Abstract
Soda lake sediments usually contain high concentrations of sulfide indicating active sulfate reduction. Monitoring of sulfate-reducing bacteria (SRB) in soda lakes demonstrated a dominance of two groups of culturable SRB belonging to the order Desulfovibrionales specialized in utilization of inorganic electron donors, such as formate, H2 and thiosulfate. The most interesting physiological trait of the novel haloalkaliphilic SRB isolates was their ability to grow lithotrophically by dismutation of thiosulfate and sulfite. All isolates were obligately alkaliphilic with a pH optimum at 9.5–10 and moderately salt tolerant. Among the fifteen newly isolated strains, four belonged to the genus Desulfonatronum and the others to the genus Desulfonatronovibrio. None of the isolates were closely related to previously described species of these genera. On the basis of phylogenetic, genotypic and phenotypic characterization of the novel soda lake SRB isolates, two novel species each in the genera Desulfonatronum and Desulfonatronovibrio are proposed.
Similar content being viewed by others
Introduction
Soda lakes are naturally occurring highly alkaline and saline habitats containing sodium carbonate at high concentrations ensuring a stable extremely high pH. Despite these extreme conditions, soda lakes are, usually, highly productive. The high primary productivity, often high sulfate concentrations and the repression of autotrophic methanogens by high salt concentrations (Ryu et al. 2004; Waldron et al. 2007) are at the basis of an active sulfur cycle in the soda lake sediments which frequently contain millimolar concentrations of free sulfide and FeS in the upper 20 cm profile. The oxidative part of the cycle is driven by well-studied aerobic chemolithoautotrophic and anoxygenic phototrophic haloalkaliphilic sulfide-oxidizing Gammaproteobacteria (Sorokin et al. 2006; Gorlenko 2007). In contrast, studies on the reductive part of the sulfur cycle in soda lakes are scarce and limited to measurements of sulfate reduction rates (Gorlenko et al. 1999; Sorokin et al. 2004; Kulp et al. 2006, 2007). The latter indicated relatively high rates of sulfate reduction at low to moderate salinity, while the actual presence of sulfate-reducing bacteria (SRB) was confirmed by the detection of the key functional gene of the dissimilatory sulfite reductase (dsrAB). Two of such investigations demonstrated the dominance of the order Desulfovibrionales sequences belonging to two genera, which were also obtained in pure culture from soda lakes, i.e., Desulfonatronum and Desulfonatronovibrio (Scholten et al. 2005; Foti et al. 2007, 2008). The total four characterized species in two genera are moderately salt-tolerant alkaliphiles with a very restricted substrate spectrum limited to H2 and formate for Desulfonatronovibrio (Zhilina et al. 1997) and H2, formate and ethanol/lactate for Desulfonatronum (Pikuta et al. 1998; 2003; Zhilina et al. 2005). Recent attempts to enrich SRB from hypersaline soda lakes at saturating soda concentrations resulted in the identification and description of a novel genus Desulfonatronospira (Sorokin et al. 2008), a member of the family Desulfohalobiaceae within the order Desulfovibrionales (Kuever et al. 2005). The organism is unique in its ability to grow chemolithoautotrophically in saturated soda brines.
Our systematic study of the sulfidogenesis in soda lakes demonstrated high rates of sulfate and thiosulfate reduction at moderate salinity which were further stimulated by the addition of formate, but not by the addition of H2 (Sorokin et al. 2010a). The microbiology work resulted in the isolation of 15 novel lithotrophic SRB strains from soda lake sediments growing optimally at a pH of 10. All of them belonged to the already known genera Desulfonatronum and Desulfonatronovibrio, but none were identical to the described species. In this paper, the properties of the novel lithotrophic SRB isolates are described and four novel species are proposed.
Methods
Samples
Sediment samples (2–20 cm depth) were obtained from 5 moderate and hypersaline soda lakes in Kulunda Steppe (south-eastern Siberia, Altai, Russia) in July 2007–2010 and from Owens hypersaline alkaline Lake (California) in 2008. The main brine properties of the lakes are given in Table 1.
