (2009). RiceArrayNet: a database for correlating gene expression from transcriptome profiling, and its application to the analysis of coexpressed genes in rice. Plant Physiol. 151, 16–33.
Liu, H.Y., Dai, J.R., Feng, D.R., Liu, B., Wang, H.B., and Wang, J.F. (2010). Characterization of a novel plantain Asr gene, MpAsr, that is regulated in response to infection of Fusarium oxysporum f. sp. cubense and abiotic stresses. J. Integr. Plant Biol. 52, 315–323.
Lopez-Ochoa, L., Acevedo-Hernandez, G., Martinez-Hernandez, A., Arguello-Astorga, G., and Herrera-Estrella, L. (2007). Structural relationships between diverse cis-acting elements are critical for the functional properties of a rbcS minimal light regulatory unit. J. Exp. Bot. 58, 4397–4406.
Maleck, K., Levine, A., Eulgem, T., Morgan, A., Schmid, J., Lawton, K.A., Dangl, J.L., and Dietrich, R.A. (2000). The transcriptome of Arabidopsis thaliana during systemic acquired resistance. Nat. Genet. 26, 403–410.
Maruyama-Nakashita, A., Nakamura, Y., Watanabe-Takahashi, A., Inoue, E., Yamaya, T., and Takahashi, H. (2005). Identification of a novel cis-acting element conferring sulfur deficiency response in Arabidopsis roots. Plant J. 42, 305–314.
Maskin, L., Gubesblat, G.E., Moreno, J.E., Carrari, F.O., Frankel, N., Sambade, A., Rossi, M., and Iusem, N.D. (2001). Differential expression of the members of the Asr gene family in tomato (Lycopersicon esculentum). Plant Sci. 161, 739–746.
Morikami, A., Matsunaga, R., Tanaka, Y., Suzuki, S., Mano, S., and Nakamura, K. (2005). Two cis-acting regulatory elements are involved in the sucrose-inducible expression of the sporamin gene promoter from sweet potato in transgenic tobacco. Mol. Genet. Genomics 272, 690–699.
Ogawa, M., Hanada, A., Yamauchi, Y., Kuwahara, A., Kamiya, Y., and Yamaguchi, S. (2003). Gibberellin biosynthesis and response during Arabidopsis seed germination. Plant Cell 15, 1591–1604.
Padmanabhan, V., Dias, D.M., and Newton, R.J. (1997). Expression analysis of a gene family in loblolly pine (Pinus taeda L.) induced by water deficit stress. Plant Mol. Biol. 35, 801–807.
Palm, C.J., Costa, M.A., An, G., and Ryan, C.A. (1990). Woundinducible nuclear protein binds DNA fragments that regulate a proteinase inhibitor II gene from potato. Proc. Natl. Acad. Sci. USA 87, 603–607.
Pantuwan, G., Fukai, S., Cooper, M., Rajatasereekul, S., and O’Toole, J.C.O. (2002). Yield response of rice (Oryza sativa L.) genotypes to drought under rain fed low land: 3. Plant factors contributing to drought resistance. Field Crops Res. 73, 181–200.
Phi-Van, L., and Stratling, W.H. (1996). Dissection of the ability of the chicken lysozyme gene 5′ matrix attachment region to stimulate transgene expression and to dampen position effects. Biochemistry 35, 10735–10742.
Philippe, R., Courtois, B., McNally, K.L., Mournet, P., El-Malki, R., Le Paslier, M.C., Fabre, D., Billot, C., Brunel, D., Glaszmann, J.C., et al. (2010). Structure, allelic diversity and selection of Asr genes, candidate for drought tolerance, in Oryza sativa L. and wild relatives. Theor. Appl. Genet. 121, 769–787.
Price, A., and Courtois, B. (1999). Mapping QTLs associated with drought resistance in rice: Progress, problems and prospects. Plant Growth Regulation 29, 123–133.