Enrichment and cultivation of haloalkaliphilic SRB
Anaerobic enrichment and routine cultivation of SRB from soda lake sediments were performed at 30°C on a mineral base medium containing in total either 0.6 or 2 M of total Na+ and strongly buffered at pH 10: 22 (or 95) g l−1 of Na2CO3, 8 (or 15) g l−1 of NaHCO3, 6 (or 16) g l−1 of NaCl, and 1 g l−1 of K2HPO4. After sterilization, the medium was supplemented with 4 mM NH4Cl, 1 mM MgSO4, 20 mg l−1 of yeast extract and 1 ml l−1 of each solution of acidic trace metals and vitamins (Pfennig and Lippert 1966). Also, 1 ml of basic filter-sterilized Se/W solution (Widdel and Bak 1992) was supplied. In some cases, when a possibility of autotrophic growth was tested, yeast extract was omitted. Electron donors and acceptors were used at 20 mM concentrations (for sulfite the concentration was 10 mM). 1 mM HS− was added as a reductant. Routine cultivation was performed in 15 ml Hungate tubes with 10 ml medium made anoxic by several cycles of flushing with argon and evacuation. Growth was monitored by sulfide production. When the sulfide concentration in the enrichments exceeded 5 mM, the cultures were transferred into new medium at 1:100 dilution. After 2–3 successful transfers, the enrichments were serially diluted up to 10−11. Colonies were obtained only for the Desulfonatronum isolates. The alkaline solid medium was prepared by 1:1 mixing of the liquid alkaline medium with 4% (w/v) washed agar at 50°C. After cooling to 45°C, the medium was supplied with 2 mM HS−/100 μM dithionite, inoculated and poured into Petri dishes in an anaerobic chamber. After solidifying, the agar plates were placed into closed jars and incubated outside the chamber with oxygen-scavenging catalyzer (Oxoid). The final culture purity was checked by microscopy and the absence of growth on rich media without electron acceptors.
The pH dependence was examined at a Na+ content of 0.6 M, using the following filter-sterilized mineral media: for pH 6–8, 0.1 M HEPES and NaCl; for pH 8.5–11, a mixture of sodium bicarbonate/sodium carbonate containing 0.1 M NaCl. Growth resulted in a shift of initial pH values, especially at pH extremes (from 7 to 7.5, from 7.5 to 7.9, from 8 to 8.3, from 8.5 to 8.8, from 11 to 10.65, from 10.75 to 10.5, from 10.5 to 10.3). Therefore, final pH values were taken to indicate a suitable range for growth. To study the influence of salt concentration on growth, mineral sodium carbonate bases with pH 10 containing 0.1 and 3.0 M of total Na+ were mixed in different proportions.
Analyses
Sulfide was precipitated in 10% (w/v) Zn acetate and analyzed by the methylene blue method after separation from the supernatant (Trüper and Schlegel 1964). Thiosulfate and sulfite were determined after removal of ZnS by acidic iodimetric titration. The amount of cell proteins was measured by the Lowry method (Lowry et al. 1951) after removal of interfering FeS from the cell pellet by a double wash with 0.5 M NaCl, pH 4. Polar lipids were extracted from freeze-dried biomass by acidic methanol and their fatty acid composition examined with GC–MS according to Zhilina et al. (1997). Organic compatible solute composition in strain AHT9 grown at 3 M total Na+ was analyzed by 13C-NMR as described previously (Banciu et al. 2008). Phase contrast photomicrographs were obtained with a Zeiss Axioplan Imaging 2 microscope (Göttingen, Germany). For electron microscopy of total cells they were suspended in 0.1–0.5 M NaCl, fixed with glutaraldehyde (final 3% v/v) and contrasted with 1% (w/v) neutralized phospotungstic acid.
Genetic and phylogenetic analysis
Isolation of genomic DNA and determination of the G+C content of the DNA from pure cultures was performed according to Marmur (1961) and Marmur and Doty (1962). DNA–DNA hybridization was performed by thermal spectrophotometry according to De Ley et al. (1970). For molecular analysis, the DNA was extracted from the cells using the UltraClean Microbial DNA Isolation kit (MoBio Laboratories Inc., Carlsbad, CA, USA) following the manufacturer’s instructions. The nearly complete 16S rRNA gene was obtained from pure cultures using general bacterial primers 11f-1492r (Lane 1991). The PCR products were purified using the Qiagen Gel Extraction Kit (Qiagen, the Netherlands). The sequences were first compared to all sequences stored in GenBank using the BLAST algorithm and were consequently aligned using CLUSTAL W. A phylogenetic tree was reconstructed using TREECON W package and the neighbor-joining algorithm.
Results and discussion
Enrichment and isolation of pure cultures of lithotrophic SRB from soda lake sediments
Standard electron donors (H2/acetate, lactate, EtOH) for enrichment and isolation of SRB with sulfate as electron acceptor, although supporting sulfate reduction, were very inefficient in case of soda lakes. In all such cultures acetogens (Tindallia, in particular) were taking over and it was not possible to purify the presented SRB by serial dilution. We can only speculate about the reasons behind the phenomenon that acetogens may take selective advantage of utilization of sodium-based bioenergetics at such conditions (Müller 2003). There were two types of conditions eliminating this problem. In case of the mentioned enrichment conditions a switch to formate as electron donor and, in case of EtOH to sulfite as electron acceptor resulted in the suppression of acetogens and eventually allowed the isolation of pure cultures of SRB. Likewise, the most active and successful enrichments were started with formate as electron donor. The second, even more selective, set of enrichment conditions was elimination of organic electron donor leaving only acetate as a carbon source and replacement of sulfate to thiosulfate. This approach selected for a very specific metabolism presented only in a few SRB (Bak and Pfennig 1987; Rabus et al. 2006)—inorganic dismutation (disproportionation of thiosulfate):
Apparently, in the case of the SRB from the soda lakes, it was very successful, allowing the isolation of pure cultures in a single serial dilution series. With these 2 approaches, fifteen strains of SRB were obtained from the soda lake sediments at pH 10 and salinities of 0.6 and 2 M of total Na+. Most of the strains were recovered from the soda lakes of the Kulunda Steppe at moderate salt conditions. The enrichments and isolation process at 2 M Na+ were much less successful due to a very slow development of the enrichments and final failure to purify the SRB component. A summary of the results is presented in Table 2.
Identification of the isolates
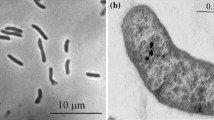
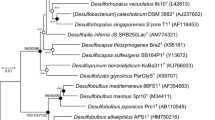
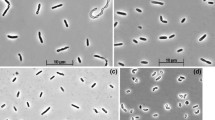
According to the phylogenetic analysis, the isolates belonged to the genera Desulfonatronovibrio (which was the dominant group), and Desufonatronum in the order Desulfovibrionales (Fig. 1). However, none of the isolates were closely related (i.e., <97% 16S rRNA sequence similarity) to described species. The eleven Desulfonatronovibrio isolates formed two subgroups. One included 10 strains with small cells (Fig. 2) and enriched either with organic electron donors or under thiosulfate dismutation conditions. The DNA–DNA homology within this group was in the range from 54 to 90%. The second phylotype was represented by a single strain AHT22 with large cells and isolated at thiosulfate dismutation conditions. Interestingly, it was co-enriched with a member of the group 1, strain AHT21 (Fig. 2; Supplementary Figure 1), and was finally separated from the numerically dominating AHT21 by replacing thiosulfate to sulfite. Both groups had relatively low values of the DNA homology (<40%) with the type species D. hydrogenovorans.
Phylogenetic position based on 16S rRNA gene sequence analysis of lithotrophic SRB isolates from soda lake sediments within the Deltaproteobacteria. The type strains are marked by (T) and the other haloalkaliphilic SRB species are in bold. The family and order definition is according to Kuever et al. (2005). The scale bar represents 5 nucleotide changes per 100 nucleotides. The percentage of bootstraps was derived from 1000 resampling using neighbor-joining algorithm, only values greater than 90% are indicated as black nodes on the branches
Cell morphology (phase contrast microphotographs of novel SRB isolates from soda lakes. a–e Desulfonatronovibrio isolates, f–i Desulfonatronum isolates. a Strain AHT9, b strain ASO3-2, c strain ASO4-5, d mixed thiosulfate-dismutating enrichment culture containing two different species of Desulfonatronovibrio (strains AHT20 and AHT22), e strain AHT22, f strain ASO4-1, h strain ASO4-2, g strain ASO3-6, i strain AHT30
The common occurrence and apparent easiness for growth of the Desulfonatronovibrio strains from the soda lake habitats by dismutation is an important ecological trait, since this pathway has hitherto been shown only in resting cells of this SRB species (Sydow et al. 2002). Until now, growth by thiosulfate dismutation had been shown exclusively for the species Desulfonatronum thiodismutans (Pikuta et al. 2003) and, later for extremely natronophilic genus Desulfonatronospira (Sorokin et al. 2008).
Four SRB isolates formed two novel phylotypes within the genus Desulfonatronum (Fig. 1). They were enriched and isolated only at moderate salinity using several electron donors, including ethanol and pyruvate, and sulfite as electron acceptor, apart from the standard combination of formate as e-donor, acetate as C-source and sulfate as e-acceptor. In the latter case, it is not completely clear how they could outcompete the Desulfonatronovibrio spp. It is interesting that, despite the ability of pure cultures within this group to grow by thiosulfate dismutation, they never dominated in the dismutating enrichments. In a syntrophic culture on benzoate and sulfate, the sulfide production was associated with the activity of members of the genus Desulfonatronum. The dominant SRB, strain AHT30, obtained from this culture, was affiliated with one of the two novel subgroups within the genus. The strains have bigger cells than most of the Desulfonatronovibrio isolates and are motile by a single polar flagellum. The cells have a strong tendency for aggregation, especially when grown on EtOH (Fig. 2; Supplementary Figure 1). The DNA–DNA homology between the representatives of two phylotypes was within the range of 40–50% and below 35% with the type species of the genus (D. lacustre).