Ricardi, M.M., Guaimas, F.F., Gonzalez, R.M., Burrieza, H.P., Lopez-Fernandez, M.P., Jares-Erijman, E.A., Estevez, J.M., and Iusem, N.D. (2012). Nuclear import and dimerization of tomato ASR1, a water stress-inducible protein exclusive to plants. PLoS One 7, e41008.
Rom, S., Gilad, A., Kalifa, Y., Konrad, Z., Karpasas, M.M., Goldgur, Y., and Bar-Zvi, D. (2006). Mapping the DNA- and zinc-binding domains of ASR1 (abscisic acid stress ripening), an abioticstress regulated plant specific protein. Biochimie 88, 621–628.
Rossi, M., Carrari, F., Cabrera-Ponce, J.L., Vazquez-Rovere, C., Herrera-Estrella, L., Gudesblat, G., and Iusem, N.D. (1998). Analysis of an abscisic acid (ABA)-responsive gene promoter belonging to the Asr gene family from tomato in homologous and heterologous systems. Mol. Gen. Genet. 258, 1–8.
Saumonneau, A., Agasse, A., Bidoyen, M.T., Lallemand, M., Cantereau, A., Medici, A., Laloi, M., and Atanassova, R. (2008). Interaction of grape ASR proteins with a DREB transcription factor in the nucleus. FEBS Lett. 582, 3281–3287.
Shen, Q., Zhang, P., and Ho, T.H. (1996). Modular nature of abscisic acid (ABA) response complexes: composite promoter units that are necessary and sufficient for ABA induction of gene expression in barley. Plant Cell 8, 1107–1119.
Shinozaki, K., and Yamaguchi-Shinozaki, K. (1997). Gene expression and signal transduction in water-stress response. Plant Physiol. 115, 327–334.
Shinozaki, K., and Yamaguchi-Shinozaki, K. (2000). Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr. Opin. Plant Biol. 3, 217–223.
Shkolnik, D., and Bar-Zvi, D. (2008). Tomato ASR1 abrogates the response to abscisic acid and glucose in Arabidopsis by competing with ABI4 for DNA binding. Plant Biotechnol. J. 6, 368–378.
Silhavy, D., Hutvagner, G., Barta, E., and Banfalvi, Z. (1995). Isolation and characterization of a water-stress-inducible cDNA clone from Solanum chacoense. Plant Mol. Biol. 27, 587–595.
Simpson, S.D., Nakashima, K., Narusaka, Y., Seki, M., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2003). Two different novel cis-acting elements of erd1, a clpA homologous Arabidopsis gene function in induction by dehydration stress and darkinduced senescence. Plant J. 33, 259–270.
Sun, T.P., and Gubler, F. (2004). Molecular mechanism of gibberellin signaling in plants. Annu. Rev. Plant Biol. 55, 197–223.
Sutoh, K., and Yamauchi, D. (2003). Two cis-acting elements necessary and sufficient for gibberellin-upregulated proteinase expression in rice seeds. Plant J. 34, 635–645.
Takasaki, H., Mahmood, T., Matsuoka, M., Matsumoto, H., and Komatsu, S. (2008). Identification and characterization of a gibberellin-regulated protein, which is ASR5, in the basal region of rice leaf sheaths. Mol. Genet. Genomics 279, 359–370.
Toyofuku, K., Umemura, T., and Yamaguchi, J. (1998). Promoter elements required for sugar-repression of the RAmy3D gene for alpha-amylase in rice. FEBS Lett. 428, 275–280.
Ulmasov, T., Hagen, G., and Guilfoyle, T.J. (1999). Dimerization and DNA binding of auxin response factors. Plant J. 19, 309–319.
Uno, Y., Furihata, T., Abe, H., Yoshida, R., Shinozaki, K., and Yamaguchi-Shinozaki, K. (2000). Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc. Natl. Acad. Sci. USA 97, 11632–11637.