The phylogenetic affiliation of the novel isolates confirmed the results of our previous culture-independent molecular studies in which multiple unknown dsrAB-based phylotypes of Desulfonatronovibrio and Desulfonatronum were detected in the Kulunda soda lakes (Foti et al. 2007).
Comparison of the PLFA profiles with the type strains (Supplementary Table) showed substantial differences between the representative strains of the two novel Desulfonatronovibrio phylotypes and between them and the type species confirming the phylogenetic differentiation. In contrast, two novel phylotypes of Desulfonatronum had similar PLFA profiles with each other and with the other species within the genus, indicating a higher level of conservation of this feature in this particular SRB group.
Growth and metabolic characteristics of the isolates
All isolates were able to grow with H2 and formate as electron donors, either autotrophically (also in the absence of yeast extract in the standard media) or in the presence of acetate as C-source, which is a common trend in the order Desulfovibrionales including the alkaliphilic genera Desulfonatronovibrio and Desulfonatronum (Table 3). However, there were also important growth characteristics not shown for the previously described species in these genera, particularly fermentation of pyruvate and growth by dismutation of thiosulfate/sulfite in the genus Desulfonatronovibrio. In general, the biomass growth of the novel isolates of this genus was very poor but they produced relatively high concentrations of sulfide (up to 25 mM), especially when thiosulfate served as the electron acceptor. The maximum growth rate was observed for strain AHT22 growing with formate/acetate and thiosulfate (0.040 h−1). Desulfonatronum isolates grew best with EtOH and sulfite/thiosulfate with maximum growth rate of 0.055 h−1 (strain ASO3-6), while sulfide production was still maximal with formate as the electron donor and thiosulfate as electron acceptor (up to 30 mM) (Table 4).
pH and salt profiles
All isolates grew well at pH 10. The pH profiles for growth obtained for several representatives indicated that they belonged to obligate alkaliphiles with a pH optimum within the range from 9.3 to 10 (Fig. 3a, c). As is typical for the soda lake alkaliphiles, none of the isolated SRB required NaCl for growth and, therefore, must be regarded as (natrono)philes, rather than (halo)philes. In respect to salt tolerance, the members of the genus Desulfonatronovibrio were clearly more adapted than the Desulfonatronum isolates to grow at salt concentrations above 1 M Na+. Several Desulfonatronovibrio strains were even capable of slow growth at Na+ concentrations 2.5–3.0 M, which is higher than reported for the type species (Fig. 3b, d). That probably explains the fact that in enrichments at 2 M Na+ only the members of Desulfonatronovibrio were selected. Compatible solute analysis (Supplementary Figure 2) of the most salt-tolerant Desulfonatronovibrio strain AHT9 grown at 2.5 M Na+ demonstrated the presence of two types of organic osmolytes: sucrose (5% w/w) and N-AGGN (N-acetylglutaminylglutamine amide) (approx. 2% w/w). The latter is a relatively rare type that is present in non-halophilic bacteria stressed by salts (Sagot et al. 2010). In this respect, Desulfonatronovibrio is significantly different from closely related extremely natronophilic genus Desulfonatronospira with a similar metabolism, which synthesizes glycine betaine as the main compatible solute (Sorokin et al. 2008).
Activity of resting cells grown at different conditions
One of the common trends in majority of the tested strains was a very active elemental sulfur reduction by nongrowing washed cells, while none of the strains could use elemental sulfur as an electron acceptor during growth (Tables 5, 6). This has already been observed for other haloalkaliphilic SRB species from soda lakes (Sorokin et al. 2008, 2010b). A possible explanation is the toxicity for growth of polysulfide formed as a stable product during sulfur reduction in the abiotic reaction with sulfide. Since sulfur reduction occurs with comparable rates both in thiosulfate and sulfate/sulfite-grown cells, it may be concluded that either it is a property of the sulfate-reducing system or that there is a constitutive sulfur-reductase. The former is more likely.
Another interest in performing activity tests was to compare cells grown with external electron donors, such as formate, and at dismutating conditions. The results proved that for the novel Desulfonatronovibrio and Desulfonatronum isolates thiosulfate and sulfite dismutation is active even if the cells were grown with an external electron donor indicating that the enzymes involved in dismutation are part of the “normal” sulfate reduction pathway (Tables 5, 6). On the other hand, there were clear differences in the dismutation activity of cells grown at different conditions and between strains with different isolation history. In the two most actively dismutating strains, Desulfonatronum ASO3-6 and Desulfonatronovibrio AHT22, cells grown at thiosulfate-dismutating conditions reacted only slightly to the addition of an external electron donor (formate) in case of thiosulfate reduction, while sulfite dismutation was completely inhibited in the presence of formate (i.e., the cells shifted to dissimilatory sulfite reduction). When the cells of these two organisms were grown at thiosulfate/sulfate-reducing conditions with formate as the electron donor, they could actively dismutate both thiosulfate and sulfite, but the addition of formate had pronounced inhibitory effect on thiosulfate dismutation. The results indicated that the dismutating and sulfate/thiosulfate-reducing systems in studied haloalkaliphilic SRB are, at least partly, overlapped. The only recent detailed analysis of the neutraphilic SRB Desulfocapsa sulfoexigens capable of growth by dismutation of thiosulfate and sulfur demonstrated that the thiosulfate-dismutating system most probably includes a combination of sulfate-reducing enzymes working in reverse (APS reductase, ATP sulfurilase), thiosulfate reductase and sulfite dehydrogenase which is the enzyme of the sulfur-oxidizing bacteria (Frederiksen and Finster 2003). The necessity for the latter may reflect the observed influence of growth history on the dismutating activity of the cells grown at different conditions.
In conclusion, this work demonstrated increased diversity of the lithotrophic SRB populations belonging to the genera Desulfonatronum and Desulfonatronovibrio in soda lake sediments. They are obligately alkaliphilic and moderately salt tolerant with a definite tendency for growth by thiosulfate dismutation, which has not been previously demonstrated for the genus Desulfonatronovibrio. On the basis of distinct phylogenetic, genetic and phenotypic properties, 11 Desulfonatronovibrio isolates are proposed to be accommodated in two novel species, D. thiodismutans and D. magnus. Likewise, four Desulfonatronum isolates are proposed as two novel species, D. thioautotrophicum and D. thiosulfatophilum.
References
Bak F, Pfennig N (1987) Chemolithotrophic growth of Desulfovibrio sulfodismutans sp. nov. by disproportionation of inorganic sulfur compounds. Arch Microbiol 147:184–189
Banciu HL, Sorokin DY, Tourova TP, Galinski EA, Muntyan MS, Kuenen JG, Muyzer G (2008) Influence of salts and pH on growth and activity of a novel facultatively alkaliphilic, extremely salt-tolerant, obligately chemolithoautotrophic sulfur-oxidizing gamma-proteobacterium Thioalkalibacter halophilus gen. nov., sp. nov. from south-western Siberian soda lakes. Extremophiles 12:391–404
De Ley J, Caffon H, Reinaerts A (1970) The quantitative measurements of hybridisation DNA from renaturation rates. Eur J Biochem 12:133–140
Foti M, Sorokin DY, Lomans B, Mussman M, Zakharova EE, Pimenov NV, Kuenen JG, Muyzer G (2007) Diversity, activity and abundance of sulfate-reducing bacteria in saline and hypersaline soda lakes. Appl Environ Microbiol 73:2093–2100
Foti M, Sorokin DY, Zacharova EE, Pimenov NV, Kuenen JG, Muyzer G (2008) Bacterial diversity and activity along a salinity gradient in soda lakes of the Kulunda Steppe (Altai, Russia). Extremophiles 12:133–145
Frederiksen T-M, Finster K (2003) Sulfite oxido-reductase is involved in the oxidation of sulfite in Desulfocapsa sulfoexigens during disproportionation of thiosulfate and elemental sulfur. Biodegradation 14:189–198
Gorlenko VM (2007) Anoxygenic phototrophic bacteria from soda lakes. In: Transactions of the Winogradsky Institute of Microbiology, vol XIV. Nauka, Moscow (in Russian), pp 159–183