Vaidyanathan, R., Kuruvilla, S., and Thomas, G. (1999). Characterization and expression pattern of an abscisic acid and osmotic stress responsive gene from rice. Plant Sci. 140, 21–30.
Verslues, P.E., and Zhu, J.K. (2005). Before and beyond ABA: upstream sensing and internal signals that determine ABA accumulation and response under abiotic stress. Biochem. Soc. Trans. 33, 375–379.
Villain, P., Mache, R., and Zhou, D.X. (1996). The mechanism of GT element-mediated cell type-specific transcriptional control. J. Biol. Chem. 271, 32593–32598.
Virlouvet, L., Jacquemot, M.P., Gerentes, D., Corti, H., Bouton, S., Gilard, F., Valot, B., Trouverie, J., Tcherkez, G., Falque, M., et al. (2011). The ZmASR1 protein influences branched-chain amino acid biosynthesis and maintains kernel yield in maize under water-limited conditions. Plant Physiol. 157, 917–936.
Washio, K. (2003). Functional dissections between GAMYB and Dof transcription factors suggest a role for protein-protein associations in the gibberellin-mediated expression of the RAmy1A gene in the rice aleurone. Plant Physiol. 133, 850–863.
Wu, C., Washida, H., Onodera, Y., Harada, K., and Takaiwa, F. (2000). Quantitative nature of the Prolamin-box, ACGT and AACA motifs in a rice glutelin gene promoter: minimal cis-element requirements for endosperm-specific gene expression. Plant J. 23, 415–421.
Xue, G.P. (2002). An AP2 domain transcription factor HvCBF1 activates expression of cold-responsive genes in barley through interaction with a (G/a)(C/t)CGAC motif. Biochim. Biophys. Acta 1577, 63–72.
Yamagata, H., Yonesu, K., Hirata, A., and Aizono, Y. (2002) TGTCACA motif is a novel cis-regulatory enhancer element involved in fruit-specific expression of the cucumisin gene. J. Biol. Chem. 277, 11582–11590.
Yang, T., and Poovaiah, B.W. (2002). A calmodulin-binding/CGCG box DNA-binding protein family involved in multiple signaling pathways in plants. J. Biol. Chem. 277, 45049–45058.
Yang, L., Zheng, B., Mao, C., Qi, X., Liu, F., and Wu, P. (2004). Analysis of transcripts that are differentially expressed in three sectors of the rice root system under water deficit. Mol. Genet. Genomics 272, 433–442.
Yang, C.Y., Chen, Y.C., Jauh, G.Y., and Wang, C.S. (2005). A Lily ASR protein involves abscisic acid signaling and confers drought and salt resistance in Arabidopsis. Plant Physiol. 139, 836–846.
Yang, C.Y., Wu, C.H., Jauh, G.Y., Huang, J.C., Lin, C.C., and Wang, C.S. (2008). The LLA23 protein translocates into nuclei shortly before desiccation in developing pollen grains and regulates gene expression in Arabidopsis. Protoplasma 233, 241–254.
Zeeberg, B.R., Feng, W., Wang, G., Wang, M.D., Fojo, A.T., Sunshine, M., Narasimhan, S., Kane, D.W., Reinhold, W.C., Lababidi, S., et al. (2003). GoMiner: a resource for biological interpretation of genomic and proteomic data. Genome Biol. 4, R28.
Zhang, Z.L., Xie, Z., Zou, X., Casaretto, J., Ho, T.H., and Shen, Q.J. (2004). A rice WRKY gene encodes a transcriptional repressor of the gibberellin signaling pathway in aleurone cells. Plant Physiol. 134, 1500–1513.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Joo, J., Lee, Y.H., Kim, YK. et al. Abiotic stress responsive rice ASR1 and ASR3 exhibit different tissue-dependent sugar and hormone-sensitivities. Mol Cells 35, 421–435 (2013). https://doi.org/10.1007/s10059-013-0036-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10059-013-0036-7