Gorlenko VM, Namsaraev BB, Kulyrova AV, Zavarzina DG, Zhilina TN (1999) Activity of sulfate-reducing bacteria in the sediments of the soda lakes in south-east Transbaikal area. Microbiology (Moscow, English Translation) 68:580–586
Kuever J, Rayney FA, Widdel F (2005) Order II. Desulfovibrionales ord. nov. In: Garrity GM, Brenner DJ, Krieg NR, Staley JT (eds) Bergey’s manual of systematic bacteriology, vol 2, part C, Class IV Deltaproteobacteria. Springer, New York, pp 926–936
Kulp TR, Hoeft SE, Miller LG, Saltikov C, Murphy JN, Han S, Lanoil B, Oremland RS (2006) Dissimilatory arsenate and sulfate reduction in sediments of two hypersaline, arsenic-rich soda lakes: Mono and Searles Lakes, California. Appl Environ Microbiol 72:6514–6526
Kulp TR, Han S, Saltikov CV, Lanoil BD, Zargar K, Oremland RS (2007) Effects of imposed salinity gradients on dissimilatory arsenate reduction, sulfate reduction, and other microbial processes in sediments from two California soda lakes. Appl Environ Microbiol 73:5130–5137
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–177
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with Folin phenol reagent. J Biol Chem 193:265–275
Marmur J (1961) A procedure for isolation of DNA from microorganisms. J Mol Biol 3:208–214
Marmur J, Doty P (1962) Determination of the base composition of deoxyribonucleic acid from microorganisms. J Mol Biol 5:109–118
Müller V (2003) Energy conservation in acetogenic bacteria. Appl Environ Microbiol 69:6345–6353
Pfennig N, Lippert KD (1966) Über das Vitamin B12—bedürfnis phototropher Schwefel bacterien. Arch Microbiol 55:245–256
Pikuta EV, Zhilina TN, Zavarzin GA, Kostrikina NA, Osipov GA, Rainey FA (1998) Desulfonatronum lacustre gen. nov., sp. nov.: a new alkaliphilic sulfate-reducing bacterium utilizing ethanol. Microbiology (Moscow, English translation) 67:105–113
Pikuta EV, Hoover RB, Bej AK, Marsic D, Whitman WB, Cleland D, Krader P (2003) Desulfonatronum thiodismutans sp. nov., a novel alkaliphilic, sulfate-reducing bacterium capable of lithoautotrophic growth. Int J Syst Evol Microbiol 53:1327–1332
Rabus R, Hansen TA, Widdel F (2006) Dissimilatory sulfate- and sulfur-reducing prokaryotes. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes. Ecophysiology and biochemistry, vol 2. Springer, New York, pp 659–768
Ryu J-H, Dahlgren RA, Gao S, Tanji KK (2004) Characterization of redox processes in shallow groundwater of Owens dry lake, California. Environ Sci Technol 38:5950–5957
Sagot B, Gaysinski M, Mehiric M, Guigonisd J-M, Le Ruduliera D, Alloinga G (2010) Osmotically induced synthesis of the dipeptide N-acetylglutaminylglutamine amide is mediated by a new pathway conserved among bacteria. PNAS 107:12652–12657
Scholten JCM, Joye SB, Hollibaugh JT, Murrell JC (2005) Molecular analysis of the sulfate reducing and archaeal community in a meromictic soda lake (Mono Lake, California) by targeting 16S rRNA, mcrA, apsA, and dsrAB genes. Microb Ecol 50:29–39
Sorokin DY, Gorlenko VM, Namsaraev BB, Namsaraev ZB, Lysenko AM, Eshinimaev BT, Khmelenina VN, Trotsenko YA, Kuenen JG (2004) Prokaryotic communities of the north-eastern Mongolian soda lakes. Hydrobiologia 522:235–248
Sorokin DY, Banciu H, Robertson LA, Kuenen JG (2006) Haloalkaliphilic sulfur-oxidizing bacteria. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes. Ecophysiology and biochemistry, vol 2. Springer, New York, pp 969–984
Sorokin DY, Tourova TP, Henstra AM, Stams AJM, Galinski EA, Muyzer G (2008) Sulfidogenesis at extremely haloalkaline conditions by Desulfonatronospira thiodismutans gen. nov., sp. nov., and Desulfonatronospira delicata sp. nov.—a novel lineage of Deltaproteobacteria from hypersaline soda lakes. Microbiology (UK) 154:1444–1453
Sorokin DY, Rusanov II, Pimenov NV, Tourova TP, Abbas B, Muyzer G (2010a) Sulfidogenesis at extremely haloalkaline conditions in soda lakes of Kulunda Steppe (Altai, Russia). FEMS Microbiol Ecol 73:278–290
Sorokin DY, Detkova EN, Muyzer G (2010b) Propionate and butyrate dependent bacterial sulfate reduction at extremely haloalkaline conditions and description of Desulfobotulus alkaliphilus sp. nov. Extremophiles 14:71–77
Sydow U, Wohland P, Wolke I, Cypionka H (2002) Bioenergetics of the alkaliphilic sulfate-reducing bacterium Desulfonatronovibrio hydrogenovorans. Microbiology 148:853–860
Trüper HG, Schlegel HG (1964) Sulfur metabolism in Thiorhodaceae. 1. Quantitative measurements on growing cells of Chromatium okenii. Ant van Leeuwenhoek 30:225–238
Waldron PJ, Petsch ST, Martini AM, Nüsslein K (2007) Salinity constraints on subsurface archaeal diversity and methanogenesis in sedimentary rock rich in organic matter. Appl Environ Microbiol 73:4171–4179
Widdel F, Bak F (1992) Gram-negative mesophilic sulfate-reducing bacteria. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer K-H (eds) The prokaryotes, vol 3, 2nd edn. Springer, New York, pp 3352–3378
Zhilina TN, Zavarzin GA, Rainey FA, Pikuta EN, Osipov GA, Kostrikina NA (1997) Desulfonatronovibrio hydrogenovorans gen. nov., sp. nov., an alkaliphilic, sulfate-reducing bacterium. Int J Syst Bacteriol 47:144–149
Zhilina TN, Zavarzina DG, Kuever J, Lysenko AM, Zavarzin GA (2005) Desulfonatronum cooperativum sp. nov., a novel hydrogenotrophic, alkaliphilic, sulfate-reducing bacterium, from a syntrophic culture growing on acetate. Int J Syst Evol Microbiol 55:1001–1006
Acknowledgments
This work was supported by RFBR (Grant 10-04-00152). We are grateful to Marlene Stein (Institute of Microbiology and Biotechnology, University of Bonn) for the NMR analysis of compatible solutes.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Oren.
Nucleotide sequence accession number: The GenBank/EMBL accession numbers of the 16S rRNA gene sequences determined in this study are FJ469577-FJ469580, GU196826-GU196832, HM750217 and GQ863493.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Appendix
Appendix
Description of Desulfonatronovibrio thiodismutans sp. nov.
(thi.o.dis.mu’tans Gr. n. theion (Latin transliteration thium), sulfur; L. particle dis, in two, apart; L. part. adj. mutans, changing, altering; N.L. part. adj. thiodismutans, dismutating sulfur compounds)
Cells are vibrio-shaped, 0.3–0.4 × 1–3 μm, sometimes forming long coils, motile by means of a single polar flagellum. Strictly anaerobic, utilizing H2, formate and pyruvate as electron donor and sulfate, sulfite and thiosulfate as electron acceptor. Elemental sulfur can be reduced by resting cells. Lithoautotrophic growth occurs in some of the strains on H2 and formate. Can grow by pyruvate fermentation and by inorganic fermentation (dismutation, disproportionation) of thiosulfate and sulfite either autotrophically or in the presence of acetate as C-source. Elemental sulfur can be reduced by resting cells but no growth was observed. Obligately alkaliphilic with a pH range for growth between 8.5 and 10.5 (opt. 9.5–10.0). Moderately salt-tolerant with a total Na+ range for growth at pH 10 from 0.2 to 2.0–3.0 M (optimum at 0.4–0.6 M). Mesophilic, with a maximum temperature for growth at 40–42°C. The dominant PLFA are C18:0 and C18:1ω7c. The G+C content of the genomic DNA is within the range of 41.8–42.9 mol% (T m). Includes eleven isolates from sediments of soda lakes in Kulunda Steppe (Altai, Russia) and Owens Lake (California). The type strain AHT9T (DSM 21540T = UNIQEM U754T) was isolated from the soda lake Tanatar-5 in south-eastern Siberia. The GenBank 16S rRNA gene sequence accession number of the type strain is FJ469579.
Description of Desulfonatronovibrio magnus sp. nov.
(mag’nus L. masc. adj. magnus, large, great)
Cells are vibrio-shaped, 0.8–1.0 × 2–3 μm, motile by means of a single polar flagellum. Strictly anaerobic, utilizing H2, formate and pyruvate as electron donor and sulfate, sulfite and thiosulfate as electron acceptor. Elemental sulfur can be reduced by resting cells but no growth was observed. Autotrophic growth is not observed. Can grow by pyruvate fermentation and by inorganic fermentation (dismutation, disproportionation) of thiosulfate and sulfite in the presence of acetate as C-source. Obligately alkaliphilic with a pH range for growth between 8.5 and 10.5 (opt. 10.0). Moderately salt-tolerant with a total Na+ range for growth at pH 10 from 0.3 to 2.0 M (optimum at 0.4 M). Mesophilic, with a maximum temperature for growth at 41°C. The dominant PLFA include iso15:0, anteiso15:0, iso16:0, 16:0, iso17:1ω8, anteiso17:1ω7 and 18:1ω7c. The G+C content of the genomic DNA is 43.0 mol% (T m). The type strain is AHT22T (DSM 24400T = UNIQEM U844T), and isolated from sediments of soda lake Tanatar-5 in Kulunda Steppe (Altai, Russia). The GenBank 16S rRNA gene sequence accession number of the type strain is GU196831.
Amended description of the genus Desulfonatronovibrio (Zhilina et al. 1997)
In addition to the single strain-based genus diagnosis, the investigation of multiple sets of novel isolates showed that some representatives of this genus could grow autotrophically with H2 and formate and by pyruvate fermentation. Furthermore, the ability to grow by thiosulfate/sulfite dismutation, either autotrophically or in the presence of acetate as C-source, is a common property for this taxon.
Description of Desulfonatronum thioautotrophicum sp. nov.
(thi.o.au.to’tro.phi.cum Gr. n. theion (Latin transliteration thium), sulfur; Gr. pref. auto, self; Gr. neut. adj. trophikon, nursing, tending; N.L. neut. adj. thioautotrophicum, autotrophic with sulfur compounds)
Cells are vibrio-shaped, 0.5–0.6 × 2–4 μm, sometimes in chains and aggregates, motile by means of a single thick (tube-like) polar flagellum. Strictly anaerobic, utilizing H2, formate, EtOH, lactate and pyruvate as electron donor and sulfate, sulfite and thiosulfate as electron acceptor. Elemental sulfur can be reduced by resting cells but no growth was observed. Lithoautorophic growth occurs on H2 and formate. Can grow by pyruvate fermentation and by inorganic fermentation (dismutation, disproportionation) of thiosulfate and sulfite autotrophically. Obligately alkaliphilic with a pH range for growth between 8.3 and 10.5 (opt. 9.3–10.0). Moderately salt-tolerant with a total Na+ range for growth at pH 10 from 0.1 to 2.3 M (optimum at 0.4–0.6 M). Mesophilic, with a maximum temperature for growth at 40–41°C. The dominant PLFA are iso17:1ω8, iso15:0, 18:1ω7c iso15:0 and C14:0. The G+C content of the genomic DNA is 55.7–57.0 mol% (T m). Includes 2 isolates from sediments of soda lakes in Kulunda Steppe (Altai, Russia). The type strain ASO4-1T (DSM 21337T = UNIQEM U756T) was isolated from the soda lake Tanatar-1. The GenBank 16S rRNA gene sequence accession number of the type strain is FJ469577.
Description of Desulfonatronum thiosulfatophilum sp. nov.
(thi.o.sul.fa.to’phi.lum. N.L. n. thiosulfatum thiosulfate; Gr. adj. philos loving; N.L. neut. adj. thiosulfatophilum thiosulfate-loving)
Cells are vibrio-shaped, 0.4–0.5 × 1.5–4 μm, mostly doubled, motile by means of a single polar flagellum. Strictly anaerobic, utilizing H2, formate, EtOH, lactate and pyruvate as electron donor and sulfate, sulfite and thiosulfate as electron acceptor. Elemental sulfur can be reduced by resting cells but no growth was observed. It favors thiosulfate as the electron acceptor over sulfate and sulfite. Lithoautorophic growth is not observed. Can grow by pyruvate fermentation and by inorganic fermentation (dismutation, disproportionation) of thiosulfate and sulfite in the presence of acetate as C-source. Obligately alkaliphilic with a pH range for growth between 8.0 and 10.4 (opt. 9.5). Moderately salt-tolerant with a total Na+ range for growth at pH 10 from 0.1 to 1.3 M (optimum at 0.3 M). Mesophilic, with a maximum temperature for growth at 40°C. The dominant PLFA are 16:1ω7c, iso17:1ω8, 18:1ω7c, iso15:0 and C14:0. The G+C content of the genomic DNA is 55.7–57.0 mol% (T m). Includes 2 isolates from sediments of soda lakes in Kulunda Steppe (Altai, Russia). The type strain ASO4-2T (DSM 21338T = UNIQEM U757T) was isolated from the soda lake Picturesque. The GenBank 16S rRNA gene sequence accession number of the type strain is FJ469578.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Sorokin, D.Y., Tourova, T.P., Kolganova, T.V. et al. Culturable diversity of lithotrophic haloalkaliphilic sulfate-reducing bacteria in soda lakes and the description of Desulfonatronum thioautotrophicum sp. nov., Desulfonatronum thiosulfatophilum sp. nov., Desulfonatronovibrio thiodismutans sp. nov., and Desulfonatronovibrio magnus sp. nov.. Extremophiles 15, 391–401 (2011). https://doi.org/10.1007/s00792-011-0370-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-011-0370-7